Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
7009: Hungry, hungry macrophages
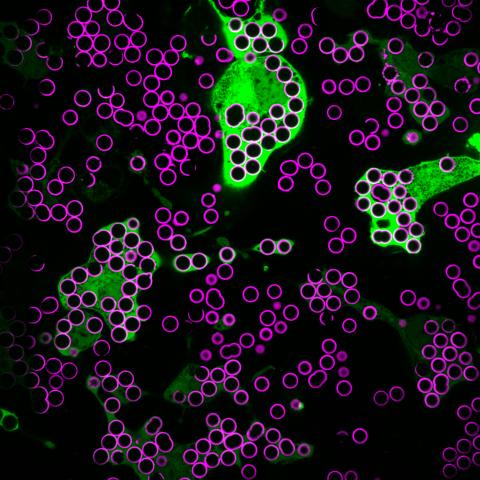
7009: Hungry, hungry macrophages
Macrophages (green) are the professional eaters of our immune system. They are constantly surveilling our tissues for targets—such as bacteria, dead cells, or even cancer—and clearing them before they can cause harm. In this image, researchers were testing how macrophages responded to different molecules that were attached to silica beads (magenta) coated with a lipid bilayer to mimic a cell membrane.
Find more information on this image in the NIH Director’s Blog post "How to Feed a Macrophage."
Find more information on this image in the NIH Director’s Blog post "How to Feed a Macrophage."
Meghan Morrissey, University of California, Santa Barbara.
View Media
3675: NCMIR kidney-1
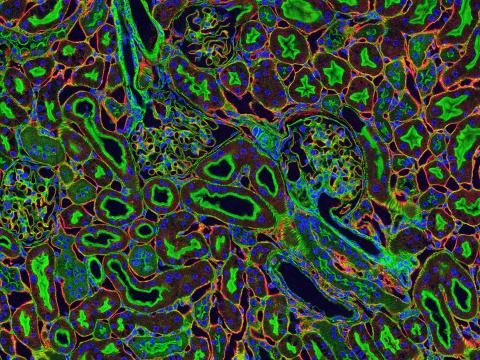
3675: NCMIR kidney-1
Stained kidney tissue. The kidney is an essential organ responsible for disposing wastes from the body and for maintaining healthy ion levels in the blood. It also secretes two hormones, erythropoietin (EPO) and calcitriol (a derivative of vitamin D), into the blood. It works like a purifier by pulling break-down products of metabolism, such as urea and ammonium, from the blood stream for excretion in urine. Related to image 3725.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
1306: Vesicular shuttle model
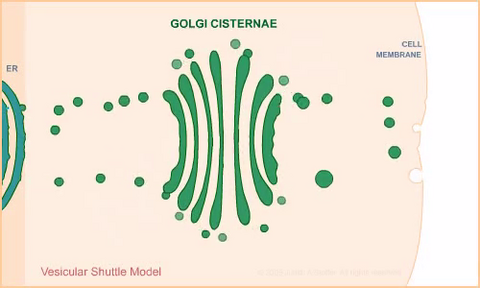
1306: Vesicular shuttle model
Animation for the vesicular shuttle model of Golgi transport.
Judith Stoffer
View Media
6593: Cell-like compartments from frog eggs 6
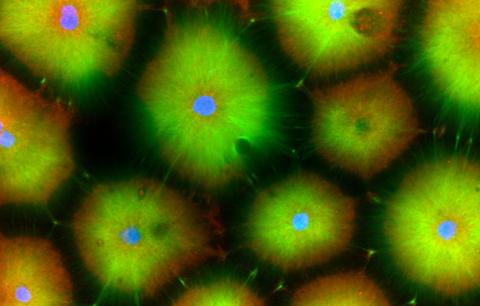
6593: Cell-like compartments from frog eggs 6
Cell-like compartments that spontaneously emerged from scrambled frog eggs, with nuclei (blue) from frog sperm. Endoplasmic reticulum (red) and microtubules (green) are also visible. Image created using confocal microscopy.
For more photos of cell-like compartments from frog eggs view: 6584, 6585, 6586, 6591, 6592.
For videos of cell-like compartments from frog eggs view: 6587, 6588, 6589, and 6590.
Xianrui Cheng, Stanford University School of Medicine.
View Media
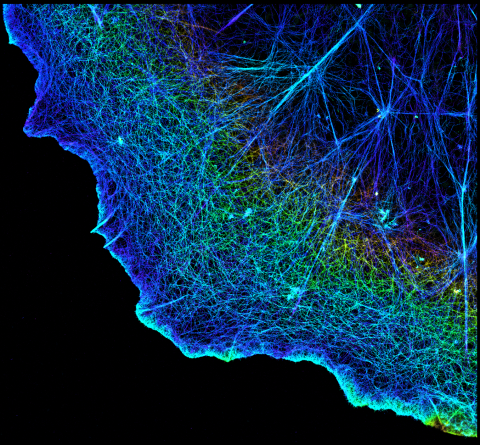
3749: 3D image of actin in a cell
3749: 3D image of actin in a cell
Actin is an essential protein in a cell's skeleton (cytoskeleton). It forms a dense network of thin filaments in the cell. Here, researchers have used a technique called stochastic optical reconstruction microscopy (STORM) to visualize the actin network in a cell in three dimensions. The actin strands were labeled with a dye called Alexa Fluor 647-phalloidin. This image appears in a study published by Nature Methods, which reports how researchers use STORM to visualize the cytoskeleton.
Xiaowei Zhuang, Howard Hughes Medical Institute, Harvard University
View Media
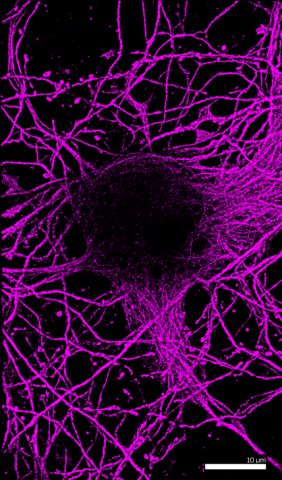
6890: Microtubules in hippocampal neurons
6890: Microtubules in hippocampal neurons
Microtubules (magenta) in neurons of the hippocampus, a part of the brain involved in learning and memory. Microtubules are strong, hollow fibers that provide structural support to cells. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6889, 6891, and 6892.
