Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
1021: Lily mitosis 08
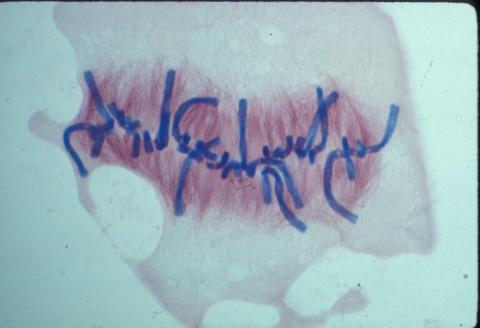
1021: Lily mitosis 08
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and lined up.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1018, and 1019.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1018, and 1019.
Andrew S. Bajer, University of Oregon, Eugene
View Media

2305: Beaded bacteriophage
2305: Beaded bacteriophage
This sculpture made of purple and clear glass beads depicts bacteriophage Phi174, a virus that infects bacteria. It rests on a surface that portrays an adaptive landscape, a conceptual visualization. The ridges represent the gene combinations associated with the greatest fitness levels of the virus, as measured by how quickly the virus can reproduce itself. Phi174 is an important model system for studies of viral evolution because its genome can readily be sequenced as it evolves under defined laboratory conditions.
Holly Wichman, University of Idaho. (Surface by A. Johnston; photo by J. Palmersheim)
View Media
2790: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 01
2790: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 01
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media

2409: Bacterial glucose isomerase
2409: Bacterial glucose isomerase
A crystal of bacterial glucose isomerase protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
2325: Multicolor STORM
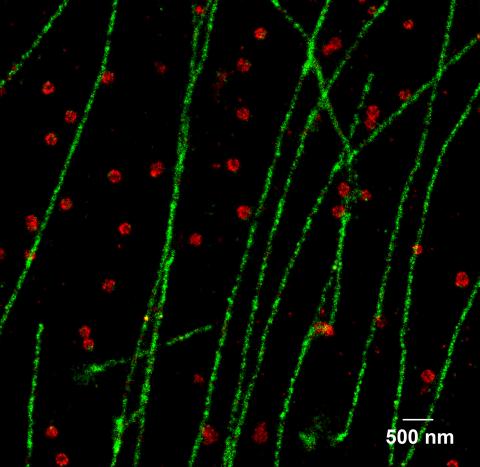
2325: Multicolor STORM
In 2006, scientists developed an optical microscopy technique enabling them to clearly see individual molecules within cells. In 2007, they took the technique, abbreviated STORM, a step further. They identified multicolored probes that let them peer into cells and clearly see multiple cellular components at the same time, such as these microtubules (green) and small hollows called clathrin-coated pits (red). Unlike conventional methods, the multicolor STORM technique produces a crisp and high resolution picture. A sharper view of how cellular components interact will likely help scientists answer some longstanding questions about cell biology.
Xiaowei Zhuang, Harvard University
View Media
3284: Neurons from human ES cells
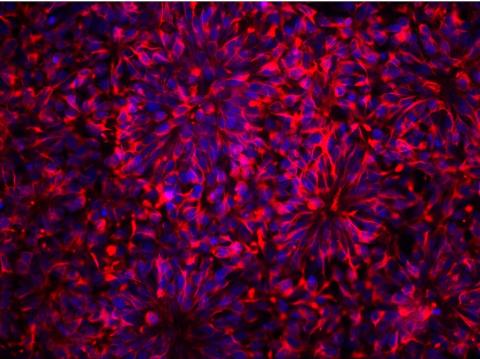
3284: Neurons from human ES cells
These neural precursor cells were derived from human embryonic stem cells. The neural cell bodies are stained red, and the nuclei are blue. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Xianmin Zeng lab, Buck Institute for Age Research, via CIRM
View Media
2365: Map of protein structures 01
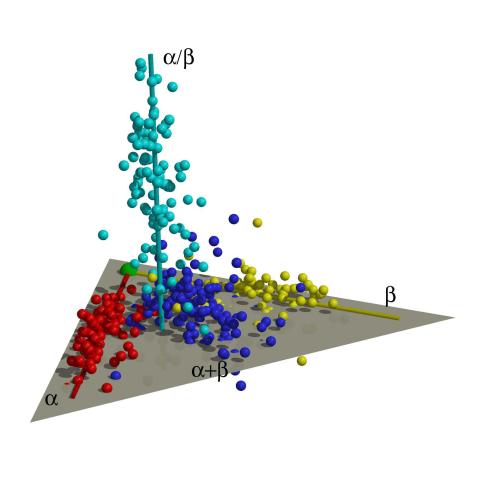
2365: Map of protein structures 01
A global "map of the protein structure universe." The Berkeley Structural Genomics Center has developed a method to visualize the vast universe of protein structures in which proteins of similar structure are located close together and those of different structures far away in the space. This map, constructed using about 500 of the most common protein folds, reveals a highly non-uniform distribution, and shows segregation between four elongated regions corresponding to four different protein classes (shown in four different colors). Such a representation reveals a high-level of organization of the protein structure universe.
