Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
1017: Lily mitosis 07
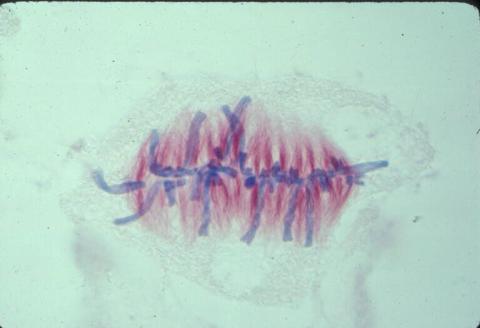
1017: Lily mitosis 07
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and have lined up in the middle of the dividing cell.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1018, 1019, and 1021.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1018, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
2307: Cells frozen in time
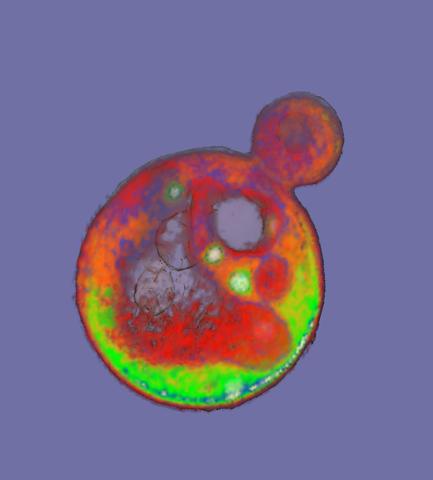
2307: Cells frozen in time
The fledgling field of X-ray microscopy lets researchers look inside whole cells rapidly frozen to capture their actions at that very moment. Here, a yeast cell buds before dividing into two. Colors show different parts of the cell. Seeing whole cells frozen in time will help scientists observe cells' complex structures and follow how molecules move inside them.
Carolyn Larabell, University of California, San Francisco, and the Lawrence Berkeley National Laboratory
View Media
2755: Two-headed Xenopus laevis tadpole
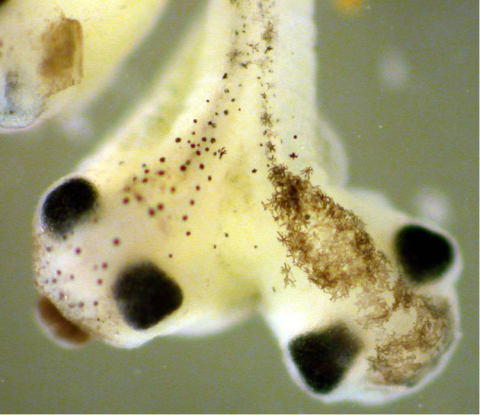
2755: Two-headed Xenopus laevis tadpole
Xenopus laevis, the African clawed frog, has long been used as a research organism for studying embryonic development. The abnormal presence of RNA encoding the signaling molecule plakoglobin causes atypical signaling, giving rise to a two-headed tadpole.
Michael Klymkowsky, University of Colorado, Boulder
View Media
7023: Dynein moving along microtubules
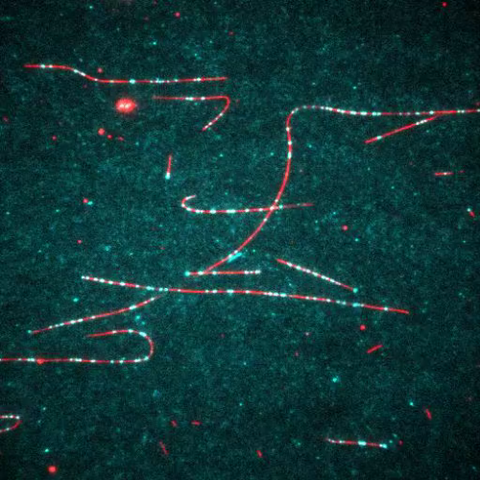
7023: Dynein moving along microtubules
Dynein (green) is a motor protein that “walks” along microtubules (red, part of the cytoskeleton) and carries its cargo along with it. This video was captured through fluorescence microscopy.
Morgan DeSantis, University of Michigan.
View Media
2441: Hydra 05
2441: Hydra 05
Hydra magnipapillata is an invertebrate animal used as a model organism to study developmental questions, for example the formation of the body axis.
Hiroshi Shimizu, National Institute of Genetics in Mishima, Japan
View Media
1284: Ion channels
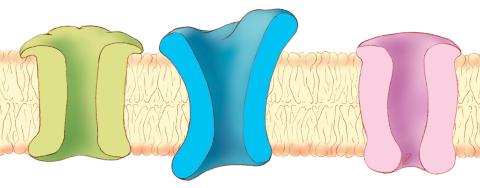
1284: Ion channels
The body uses a variety of ion channels to transport small molecules across cell membranes.
Judith Stoffer
View Media
3480: Cancer Cells Glowing from Luciferin
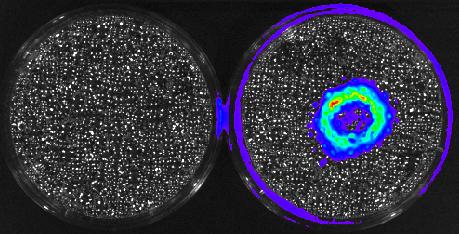
3480: Cancer Cells Glowing from Luciferin
The activator cancer cell culture, right, contains a chemical that causes the cells to emit light when in the presence of immune cells.
Mark Sellmyer, Stanford University School of Medicine
View Media
1282: Lysosomes
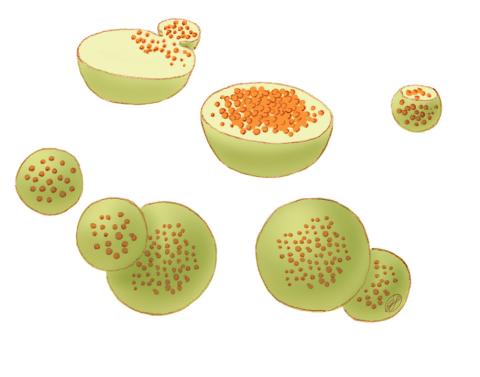
1282: Lysosomes
Lysosomes have powerful enzymes and acids to digest and recycle cell materials.