Related to images 6889, 6891, and 6892.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media

3344: Artificial cilia exhibit spontaneous beating
3344: Artificial cilia exhibit spontaneous beating
Researchers have created artificial cilia that wave like the real thing. Zvonimir Dogic and his Brandeis University colleagues combined just a few cilia proteins to create cilia that are able to wave and sweep material around--although more slowly and simply than real ones. The researchers are using the lab-made cilia to study how the structures coordinate their movements and what happens when they don't move properly. Featured in the August 18, 2011, issue of Biomedical Beat.
Zvonimir Dogic
View Media
1085: Natcher Building 05
1085: Natcher Building 05
NIGMS staff are located in the Natcher Building on the NIH campus.
Alisa Machalek, National Institute of General Medical Sciences
View Media
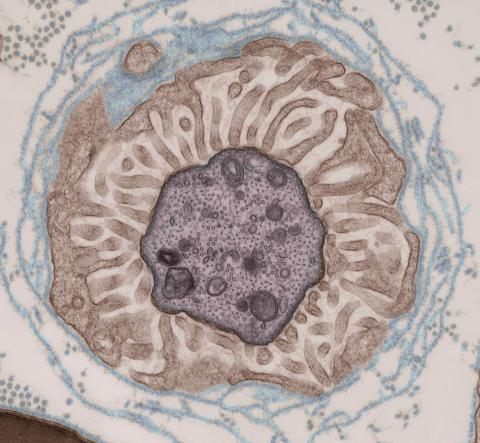
3740: Transmission electron microscopy showing cross-section of the node of Ranvier
3740: Transmission electron microscopy showing cross-section of the node of Ranvier
Nodes of Ranvier are short gaps in the myelin sheath surrounding myelinated nerve cells (axons). Myelin insulates axons, and the node of Ranvier is where the axon is exposed to the extracellular environment, allowing for the transmission of action potentials at these nodes via ion flows between the inside and outside of the axon. The image shows a cross-section through the node, with the surrounding extracellular matrix encasing and supporting the axon shown in cyan.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
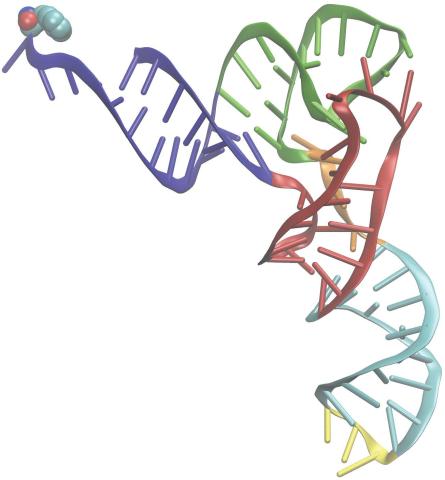
3406: Phenylalanine tRNA molecule
3406: Phenylalanine tRNA molecule
Phenylalanine tRNA showing the anticodon (yellow) and the amino acid, phenylalanine (blue and red spheres).
Patrick O'Donoghue and Dieter Soll, Yale University
View Media
3590: Fruit fly spermatids
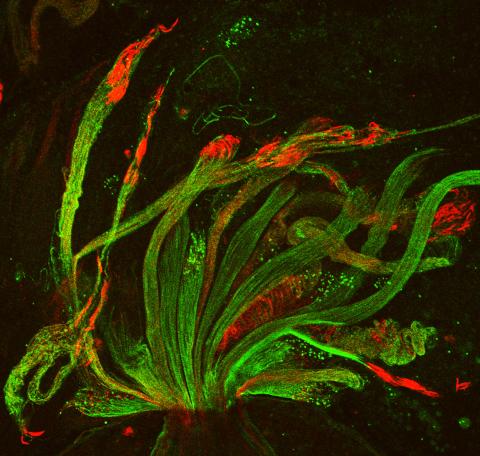
3590: Fruit fly spermatids
Developing spermatids (precursors of mature sperm cells) begin as small, round cells and mature into long-tailed, tadpole-shaped ones. In the sperm cell's head is the cell nucleus; in its tail is the power to outswim thousands of competitors to fertilize an egg. As seen in this microscopy image, fruit fly spermatids start out as groups of interconnected cells. A small lipid molecule called PIP2 helps spermatids tell their heads from their tails. Here, PIP2 (red) marks the nuclei and a cell skeleton-building protein called tubulin (green) marks the tails. When PIP2 levels are too low, some spermatids get mixed up and grow with their heads at the wrong end. Because sperm development is similar across species, studies in fruit flies could help researchers understand male infertility in humans.
Lacramioara Fabian, The Hospital for Sick Children, Toronto, Canada
View Media
3748: Cryo-electron microscopy of the dengue virus showing protective membrane and membrane proteins
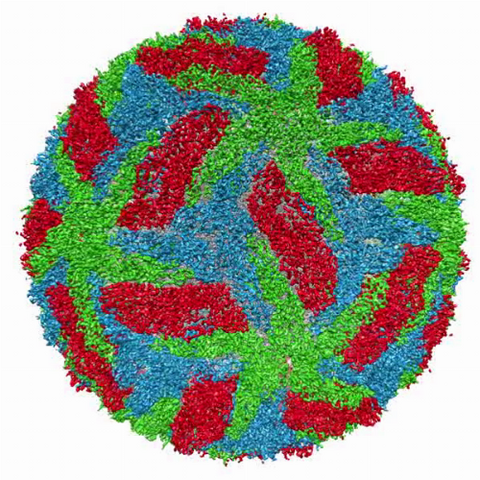
3748: Cryo-electron microscopy of the dengue virus showing protective membrane and membrane proteins
Dengue virus is a mosquito-borne illness that infects millions of people in the tropics and subtropics each year. Like many viruses, dengue is enclosed by a protective membrane. The proteins that span this membrane play an important role in the life cycle of the virus. Scientists used cryo-EM to determine the structure of a dengue virus at a 3.5-angstrom resolution to reveal how the membrane proteins undergo major structural changes as the virus matures and infects a host. For more on cryo-EM see the blog post Cryo-Electron Microscopy Reveals Molecules in Ever Greater Detail. Related to image 3756.