Berkeley Structural Genomics Center, PSI
View Media
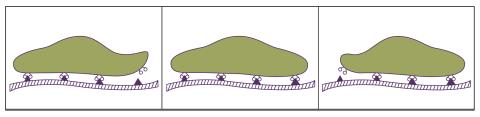
2502: Focal adhesions
2502: Focal adhesions
Cells walk along body surfaces via tiny "feet," called focal adhesions, that connect with the extracellular matrix. See image 2503 for a labeled version of this illustration.
Crabtree + Company
View Media
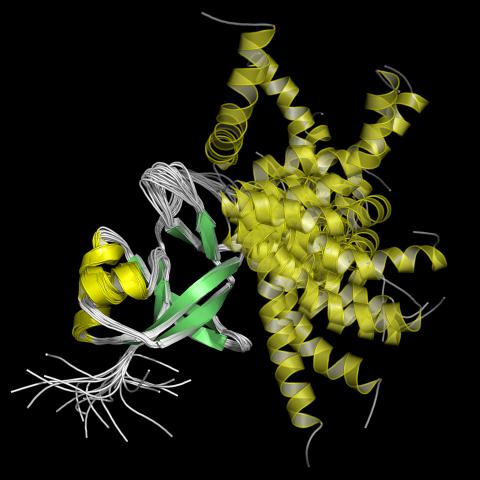
2339: Protein from Arabidopsis thaliana
2339: Protein from Arabidopsis thaliana
NMR solution structure of a plant protein that may function in host defense. This protein was expressed in a convenient and efficient wheat germ cell-free system. Featured as the June 2007 Protein Structure Initiative Structure of the Month.
Center for Eukaryotic Structural Genomics
View Media
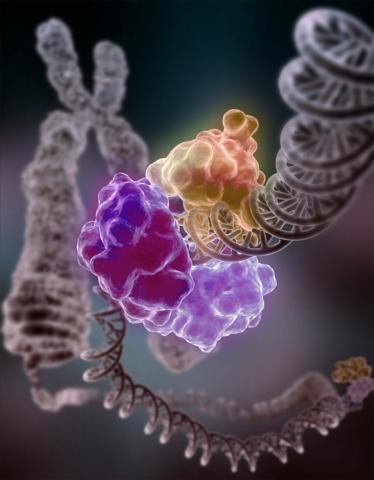
3493: Repairing DNA
3493: Repairing DNA
Like a watch wrapped around a wrist, a special enzyme encircles the double helix to repair a broken strand of DNA. Without molecules that can mend such breaks, cells can malfunction, die, or become cancerous. Related to image 2330.