Judith Stoffer
View Media
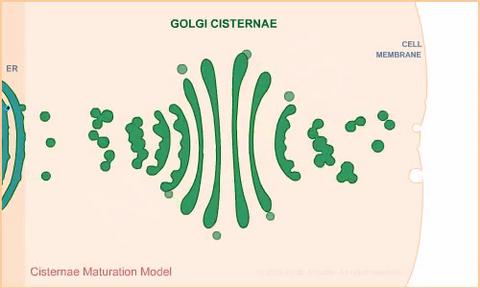
1307: Cisternae maturation model
1307: Cisternae maturation model
Animation for the cisternae maturation model of Golgi transport.
Judith Stoffer
View Media
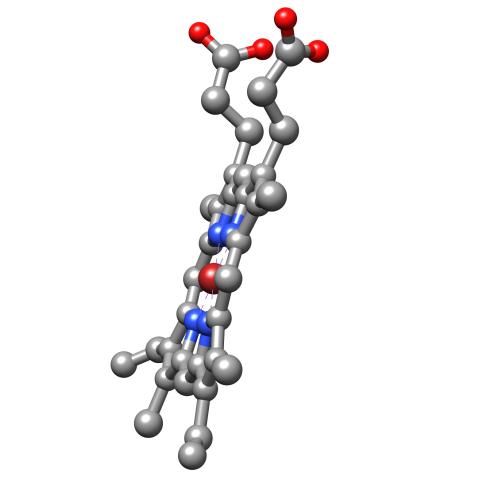
3540: Structure of heme, side view
3540: Structure of heme, side view
Molecular model of the struture of heme. Heme is a small, flat molecule with an iron ion (dark red) at its center. Heme is an essential component of hemoglobin, the protein in blood that carries oxygen throughout our bodies. This image first appeared in the September 2013 issue of Findings Magazine. View side view of heme here 3539.
Rachel Kramer Green, RCSB Protein Data Bank
View Media
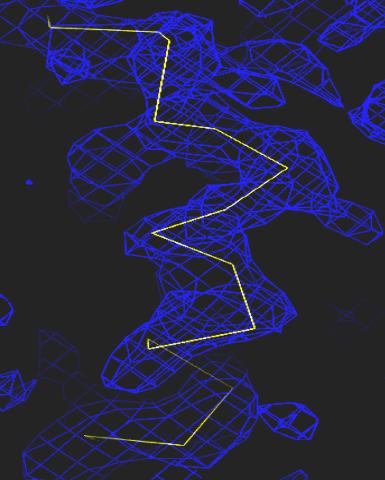

2354: Section of an electron density map
2354: Section of an electron density map
Electron density maps such as this one are generated from the diffraction patterns of X-rays passing through protein crystals. These maps are then used to generate a model of the protein's structure by fitting the protein's amino acid sequence (yellow) into the observed electron density (blue).
The Southeast Collaboratory for Structural Genomics
View Media
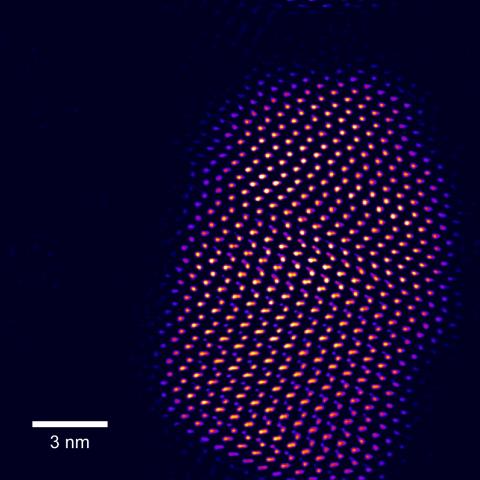

2332: Tiny points of light in a quantum dot
2332: Tiny points of light in a quantum dot
This fingertip-shaped group of lights is a microscopic crystal called a quantum dot. About 10,000 times thinner than a sheet of paper, the dot radiates brilliant colors under ultraviolet light. Dots such as this one allow researchers to label and track individual molecules in living cells and may be used for speedy disease diagnosis, DNA testing, and screening for illegal drugs.
Sandra Rosenthal and James McBride, Vanderbilt University, and Stephen Pennycook, Oak Ridge National Laboratory
View Media
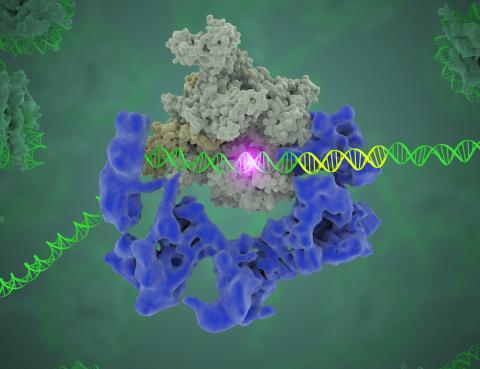
3766: TFIID complex binds DNA to start gene transcription
3766: TFIID complex binds DNA to start gene transcription
Gene transcription is a process by which the genetic information encoded in DNA is transcribed into RNA. It's essential for all life and requires the activity of proteins, called transcription factors, that detect where in a DNA strand transcription should start. In eukaryotes (i.e., those that have a nucleus and mitochondria), a protein complex comprising 14 different proteins is responsible for sniffing out transcription start sites and starting the process. This complex, called TFIID, represents the core machinery to which an enzyme, named RNA polymerase, can bind to and read the DNA and transcribe it to RNA. Scientists have used cryo-electron microscopy (cryo-EM) to visualize the TFIID-RNA polymerase-DNA complex in unprecedented detail. In this illustration, TFIID (blue) contacts the DNA and recruits the RNA polymerase (gray) for gene transcription. The start of the transcribed gene is shown with a flash of light. To learn more about the research that has shed new light on gene transcription, see this news release from Berkeley Lab. Related to video 5730.
Eva Nogales, Berkeley Lab
View Media

2702: Thermotoga maritima and its metabolic network
2702: Thermotoga maritima and its metabolic network
A combination of protein structures determined experimentally and computationally shows us the complete metabolic network of a heat-loving bacterium.
View Media
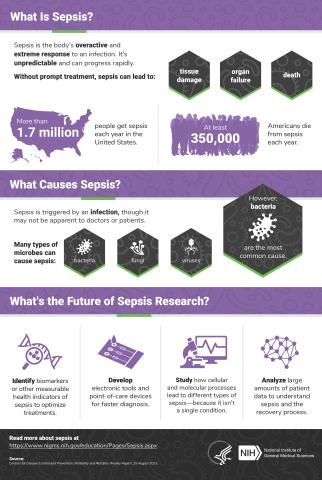

6536: Sepsis Infographic
6536: Sepsis Infographic
Sepsis is the body’s overactive and extreme response to an infection. More than 1.7 million people get sepsis each year in the United States. Without prompt treatment, sepsis can lead to tissue damage, organ failure, and death. Many NIGMS-supported researchers are working to improve sepsis diagnosis and treatment. Learn more with our sepsis featured topic page.