Hong Zhou, UCLA
View Media
6791: Yeast cells entering mitosis
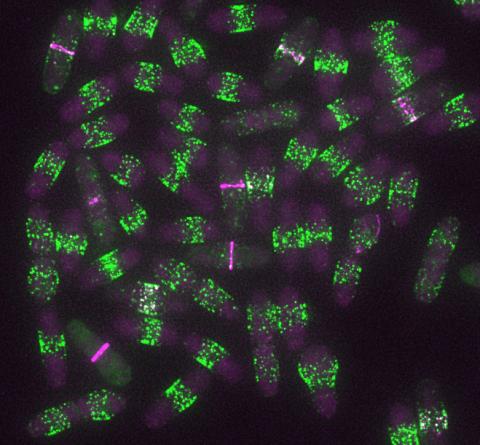
6791: Yeast cells entering mitosis
Yeast cells entering mitosis, also known as cell division. The green and magenta dots are two proteins that play important roles in mitosis. They show where the cells will split. This image was captured using wide-field microscopy with deconvolution.
Related to images 6792, 6793, 6794, 6797, 6798, and videos 6795 and 6796.
Related to images 6792, 6793, 6794, 6797, 6798, and videos 6795 and 6796.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media
5811: NCMIR Tongue 2
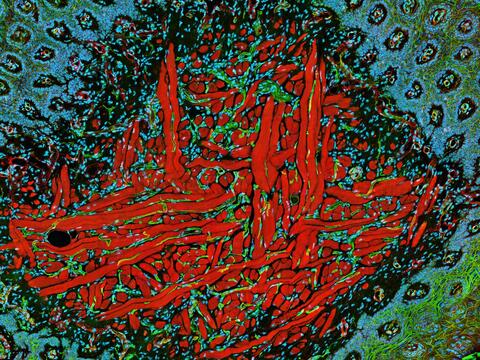
5811: NCMIR Tongue 2
Microscopy image of a tongue. One in a series of two, see image 5810
National Center for Microscopy and Imaging Research (NCMIR)
View Media
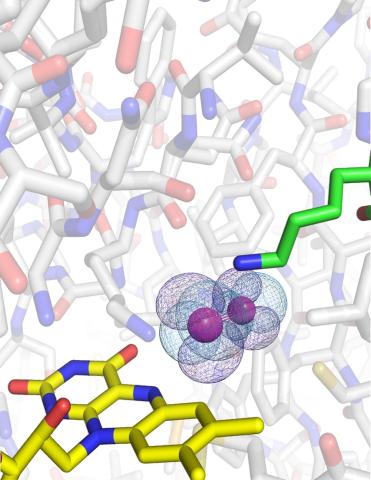
3411: O2 reacting with a flavin-dependent enzyme
3411: O2 reacting with a flavin-dependent enzyme
Department of Biological Chemistry, University of Michigan
View Media
6602: See how immune cell acid destroys bacterial proteins
6602: See how immune cell acid destroys bacterial proteins
This animation shows the effect of exposure to hypochlorous acid, which is found in certain types of immune cells, on bacterial proteins. The proteins unfold and stick to one another, leading to cell death.
American Chemistry Council
View Media

2748: Early ribbon drawing of a protein
2748: Early ribbon drawing of a protein
This ribbon drawing of a protein hand drawn and colored by researcher Jane Richardson in 1981 helped originate the ribbon representation of proteins that is now ubiquitous in molecular graphics. The drawing shows the 3-dimensional structure of the protein triose phosphate isomerase. The green arrows represent the barrel of eight beta strands in this structure and the brown spirals show the protein's eight alpha helices. A black and white version of this drawing originally illustrated a review article in Advances in Protein Chemistry, volume 34, titled "Anatomy and Taxonomy of Protein Structures." The illustration was selected as Picture of The Day on the English Wikipedia for November 19, 2009. Other important and beautiful images of protein structures by Jane Richardson are available in her Wikimedia gallery.