Tom Ellenberger, Washington University School of Medicine
View Media
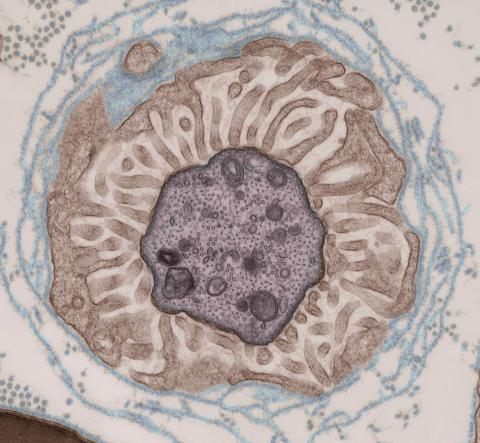
3740: Transmission electron microscopy showing cross-section of the node of Ranvier
3740: Transmission electron microscopy showing cross-section of the node of Ranvier
Nodes of Ranvier are short gaps in the myelin sheath surrounding myelinated nerve cells (axons). Myelin insulates axons, and the node of Ranvier is where the axon is exposed to the extracellular environment, allowing for the transmission of action potentials at these nodes via ion flows between the inside and outside of the axon. The image shows a cross-section through the node, with the surrounding extracellular matrix encasing and supporting the axon shown in cyan.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
3402: Hsp33 Heat Shock Protein Inactive to Active
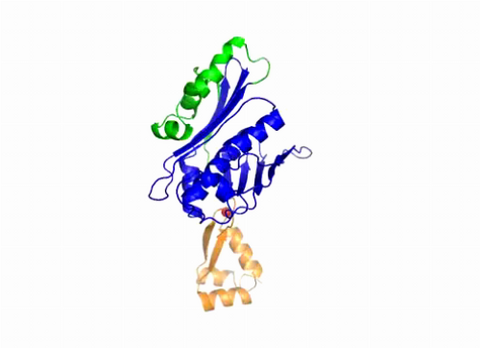
3402: Hsp33 Heat Shock Protein Inactive to Active
When the heat shock protein hsp33 is folded, it is inactive and contains a zinc ion, stabilizing the redox sensitive domain (orange). In the presence of an environmental stressor, the protein releases the zinc ion, which leads to the unfolding of the redox domain. This unfolding causes the chaperone to activate by reaching out its "arm" (green) to protect other proteins.
Dana Reichmann, University of Michigan
View Media
6991: SARS-CoV-2 nucleocapsid dimer
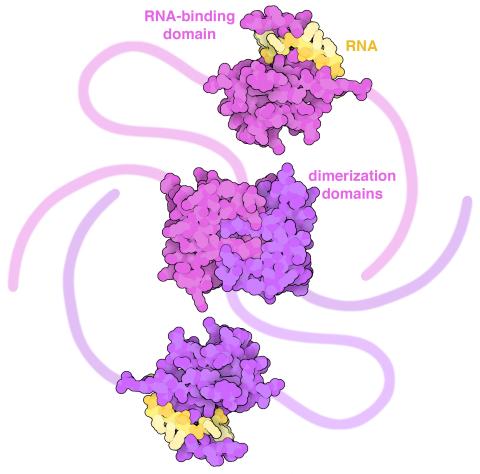
6991: SARS-CoV-2 nucleocapsid dimer
In SARS-CoV-2, the virus that causes COVID-19, nucleocapsid is a complex molecule with many functional parts. One section folds into an RNA-binding domain, with a groove that grips a short segment of the viral genomic RNA. Another section folds into a dimerization domain that brings two nucleocapsid molecules together. The rest of the protein is intrinsically disordered, forming tails at each end of the protein chain and a flexible linker that connects the two structured domains. These disordered regions assist with RNA binding and orchestrate association of nucleocapsid dimers into larger assemblies that package the RNA in the small space inside virions. Nucleocapsid is in magenta and purple, and short RNA strands are in yellow.
Find these in the RCSB Protein Data Bank: RNA-binding domain (PDB entry 7ACT) and Dimerization domain (PDB entry 6WJI).
Find these in the RCSB Protein Data Bank: RNA-binding domain (PDB entry 7ACT) and Dimerization domain (PDB entry 6WJI).
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
2507: Carbon building blocks (with examples)
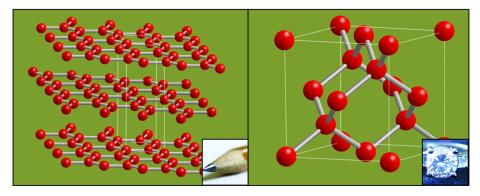
2507: Carbon building blocks (with examples)
The arrangement of identical molecular components can make a dramatic difference. For example, carbon atoms can be arranged into dull graphite (left) or sparkly diamonds (right). See image 2506 for an illustration without examples.