See 6551 for the Spanish version of this infographic.
See 6551 for the Spanish version of this infographic.
National Institute of General Medical Sciences
View Media
2791: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 02
2791: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 02
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
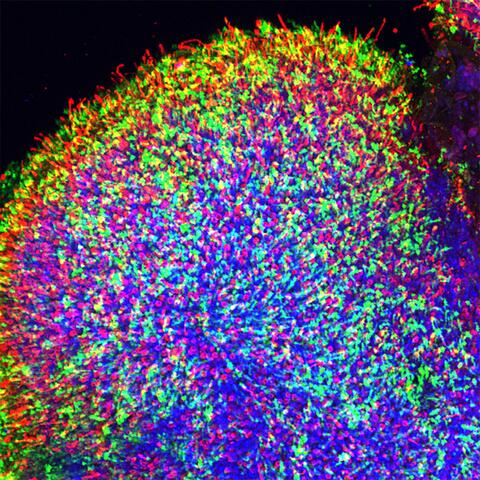
6748: Human retinal organoid
6748: Human retinal organoid
A replica of a human retina grown from stem cells. It shows rod photoreceptors (nerve cells responsible for dark vision) in green and red/green cones (nerve cells responsible for red and green color vision) in red. The cell nuclei are stained blue. This image was captured using a confocal microscope.
Kevin Eliceiri, University of Wisconsin-Madison.
View Media
6901: Mouse brain slice showing nerve cells
6901: Mouse brain slice showing nerve cells
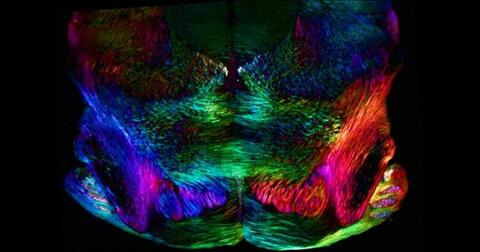
A 20-µm thick section of mouse midbrain. The nerve cells are transparent and weren’t stained. Instead, the color is generated by interaction of white polarized light with the molecules in the cells and indicates their orientation.
The image was obtained with a polychromatic polarizing microscope that shows the polychromatic birefringent image with hue corresponding to the slow axis orientation. More information about the microscopy that produced this image can be found in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak.
The image was obtained with a polychromatic polarizing microscope that shows the polychromatic birefringent image with hue corresponding to the slow axis orientation. More information about the microscopy that produced this image can be found in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media

6962: Trigonium diatom
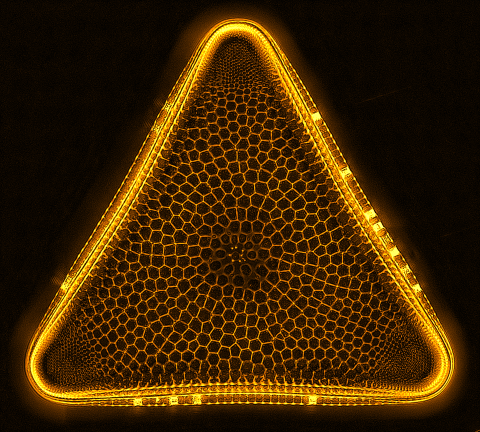
6962: Trigonium diatom
A Trigonium diatom imaged by a quantitative orientation-independent differential interference contrast (OI-DIC) microscope. Diatoms are single-celled photosynthetic algae with mineralized cell walls that contain silica and provide protection and support. These organisms form an important part of the plankton at the base of the marine and freshwater food chains. The width of this image is 90 μm.
More information about the microscopy that produced this image can be found in the Journal of Microscopy paper “An Orientation-Independent DIC Microscope Allows High Resolution Imaging of Epithelial Cell Migration and Wound Healing in a Cnidarian Model” by Malamy and Shribak.
More information about the microscopy that produced this image can be found in the Journal of Microscopy paper “An Orientation-Independent DIC Microscope Allows High Resolution Imaging of Epithelial Cell Migration and Wound Healing in a Cnidarian Model” by Malamy and Shribak.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media

6769: Culex quinquefasciatus mosquito larva
6769: Culex quinquefasciatus mosquito larva
A mosquito larva with genes edited by CRISPR. The red-orange glow is a fluorescent protein used to track the edits. This species of mosquito, Culex quinquefasciatus, can transmit West Nile virus, Japanese encephalitis virus, and avian malaria, among other diseases. The researchers who took this image developed a gene-editing toolkit for Culex quinquefasciatus that could ultimately help stop the mosquitoes from spreading pathogens. The work is described in the Nature Communications paper "Optimized CRISPR tools and site-directed transgenesis towards gene drive development in Culex quinquefasciatus mosquitoes" by Feng et al. Related to image 6770 and video 6771.
Valentino Gantz, University of California, San Diego.
View Media

1294: Stem cell differentiation
1294: Stem cell differentiation
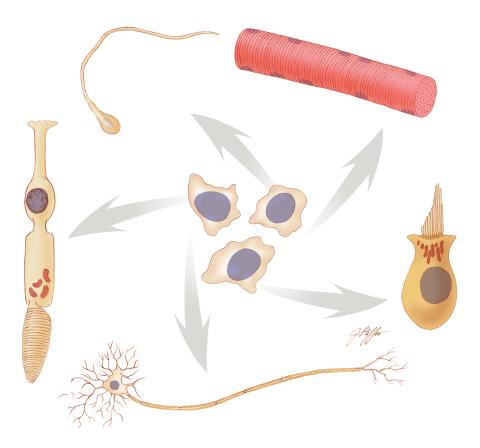
Undifferentiated embryonic stem cells cease to exist a few days after conception. In this image, ES cells are shown to differentiate into sperm, muscle fiber, hair cells, nerve cells, and cone cells.