Jane Richardson, Duke University Medical Center
View Media
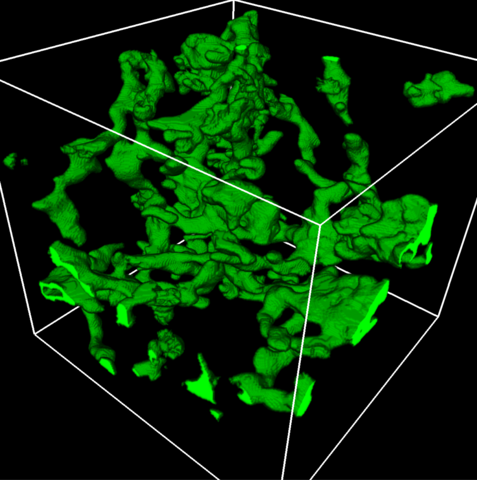
5857: 3D reconstruction of a tubular matrix in peripheral endoplasmic reticulum
5857: 3D reconstruction of a tubular matrix in peripheral endoplasmic reticulum
Detailed three-dimensional reconstruction of a tubular matrix in a thin section of the peripheral endoplasmic reticulum between the plasma membranes of the cell. The endoplasmic reticulum (ER) is a continuous membrane that extends like a net from the envelope of the nucleus outward to the cell membrane. The ER plays several roles within the cell, such as in protein and lipid synthesis and transport of materials between organelles. Shown here is a three-dimensional representation of the peripheral ER microtubules. Related to images 5855 and 5856
Jennifer Lippincott-Schwartz, Howard Hughes Medical Institute Janelia Research Campus, Virginia
View Media
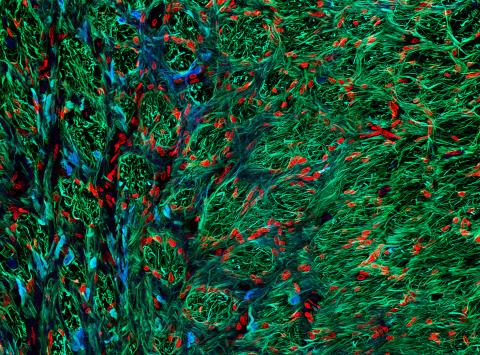
5852: Optic nerve astrocytes
5852: Optic nerve astrocytes
Astrocytes in the cross section of a human optic nerve head
Tom Deerinck and Keunyoung (“Christine”) Kim, NCMIR
View Media
2796: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 03
2796: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 03
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media

2404: Bovine milk alpha-lactalbumin (2)
2404: Bovine milk alpha-lactalbumin (2)
Crystals of bovine milk alpha-lactalbumin protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
3586: Human blood cells with Borrelia hermsii, a bacterium that causes relapsing fever
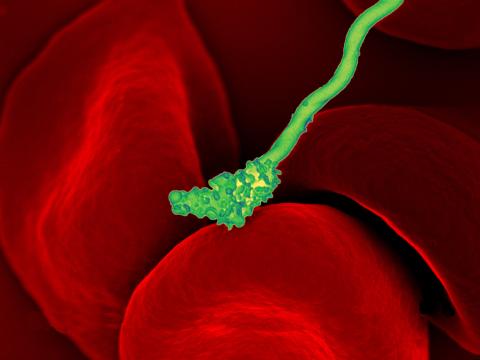
3586: Human blood cells with Borrelia hermsii, a bacterium that causes relapsing fever
Relapsing fever is caused by a bacterium and transmitted by certain soft-bodied ticks or body lice. The disease is seldom fatal in humans, but it can be very serious and prolonged. This scanning electron micrograph shows Borrelia hermsii (green), one of the bacterial species that causes the disease, interacting with red blood cells. Micrograph by Robert Fischer, NIAID.
For more information on this see, relapsing fever.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
For more information on this see, relapsing fever.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
NIAID
View Media
6807: Fruit fly ovaries
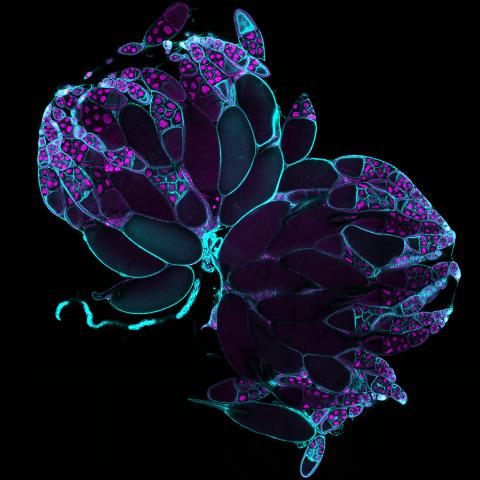
6807: Fruit fly ovaries
Fruit fly (Drosophila melanogaster) ovaries with DNA shown in magenta and actin filaments shown in light blue. This image was captured using a confocal laser scanning microscope.
Related to image 6806.
Related to image 6806.
Vladimir I. Gelfand, Feinberg School of Medicine, Northwestern University.
View Media
2365: Map of protein structures 01
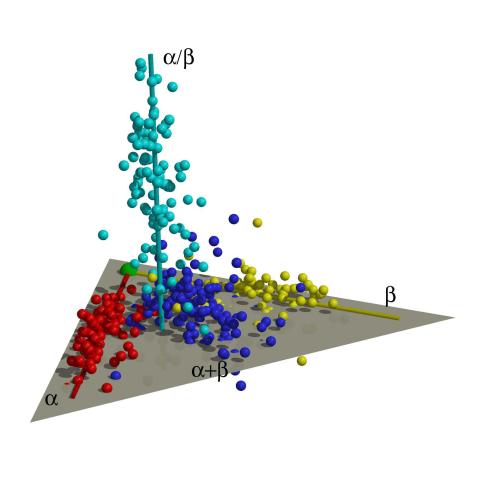
2365: Map of protein structures 01
A global "map of the protein structure universe." The Berkeley Structural Genomics Center has developed a method to visualize the vast universe of protein structures in which proteins of similar structure are located close together and those of different structures far away in the space. This map, constructed using about 500 of the most common protein folds, reveals a highly non-uniform distribution, and shows segregation between four elongated regions corresponding to four different protein classes (shown in four different colors). Such a representation reveals a high-level of organization of the protein structure universe.
Berkeley Structural Genomics Center, PSI
View Media
2733: Early development in Arabidopsis
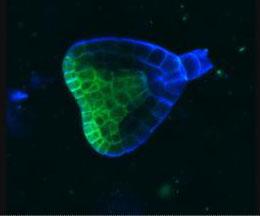
2733: Early development in Arabidopsis
Early on, this Arabidopsis plant embryo picks sides: While one end will form the shoot, the other will take root underground. Short pieces of RNA in the bottom half (blue) make sure that shoot-forming genes are expressed only in the embryo's top half (green), eventually allowing a seedling to emerge with stems and leaves. Like animals, plants follow a carefully orchestrated polarization plan and errors can lead to major developmental defects, such as shoots above and below ground. Because the complex gene networks that coordinate this development in plants and animals share important similarities, studying polarity in Arabidopsis--a model organism--could also help us better understand human development.