Crabtree + Company
View Media
3646: Cells lining the trachea
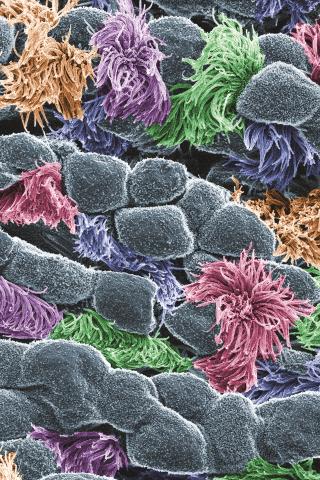
3646: Cells lining the trachea
In this image, viewed with a ZEISS ORION NanoFab microscope, the community of cells lining a mouse airway is magnified more than 10,000 times. This collection of cells, known as the mucociliary escalator, is also found in humans. It is our first line of defense against inhaled bacteria, allergens, pollutants, and debris. Malfunctions in the system can cause or aggravate lung infections and conditions such as asthma and chronic obstructive pulmonary disease. The cells shown in gray secrete mucus, which traps inhaled particles. The colored cells sweep the mucus layer out of the lungs.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Eva Mutunga and Kate Klein, University of the District of Columbia and National Institute of Standards and Technology
View Media

3728: Quorum-sensing inhibitor limits bacterial growth
3728: Quorum-sensing inhibitor limits bacterial growth
To simulate the consequences of disrupting bacterial cell-to-cell communication, called quorum sensing, in the crypts (small chambers within the colon), the researchers experimented with an inhibitor molecule (i.e., antagonist) to turn off quorum sensing in methicillin-resistant Staphylococcus aureus (MRSA), an antibiotic-resistant strain of bacteria that often causes human infections. In this experiment, a medium promoting bacterial growth flows through experimental chambers mimicking the colon environment. The chambers on the right contained no antagonist. In the left chambers, after being added to the flowing medium, the quorum-sensing-inhibiting molecules quickly spread throughout the crevices, inactivating quorum sensing and reducing colonization. These results suggest a potential strategy for addressing MRSA virulence via inhibitors of bacterial communication. You can read more about this research here.
Minyoung Kevin Kim and Bonnie Bassler, Princeton University
View Media
2626: Telomeres
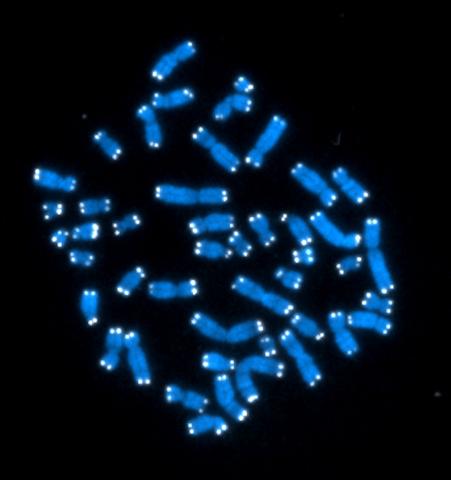
2626: Telomeres
The 46 human chromosomes are shown in blue, with the telomeres appearing as white pinpoints. The DNA has already been copied, so each chromosome is actually made up of two identical lengths of DNA, each with its own two telomeres.
Hesed Padilla-Nash and Thomas Ried, the National Cancer Institute, a part of NIH
View Media
7013: An adult Hawaiian bobtail squid
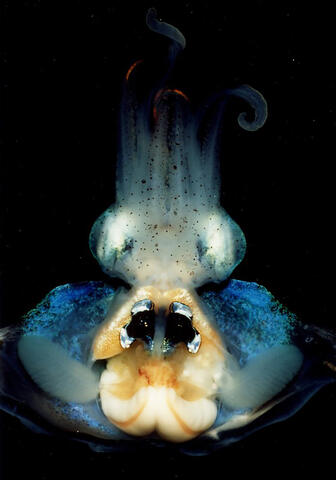
7013: An adult Hawaiian bobtail squid
An adult female Hawaiian bobtail squid, Euprymna scolopes, with its mantle cavity exposed from the underside. Some internal organs are visible, including the two lobes of the light organ that contains bioluminescent bacteria, Vibrio fischeri. The light organ includes accessory tissues like an ink sac (black) that serves as a shutter, and a silvery reflector that directs the light out of the underside of the animal.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
6603: Protein formation
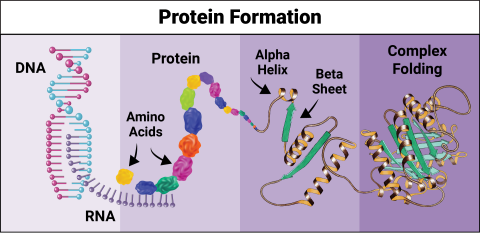
6603: Protein formation
Proteins are 3D structures made up of smaller units. DNA is transcribed to RNA, which in turn is translated into amino acids. Amino acids form a protein strand, which has sections of corkscrew-like coils, called alpha helices, and other sections that fold flat, called beta sheets. The protein then goes through complex folding to produce the 3D structure.