Judith Stoffer
View Media
2809: Vimentin in a quail embryo
2809: Vimentin in a quail embryo
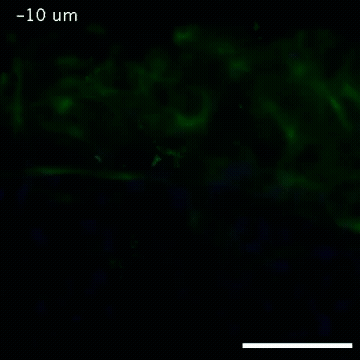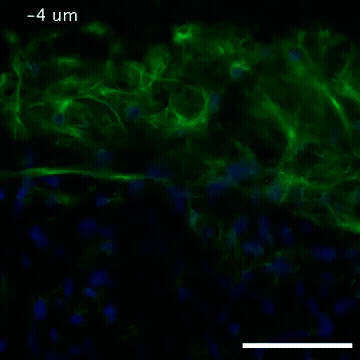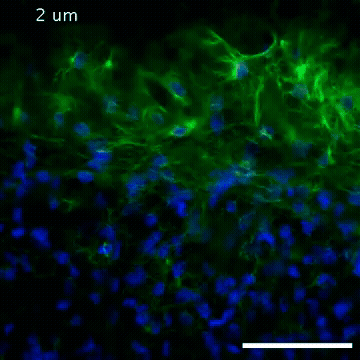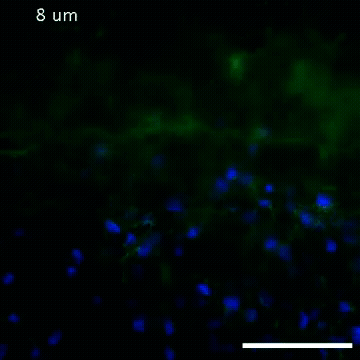
Video of high-resolution confocal images depicting vimentin immunofluorescence (green) and nuclei (blue) at the edge of a quail embryo yolk. These images were obtained as part of a study to understand cell migration in embryos. An NIGMS grant to Professor Garcia was used to purchase the confocal microscope that collected these images. Related to images 2807 and 2808.
Andrés Garcia, Georgia Tech
View Media
2320: Mapping disease spread
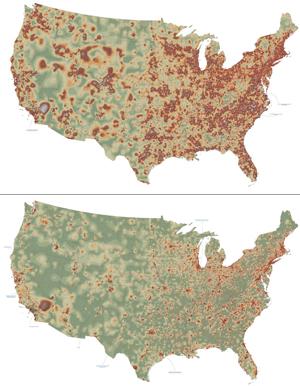
2320: Mapping disease spread
How far and fast an infectious disease spreads across a community depends on many factors, including transportation. These U.S. maps, developed as part of an international study to simulate and analyze disease spread, chart daily commuting patterns. They show where commuters live (top) and where they travel for work (bottom). Green represents the fewest number of people whereas orange, brown, and white depict the most. Such information enables researchers and policymakers to visualize how an outbreak in one area can spread quickly across a geographic region.
David Chrest, RTI International
View Media
6996: Measles virus proteins
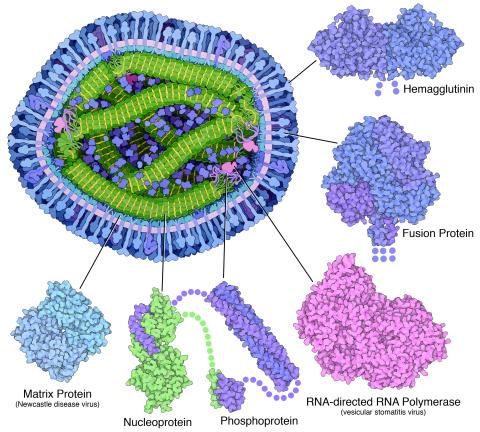
6996: Measles virus proteins
A cross section of the measles virus in which six proteins (enlarged on the outside of the virus) work together to infect cells. The measles virus is extremely infectious; 9 out of 10 people exposed will contract the disease. Fortunately, an effective vaccine protects against infection. Portions of the proteins that have not been determined are shown with dots.
Learn more about the six proteins on PDB 101’s Molecule of the Month: Measles Virus Proteins. Structures are available for the ordered regions of nucleoprotein and phosphoprotein (PDB entries 5E4V, 3ZDO, 1T6O), but the remaining regions are thought to form a flexible, random tangle. For a larger look at the measles virus, see 6995.
Learn more about the six proteins on PDB 101’s Molecule of the Month: Measles Virus Proteins. Structures are available for the ordered regions of nucleoprotein and phosphoprotein (PDB entries 5E4V, 3ZDO, 1T6O), but the remaining regions are thought to form a flexible, random tangle. For a larger look at the measles virus, see 6995.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
3747: Cryo-electron microscopy revealing the "wasabi receptor"
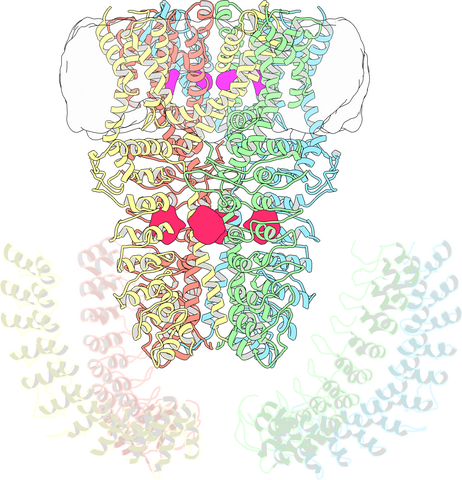
3747: Cryo-electron microscopy revealing the "wasabi receptor"
The TRPA1 protein is responsible for the burn you feel when you taste a bite of sushi topped with wasabi. Known therefore informally as the "wasabi receptor," this protein forms pores in the membranes of nerve cells that sense tastes or odors. Pungent chemicals like wasabi or mustard oil cause the pores to open, which then triggers a tingling or burn on our tongue. This receptor also produces feelings of pain in response to chemicals produced within our own bodies when our tissues are damaged or inflamed. Researchers used cryo-EM to reveal the structure of the wasabi receptor at a resolution of about 4 angstroms (a credit card is about 8 million angstroms thick). This detailed structure can help scientists understand both how we feel pain and how we can limit it by developing therapies to block the receptor. For more on cryo-EM see the blog post Cryo-Electron Microscopy Reveals Molecules in Ever Greater Detail.