Zachery R. Smith, Jeff Long lab at the Salk Institute for Biological Studies
View Media
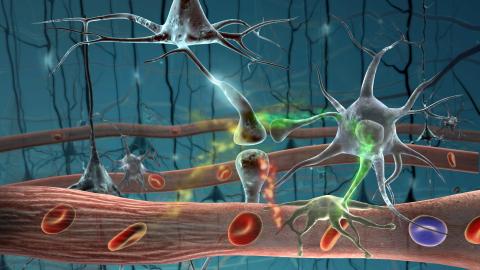
2323: Motion in the brain
2323: Motion in the brain
Amid a network of blood vessels and star-shaped support cells, neurons in the brain signal each other. The mists of color show the flow of important molecules like glucose and oxygen. This image is a snapshot from a 52-second simulation created by an animation artist. Such visualizations make biological processes more accessible and easier to understand.
Kim Hager and Neal Prakash, University of California, Los Angeles
View Media
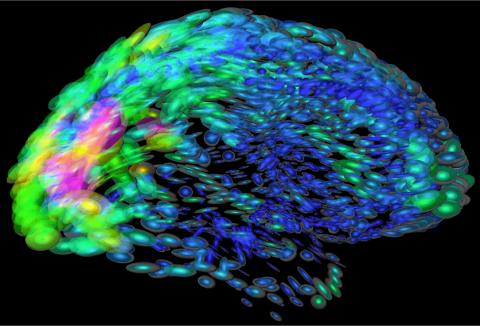
2419: Mapping brain differences
2419: Mapping brain differences
This image of the human brain uses colors and shapes to show neurological differences between two people. The blurred front portion of the brain, associated with complex thought, varies most between the individuals. The blue ovals mark areas of basic function that vary relatively little. Visualizations like this one are part of a project to map complex and dynamic information about the human brain, including genes, enzymes, disease states, and anatomy. The brain maps represent collaborations between neuroscientists and experts in math, statistics, computer science, bioinformatics, imaging, and nanotechnology.
Arthur Toga, University of California, Los Angeles
View Media
5760: Annotated TEM cross-section of C. elegans (roundworm)
5760: Annotated TEM cross-section of C. elegans (roundworm)
The worm Caenorhabditis elegans is a popular laboratory animal because its small size and fairly simple body make it easy to study. Scientists use this small worm to answer many research questions in developmental biology, neurobiology, and genetics. This image, which was taken with transmission electron microscopy (TEM), shows a cross-section through C. elegans, revealing various internal structures labeled in the image. You can find a high-resolution image without the annotations at image 5759.
The image is from a figure in an article published in the journal eLife.
The image is from a figure in an article published in the journal eLife.
Piali Sengupta, Brandeis University
View Media
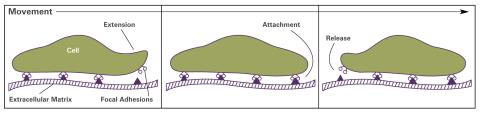
2503: Focal adhesions (with labels)
2503: Focal adhesions (with labels)
Cells walk along body surfaces via tiny "feet," called focal adhesions, that connect with the extracellular matrix. See image 2502 for an unlabeled version of this illustration.
Crabtree + Company
View Media
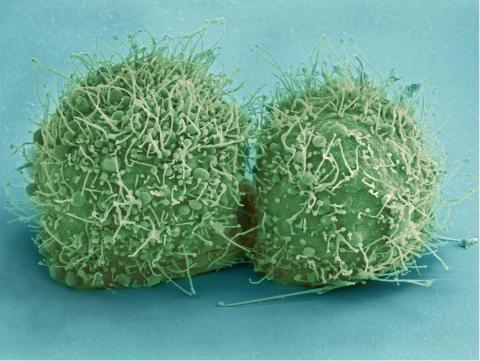
3518: HeLa cells
3518: HeLa cells
Scanning electron micrograph of just-divided HeLa cells. Zeiss Merlin HR-SEM. See related images 3519, 3520, 3521, 3522.
National Center for Microscopy and Imaging Research
View Media
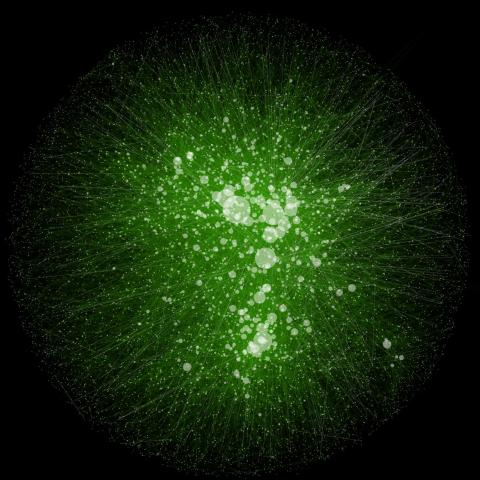

2749: Cytoscape network wiring diagram 2
2749: Cytoscape network wiring diagram 2
This image integrates the thousands of known molecular and genetic interactions happening inside our bodies using a computer program called Cytoscape. Images like this are known as network wiring diagrams, but Cytoscape creator Trey Ideker somewhat jokingly calls them "hairballs" because they can be so complicated, intricate and hard to tease apart. Cytoscape comes with tools to help scientists study specific interactions, such as differences between species or between sick and diseased cells. Related to 2737.
Trey Ideker, University of California, San Diego
View Media
3339: Single-Molecule Imaging
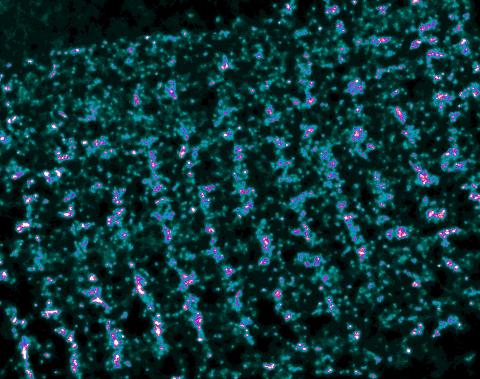
3339: Single-Molecule Imaging
This is a super-resolution light microscope image taken by Hiro Hakozaki and Masa Hoshijima of NCMIR. The image contains highlighted calcium channels in cardiac muscle using a technique called dSTORM. The microscope used in the NCMIR lab was built by Hiro Hakozaki.