NIGMS, with the folded protein illustration adapted from Jane Richardson, Duke University Medical Center
View Media

2385: Heat shock protein complex from Methanococcus jannaschii
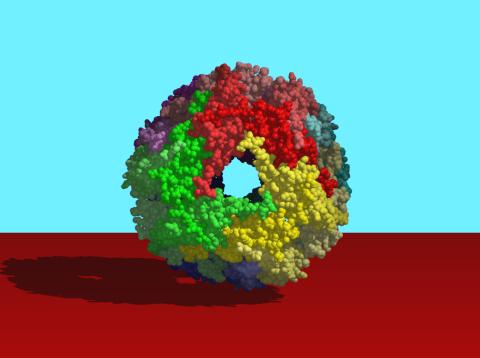
2385: Heat shock protein complex from Methanococcus jannaschii
Model based on X-ray crystallography of the structure of a small heat shock protein complex from the bacteria, Methanococcus jannaschii. Methanococcus jannaschii is an organism that lives at near boiling temperature, and this protein complex helps it cope with the stress of high temperature. Similar complexes are produced in human cells when they are "stressed" by events such as burns, heart attacks, or strokes. The complexes help cells recover from the stressful event.
Berkeley Structural Genomics Center, PSI-1
View Media

2708: Leading cells with light
2708: Leading cells with light
A blue laser beam turns on a protein that helps this human cancer cell move. Responding to the stimulus, the protein, called Rac1, first creates ruffles at the edge of the cell. Then it stretches the cell forward, following the light like a horse trotting after a carrot on a stick. This new light-based approach can turn Rac1 (and potentially many other proteins) on and off at exact times and places in living cells. By manipulating a protein that controls movement, the technique also offers a new tool to study embryonic development, nerve regeneration and cancer.
Yi Wu, University of North Carolina
View Media
1058: Lily mitosis 01
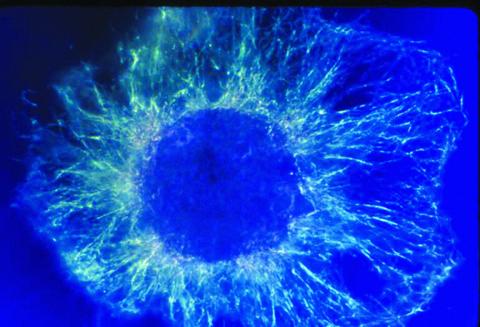
1058: Lily mitosis 01
A light microscope image shows the chromosomes, stained dark blue, in a dividing cell of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones.
Andrew S. Bajer, University of Oregon, Eugene
View Media
2574: Simulation of uncontrolled avian flu outbreak
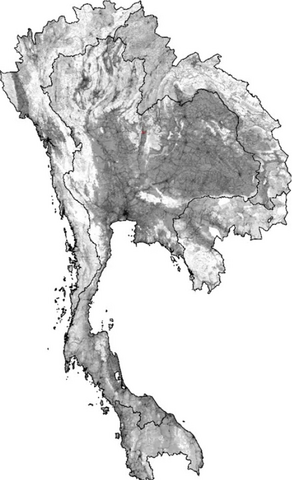
2574: Simulation of uncontrolled avian flu outbreak
This video simulation shows what an uncontrolled outbreak of transmissible avian flu among people living in Thailand might look like. Red indicates new cases while green indicates areas where the epidemic has finished. The video shows the spread of infection and recovery over 300 days in Thailand and neighboring countries.
Neil M. Ferguson, Imperial College London
View Media
2739: Tetrapolar mitosis
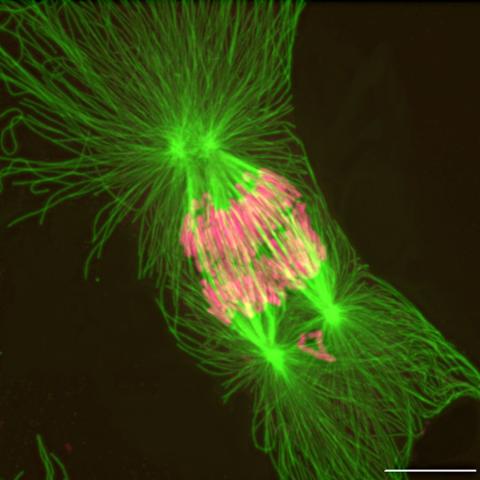
2739: Tetrapolar mitosis
This image shows an abnormal, tetrapolar mitosis. Chromosomes are highlighted pink. The cells shown are S3 tissue cultured cells from Xenopus laevis, African clawed frog.