Jean-Paul Armache, UCSF
View Media
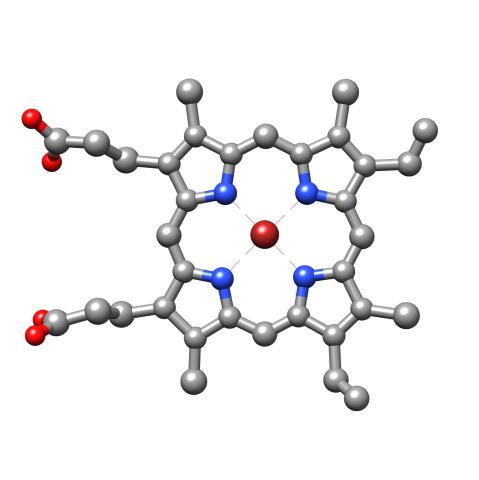
3539: Structure of heme, top view
3539: Structure of heme, top view
Molecular model of the struture of heme. Heme is a small, flat molecule with an iron ion (dark red) at its center. Heme is an essential component of hemoglobin, the protein in blood that carries oxygen throughout our bodies. This image first appeared in the September 2013 issue of Findings Magazine. View side view of heme here 3540.
Rachel Kramer Green, RCSB Protein Data Bank
View Media

3266: Biopixels
3266: Biopixels
Bioengineers were able to coax bacteria to blink in unison on microfluidic chips. This image shows a small chip with about 500 blinking bacterial colonies or biopixels. Related to images 3265 and 3268. From a UC San Diego news release, "Researchers create living 'neon signs' composed of millions of glowing bacteria."
Jeff Hasty Lab, UC San Diego
View Media
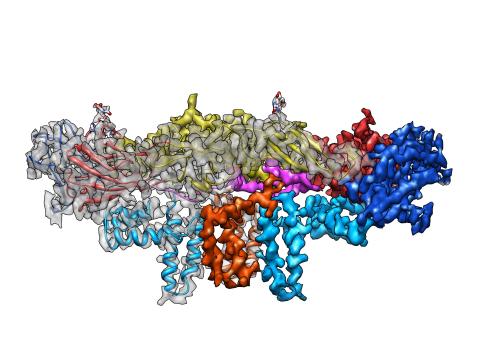
3758: Dengue virus membrane protein structure
3758: Dengue virus membrane protein structure
Dengue virus is a mosquito-borne illness that infects millions of people in the tropics and subtropics each year. Like many viruses, dengue is enclosed by a protective membrane. The proteins that span this membrane play an important role in the life cycle of the virus. Scientists used cryo-EM to determine the structure of a dengue virus at a 3.5-angstrom resolution to reveal how the membrane proteins undergo major structural changes as the virus matures and infects a host. The image shows a side view of the structure of a protein composed of two smaller proteins, called E and M. Each E and M contributes two molecules to the overall protein structure (called a heterotetramer), which is important for assembling and holding together the viral membrane, i.e., the shell that surrounds the genetic material of the dengue virus. The dengue protein's structure has revealed some portions in the protein that might be good targets for developing medications that could be used to combat dengue virus infections. For more on cryo-EM see the blog post Cryo-Electron Microscopy Reveals Molecules in Ever Greater Detail. You can watch a rotating view of the dengue virus surface structure in video 3748.
Hong Zhou, UCLA
View Media
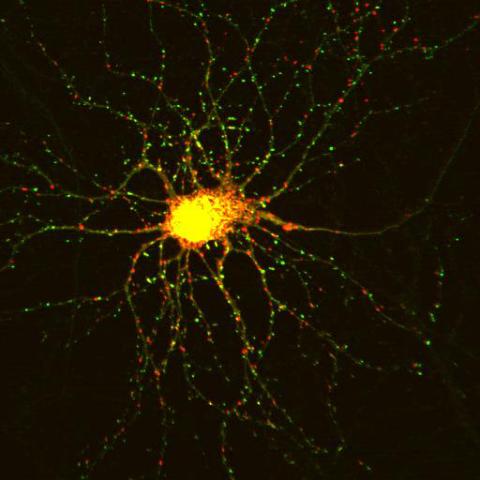
3509: Neuron with labeled synapses
3509: Neuron with labeled synapses
In this image, recombinant probes known as FingRs (Fibronectin Intrabodies Generated by mRNA display) were expressed in a cortical neuron, where they attached fluorescent proteins to either PSD95 (green) or Gephyrin (red). PSD-95 is a marker for synaptic strength at excitatory postsynaptic sites, and Gephyrin plays a similar role at inhibitory postsynaptic sites. Thus, using FingRs it is possible to obtain a map of synaptic connections onto a particular neuron in a living cell in real time.
Don Arnold and Richard Roberts, University of Southern California.
View Media
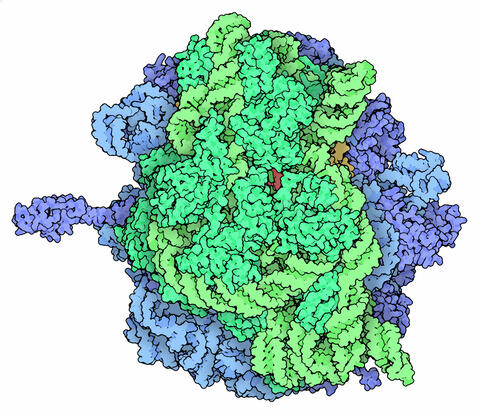
5780: Ribosome illustration from PDB
5780: Ribosome illustration from PDB
Ribosomes are complex machines made up of more than 50 proteins and three or four strands of genetic material called ribosomal RNA (rRNA). The busy cellular machines make proteins, which are critical to almost every structure and function in the cell. To do so, they read protein-building instructions, which come as strands of messenger RNA. Ribosomes are found in all forms of cellular life—people, plants, animals, even bacteria. This illustration of a bacterial ribosome was produced using detailed information about the position of every atom in the complex. Several antibiotic medicines work by disrupting bacterial ribosomes but leaving human ribosomes alone. Scientists are carefully comparing human and bacterial ribosomes to spot differences between the two. Structures that are present only in the bacterial version could serve as targets for new antibiotic medications.
From PDB’s Molecule of the Month collection (direct link: http://pdb101.rcsb.org/motm/121) Molecule of the Month illustrations are available under a CC-BY-4.0 license. Attribution should be given to David S. Goodsell and the RCSB PDB.
View Media
5878: Misfolded proteins within in the mitochondria
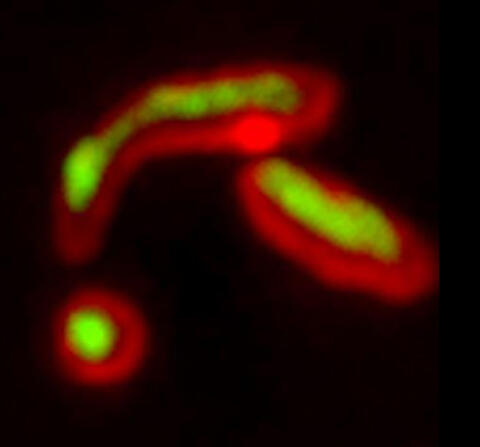
5878: Misfolded proteins within in the mitochondria
Misfolded proteins (green) within mitochondria (red). Related to video 5877.