Tom Deerinck, NCMIR
View Media
3489: Worm sperm
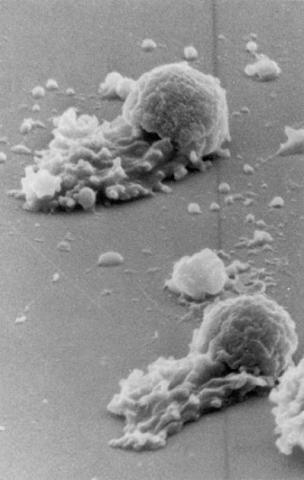
3489: Worm sperm
To develop a system for studying cell motility in unnatrual conditions -- a microscope slide instead of the body -- Tom Roberts and Katsuya Shimabukuro at Florida State University disassembled and reconstituted the motility parts used by worm sperm cells.
Tom Roberts, Florida State University
View Media

5752: Genetically identical mycobacteria respond differently to antibiotic 2
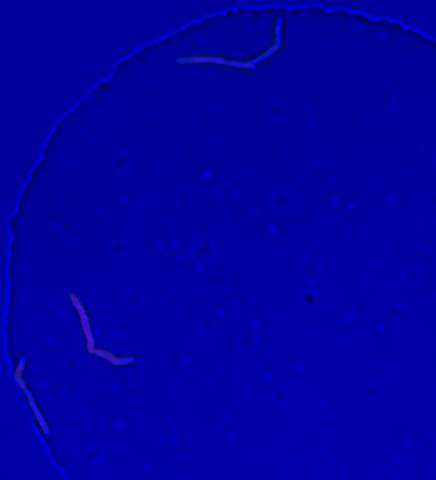
5752: Genetically identical mycobacteria respond differently to antibiotic 2
Antibiotic resistance in microbes is a serious health concern. So researchers have turned their attention to how bacteria undo the action of some antibiotics. Here, scientists set out to find the conditions that help individual bacterial cells survive in the presence of the antibiotic rifampicin. The research team used Mycobacterium smegmatis, a more harmless relative of Mycobacterium tuberculosis, which infects the lung and other organs to cause serious disease.
In this video, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to image 5751.
In this video, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to image 5751.
Bree Aldridge, Tufts University
View Media
2563: Epigenetic code (with labels)
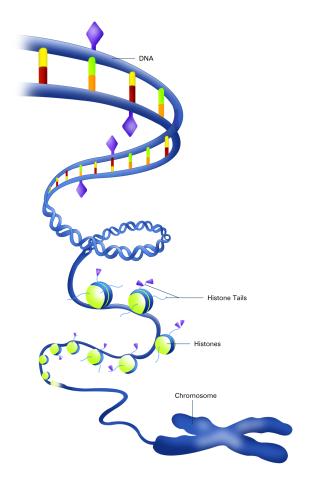
2563: Epigenetic code (with labels)
The "epigenetic code" controls gene activity with chemical tags that mark DNA (purple diamonds) and the "tails" of histone proteins (purple triangles). These markings help determine whether genes will be transcribed by RNA polymerase. Genes hidden from access to RNA polymerase are not expressed. See image 2562 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

1090: Natcher Building 10
1090: Natcher Building 10
NIGMS staff are located in the Natcher Building on the NIH campus.
Alisa Machalek, National Institute of General Medical Sciences
View Media
2518: ATP synthase (with labels)
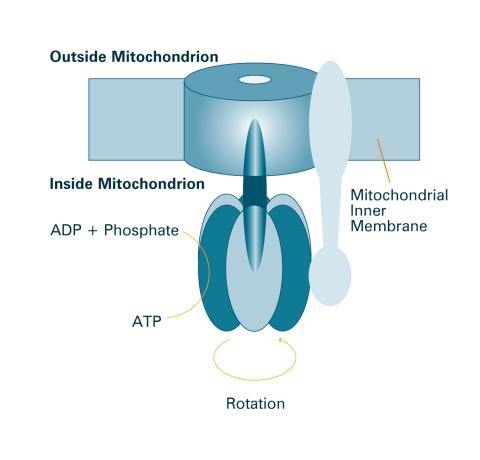
2518: ATP synthase (with labels)
The world's smallest motor, ATP synthase, generates energy for the cell. See image 2517 for an unlabeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
6893: Chromatin in human tenocyte
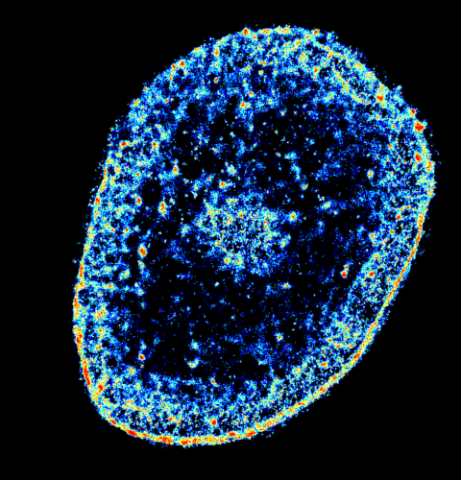
6893: Chromatin in human tenocyte
The nucleus of a degenerating human tendon cell, also known as a tenocyte. It has been color-coded based on the density of chromatin—a substance made up of DNA and proteins. Areas of low chromatin density are shown in blue, and areas of high chromatin density are shown in red. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6887 and 6888.
Related to images 6887 and 6888.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media
2350: Mandelate racemase from B. subtilis
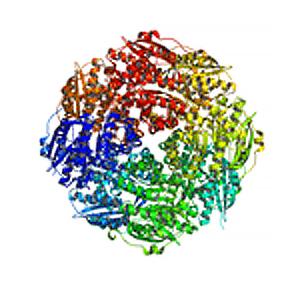
2350: Mandelate racemase from B. subtilis
Model of the mandelate racemase enzyme from Bacillus subtilis, a bacterium commonly found in soil.
New York Structural GenomiX Research Consortium, PSI
View Media
2530: Aspirin (with labels)
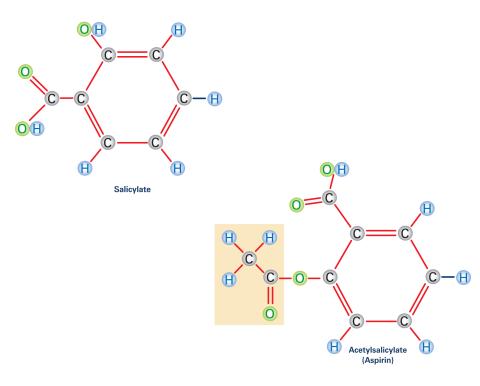
2530: Aspirin (with labels)
Acetylsalicylate (bottom) is the aspirin of today. Adding a chemical tag called an acetyl group (shaded box, bottom) to a molecule derived from willow bark (salicylate, top) makes the molecule less acidic (and easier on the lining of the digestive tract), but still effective at relieving pain. See image 2529 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
5875: Bacteriophage P22 capsid, detail
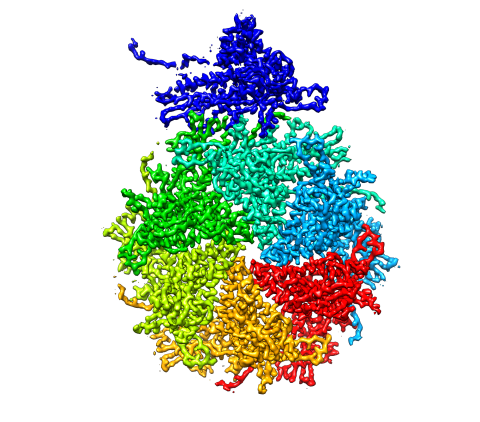
5875: Bacteriophage P22 capsid, detail
Detail of a subunit of the capsid, or outer cover, of bacteriophage P22, a virus that infects the Salmonella bacteria. Cryo-electron microscopy (cryo-EM) was used to capture details of the capsid proteins, each shown here in a separate color. Thousands of cryo-EM scans capture the structure and shape of all the individual proteins in the capsid and their position relative to other proteins. A computer model combines these scans into the image shown here. Related to image 5874.
Dr. Wah Chiu, Baylor College of Medicine
View Media
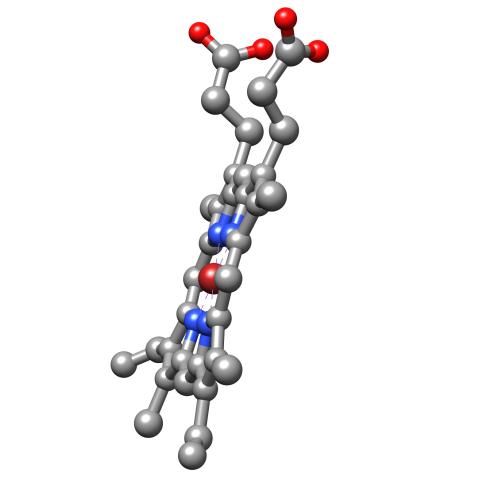
3540: Structure of heme, side view
3540: Structure of heme, side view
Molecular model of the struture of heme. Heme is a small, flat molecule with an iron ion (dark red) at its center. Heme is an essential component of hemoglobin, the protein in blood that carries oxygen throughout our bodies. This image first appeared in the September 2013 issue of Findings Magazine. View side view of heme here 3539.
Rachel Kramer Green, RCSB Protein Data Bank
View Media
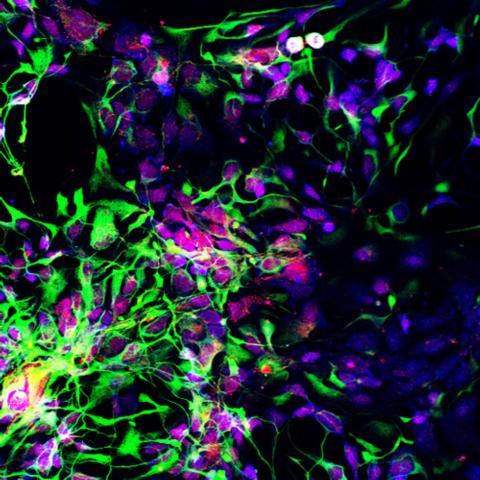
3280: Motor neuron progenitors derived from human ES cells
3280: Motor neuron progenitors derived from human ES cells
Motor neuron progenitors (green) were derived from human embryonic stem cells. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Hans Keirstead lab, University of California, Irvine, via CIRM
View Media
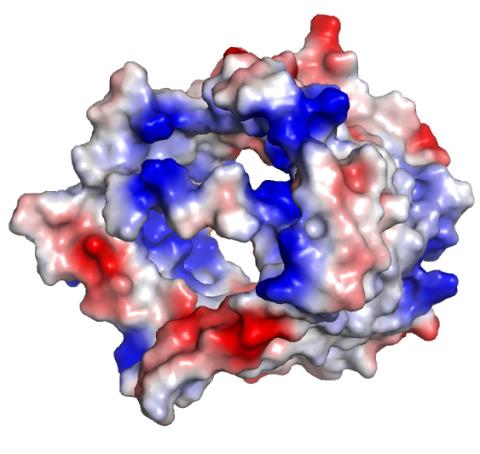
2494: VDAC-1 (3)
2494: VDAC-1 (3)
The structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to images 2491, 2495, and 2488.
Related to images 2491, 2495, and 2488.