Rong Li rong@jhu.edu Department of Chemical and Biomolecular Engineering, Whiting School of Engineering, Johns Hopkins University, Baltimore, Maryland 21218, USA.
View Media
2352: Human aspartoacylase
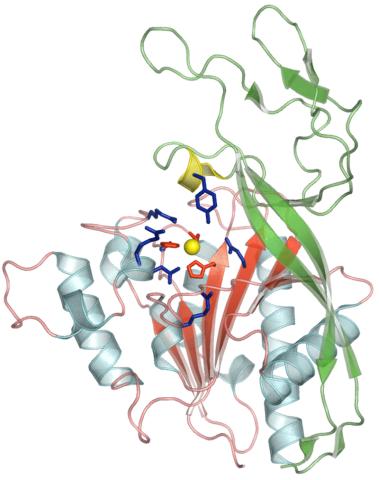
2352: Human aspartoacylase
Model of aspartoacylase, a human enzyme involved in brain metabolism.
Center for Eukaryotic Structural Genomics, PSI
View Media
3574: Cytonemes in developing fruit fly cells
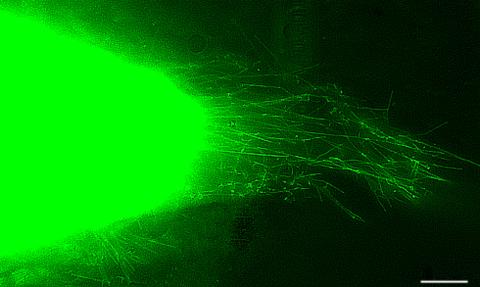
3574: Cytonemes in developing fruit fly cells
Scientists have long known that multicellular organisms use biological molecules produced by one cell and sensed by another to transmit messages that, for instance, guide proper development of organs and tissues. But it's been a puzzle as to how molecules dumped out into the fluid-filled spaces between cells can precisely home in on their targets. Using living tissue from fruit flies, a team led by Thomas Kornberg of the University of California, San Francisco, has shown that typical cells in animals can talk to each other via long, thin cell extensions called cytonemes (Latin for "cell threads") that may span the length of 50 or 100 cells. The point of contact between a cytoneme and its target cell acts as a communications bridge between the two cells.
Sougata Roy, University of California, San Francisco
View Media
5800: Mouse cerebellum in pink and blue
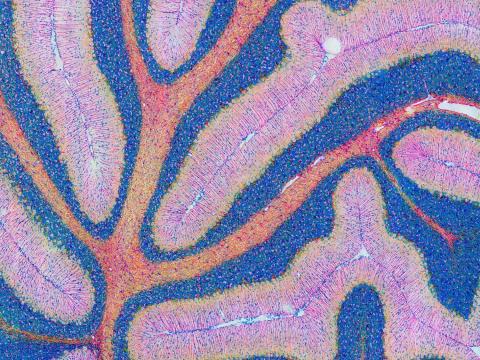
5800: Mouse cerebellum in pink and blue
The cerebellum is the brain's locomotion control center. Found at the base of your brain, the cerebellum is a single layer of tissue with deep folds like an accordion. People with damage to this region of the brain often have difficulty with balance, coordination and fine motor skills.
This image of a mouse cerebellum is part of a collection of such images in different colors and at different levels of magnification from the National Center for Microscopy and Imaging Research (NCMIR). Related to image 5795.
This image of a mouse cerebellum is part of a collection of such images in different colors and at different levels of magnification from the National Center for Microscopy and Imaging Research (NCMIR). Related to image 5795.
National Center for Microscopy and Imaging Research (NCMIR)
View Media
1247: Crab nerve cell
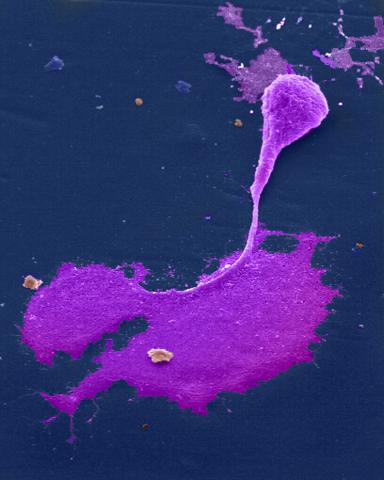
1247: Crab nerve cell
Neuron from a crab showing the cell body (bottom), axon (rope-like extension), and growth cone (top right).
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
2323: Motion in the brain
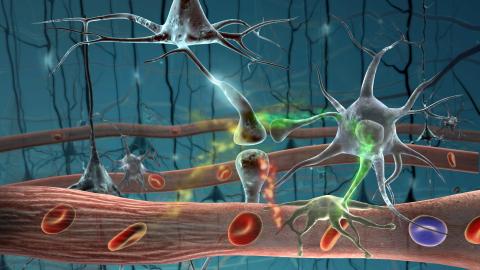
2323: Motion in the brain
Amid a network of blood vessels and star-shaped support cells, neurons in the brain signal each other. The mists of color show the flow of important molecules like glucose and oxygen. This image is a snapshot from a 52-second simulation created by an animation artist. Such visualizations make biological processes more accessible and easier to understand.
Kim Hager and Neal Prakash, University of California, Los Angeles
View Media
2764: Painted chromosomes
2764: Painted chromosomes
Like a paint-by-numbers picture, painted probes tint individual human chromosomes by targeting specific DNA sequences. Chromosome 13 is colored green, chromosome 14 is in red and chromosome 15 is painted yellow. The image shows two examples of fused chromosomes—a pair of chromosomes 15 connected head-to-head (yellow dumbbell-shaped structure) and linked chromosomes 13 and 14 (green and red dumbbell). These fused chromosomes—called dicentric chromosomes—may cause fertility problems or other difficulties in people.
Beth A. Sullivan, Duke University
View Media
3460: Prion protein fibrils 1
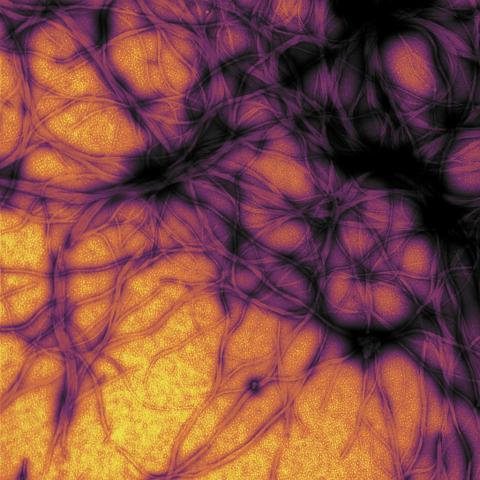
3460: Prion protein fibrils 1
Recombinant proteins such as the prion protein shown here are often used to model how proteins misfold and sometimes polymerize in neurodegenerative disorders. This prion protein was expressed in E. coli, purified and fibrillized at pH 7. Image taken in 2004 for a research project by Roger Moore, Ph.D., at Rocky Mountain Laboratories that was published in 2007 in Biochemistry. This image was not used in the publication.
Ken Pekoc (public affairs officer) and Julie Marquardt, NIAID/ Rocky Mountain Laboratories
View Media
3626: Bone cancer cell
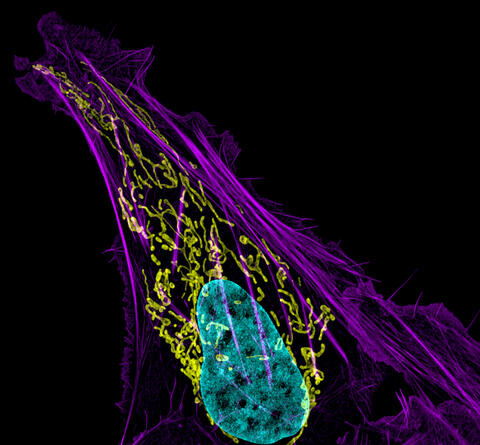
3626: Bone cancer cell
This image shows an osteosarcoma cell with DNA in blue, energy factories (mitochondria) in yellow, and actin filaments—part of the cellular skeleton—in purple. One of the few cancers that originate in the bones, osteosarcoma is rare, with about a thousand new cases diagnosed each year in the United States.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Dylan Burnette and Jennifer Lippincott-Schwartz, NICHD
View Media
2570: VDAC video 01
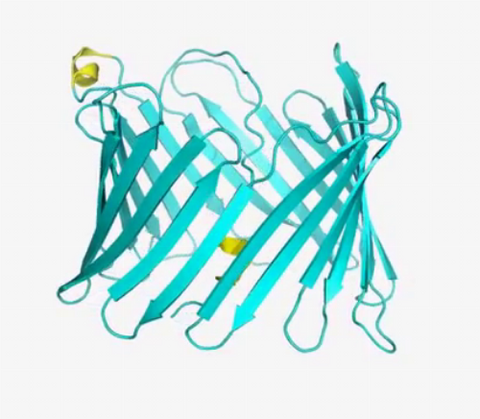
2570: VDAC video 01
This video shows the structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to videos 2571 and 2572.
Related to videos 2571 and 2572.
Gerhard Wagner, Harvard Medical School
View Media
3791: Nucleolus subcompartments spontaneously self-assemble 2
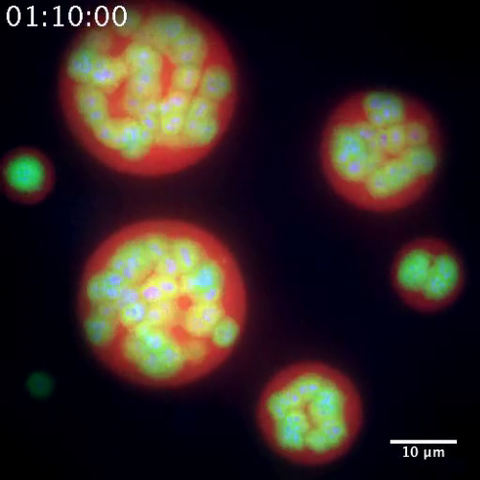
3791: Nucleolus subcompartments spontaneously self-assemble 2
The nucleolus is a small but very important protein complex located in the cell's nucleus. It forms on the chromosomes at the location where the genes for the RNAs are that make up the structure of the ribosome, the indispensable cellular machine that makes proteins from messenger RNAs.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This video of nucleoli in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows how each of the compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue) spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, image 3792 and image 3793.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This video of nucleoli in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows how each of the compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue) spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, image 3792 and image 3793.
Nilesh Vaidya, Princeton University
View Media
2566: Haplotypes
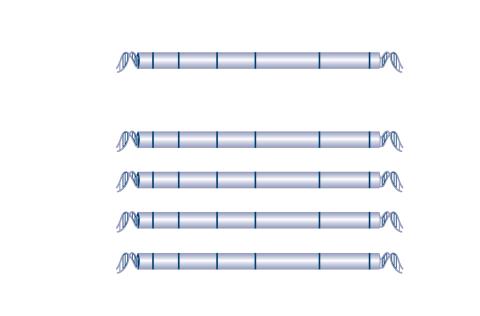
2566: Haplotypes
Haplotypes are combinations of gene variants that are likely to be inherited together within the same chromosomal region. In this example, an original haplotype (top) evolved over time to create three newer haplotypes that each differ by a few nucleotides (red). See image 2567 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

2396: Hen egg lysozyme (1)
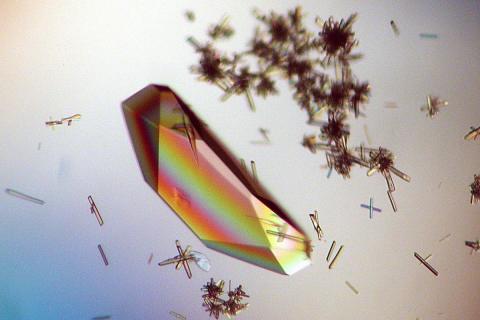
2396: Hen egg lysozyme (1)
Crystals of hen egg lysozyme protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
6774: Endoplasmic reticulum abnormalities 2
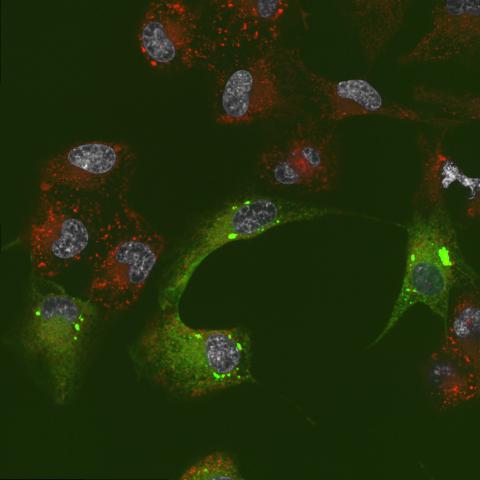
6774: Endoplasmic reticulum abnormalities 2
Human cells with the gene that codes for the protein FIT2 deleted. After an experimental intervention, they are expressing a nonfunctional version of FIT2, shown in green. The lack of functional FIT2 affected the structure of the endoplasmic reticulum (ER), and the nonfunctional protein clustered in ER membrane aggregates, seen as large bright-green spots. Lipid droplets are shown in red, and the nucleus is visible in gray. This image was captured using a confocal microscope. Related to image 6773.