Gerhard Wagner, Harvard Medical School
View Media
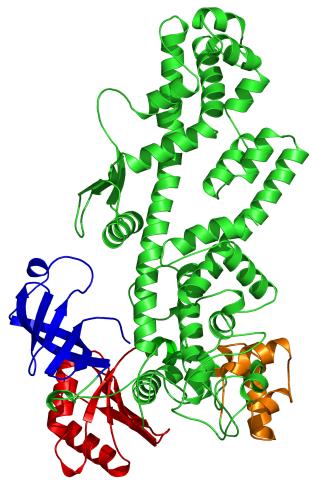
2338: Tex protein
2338: Tex protein
Model of a member from the Tex protein family, which is implicated in transcriptional regulation and highly conserved in eukaryotes and prokaryotes. The structure shows significant homology to a human transcription elongation factor that may regulate multiple steps in mRNA synthesis.
New York Structural GenomiX Research Consortium, PSI
View Media
3616: Weblike sheath covering developing egg chambers in a giant grasshopper
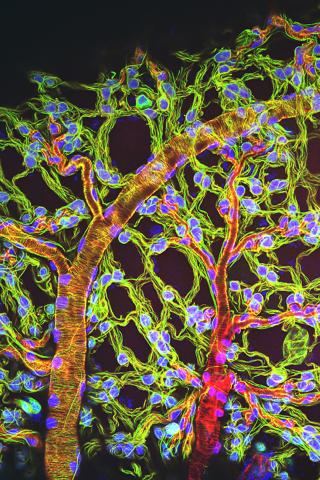
3616: Weblike sheath covering developing egg chambers in a giant grasshopper
The lubber grasshopper, found throughout the southern United States, is frequently used in biology classes to teach students about the respiratory system of insects. Unlike mammals, which have red blood cells that carry oxygen throughout the body, insects have breathing tubes that carry air through their exoskeleton directly to where it's needed. This image shows the breathing tubes embedded in the weblike sheath cells that cover developing egg chambers.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Kevin Edwards, Johny Shajahan, and Doug Whitman, Illinois State University.
View Media
2317: Fruitful dyes
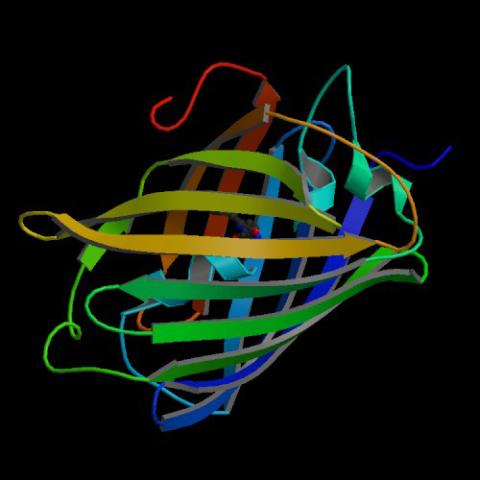
2317: Fruitful dyes
These colorful, computer-generated ribbons show the backbone of a molecule that glows a fluorescent red. The molecule, called mStrawberry, was created by chemists based on a protein found in the ruddy lips of a coral. Scientists use the synthetic molecule and other "fruity" ones like it as a dye to mark and study cell structures.
Roger Y. Tsien, University of California, San Diego
View Media
3451: Proteasome
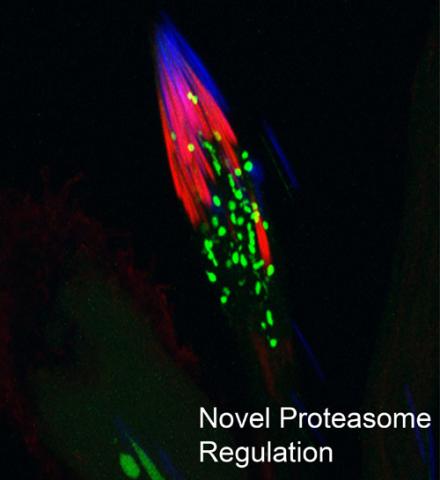
3451: Proteasome
This fruit fly spermatid recycles various molecules, including malformed or damaged proteins. Actin filaments (red) in the cell draw unwanted proteins toward a barrel-shaped structure called the proteasome (green clusters), which degrades the molecules into their basic parts for re-use.
Sigi Benjamin-Hong, Rockefeller University
View Media
1270: Glycoproteins
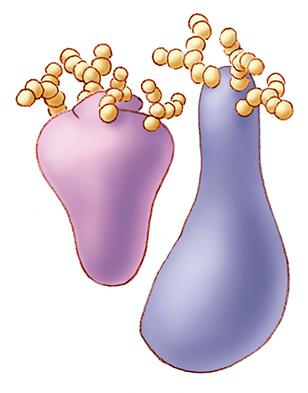
1270: Glycoproteins
About half of all human proteins include chains of sugar molecules that are critical for the proteins to function properly. Appears in the NIGMS booklet Inside the Cell.
Judith Stoffer
View Media
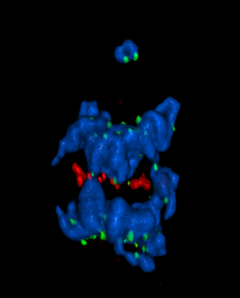
5766: A chromosome goes missing in anaphase
5766: A chromosome goes missing in anaphase
Anaphase is the critical step during mitosis when sister chromosomes are disjoined and directed to opposite spindle poles, ensuring equal distribution of the genome during cell division. In this image, one pair of sister chromosomes at the top was lost and failed to divide after chemical inhibition of polo-like kinase 1. This image depicts chromosomes (blue) separating away from the spindle mid-zone (red). Kinetochores (green) highlight impaired movement of some chromosomes away from the mid-zone or the failure of sister chromatid separation (top). Scientists are interested in detailing the signaling events that are disrupted to produce this effect. The image is a volume projection of multiple deconvolved z-planes acquired with a Nikon widefield fluorescence microscope.
This image was chosen as a winner of the 2016 NIH-funded research image call. The research that led to this image was funded by NIGMS.
Related to image 5765.
View Media
This image was chosen as a winner of the 2016 NIH-funded research image call. The research that led to this image was funded by NIGMS.
Related to image 5765.