Michel Becuwe, Harvard University.
View Media

6344: Drosophila
6344: Drosophila
Two adult fruit flies (Drosophila)
Dr. Vicki Losick, MDI Biological Laboratory, www.mdibl.org
View Media
7019: Bacterial cells aggregated above a light-organ pore of the Hawaiian bobtail squid
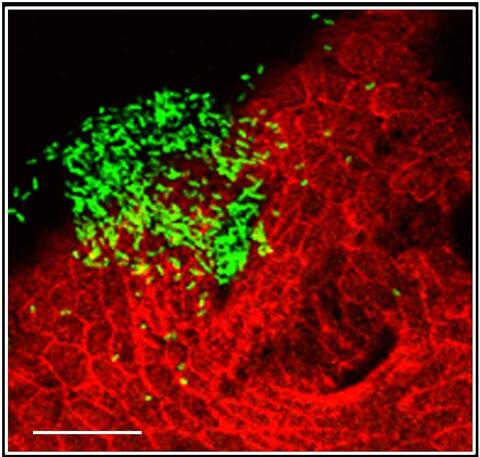
7019: Bacterial cells aggregated above a light-organ pore of the Hawaiian bobtail squid
The beating of cilia on the outside of the Hawaiian bobtail squid’s light organ concentrates Vibrio fischeri cells (green) present in the seawater into aggregates near the pore-containing tissue (red). From there, the bacterial cells (~2 mm) swim to the pores and migrate through a bottleneck into the interior crypts where a population of symbionts grow and remain for the life of the host. This image was taken using confocal fluorescence microscopy.
Related to images 7016, 7017, 7018, and 7020.
Related to images 7016, 7017, 7018, and 7020.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media

2401: Bacterial alpha amylase
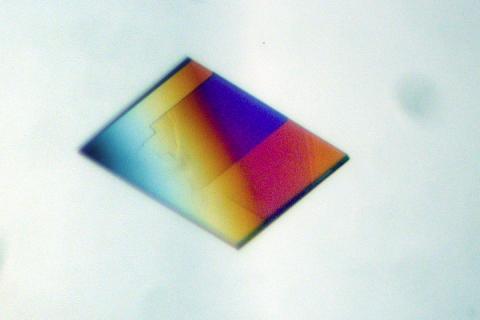
2401: Bacterial alpha amylase
A crystal of bacterial alpha amylase protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
3729: A molecular switch strips transcription factor from DNA
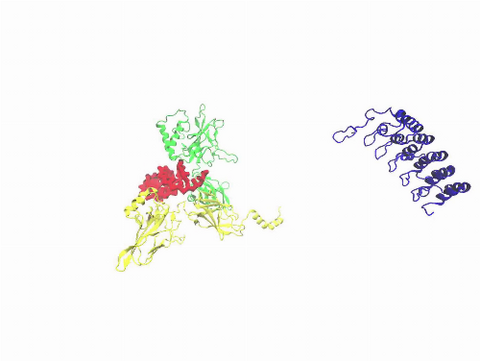
3729: A molecular switch strips transcription factor from DNA
In this video, Rice University scientists used molecular modeling with a mathematical algorithm called AWSEM (for associative memory, water-mediated, structure and energy model) and structural data to analyze how a transcription factor called nuclear factor kappa B (NFkB) is removed from DNA to stop gene activation. AWSEM uses the interacting energies of their components to predict how proteins fold. At the start, the NFkB dimer (green and yellow, in the center) grips DNA (red, to the left), which activates the transcription of genes. IkB (blue, to the right), an inhibitor protein, stops transcription when it binds to NFkB and forces the dimer to twist and release its hold on DNA. The yellow domain at the bottom of IkB is the PEST domain, which binds first to NFkB. For more details about this mechanism called molecular stripping, see here.
Davit Potoyan and Peter Wolynes
View Media
3754: Circadian rhythm neurons in the fruit fly brain
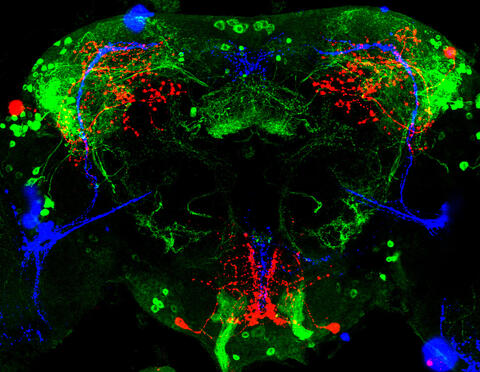
3754: Circadian rhythm neurons in the fruit fly brain
Some nerve cells (neurons) in the brain keep track of the daily cycle. This time-keeping mechanism, called the circadian clock, is found in all animals including us. The circadian clock controls our daily activities such as sleep and wakefulness. Researchers are interested in finding the neuron circuits involved in this time keeping and how the information about daily time in the brain is relayed to the rest of the body. In this image of a brain of the fruit fly Drosophila the time-of-day information flowing through the brain has been visualized by staining the neurons involved: clock neurons (shown in blue) function as "pacemakers" by communicating with neurons that produce a short protein called leucokinin (LK) (red), which, in turn, relays the time signal to other neurons, called LK-R neurons (green). This signaling cascade set in motion by the pacemaker neurons helps synchronize the fly's daily activity with the 24-hour cycle. To learn more about what scientists have found out about circadian pacemaker neurons in the fruit fly see this news release by New York University. This work was featured in the Biomedical Beat blog post Cool Image: A Circadian Circuit.
Justin Blau, New York University
View Media