Gary Gorbsky, Oklahoma Medical Research Foundation
View Media
2331: Statistical cartography
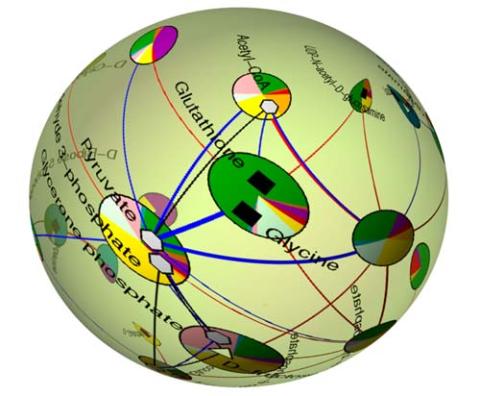
2331: Statistical cartography
Like a world of its own, this sphere represents all the known chemical reactions in the E. coli bacterium. The colorful circles on the surface symbolize sets of densely interconnected reactions. The lines between the circles show additional connecting reactions. The shapes inside the circles are landmark molecules, like capital cities on a map, that either act as hubs for many groups of reactions, are highly conserved among species, or both. Molecules that connect far-flung reactions on the sphere are much more conserved during evolution than molecules that connect reactions within a single circle. This statistical cartography could reveal insights about other complex systems, such as protein interactions and gene regulation networks.
Luis A. Nunes Amaral, Northwestern University
View Media

1313: Cell eyes clock
2364: High-throughput protein structure determination pipeline
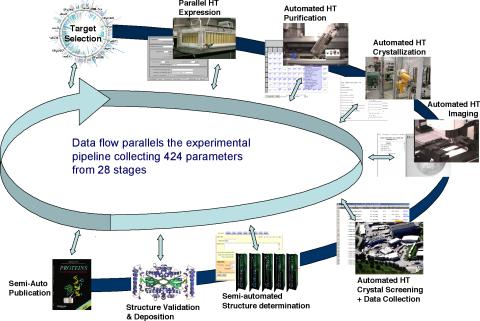
2364: High-throughput protein structure determination pipeline
This slide shows the technologies that the Joint Center for Structural Genomics developed for going from gene to structure and how the technologies have been integrated into a high-throughput pipeline, including all of the steps from target selection, parallel expression, protein purification, automated crystallization trials, automated crystal screening, structure determination, validation, and publication.
Joint Center for Structural Genomics
View Media
3746: Serum albumin structure 3
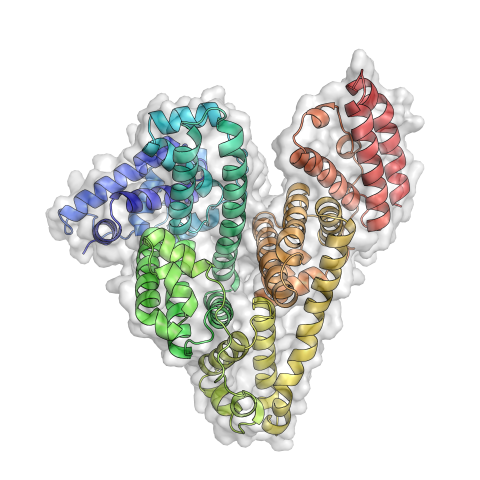
3746: Serum albumin structure 3
Serum albumin (SA) is the most abundant protein in the blood plasma of mammals. SA has a characteristic heart-shape structure and is a highly versatile protein. It helps maintain normal water levels in our tissues and carries almost half of all calcium ions in human blood. SA also transports some hormones, nutrients and metals throughout the bloodstream. Despite being very similar to our own SA, those from other animals can cause some mild allergies in people. Therefore, some scientists study SAs from humans and other mammals to learn more about what subtle structural or other differences cause immune responses in the body.
Related to entries 3744 and 3745.
Related to entries 3744 and 3745.
Wladek Minor, University of Virginia
View Media
3539: Structure of heme, top view
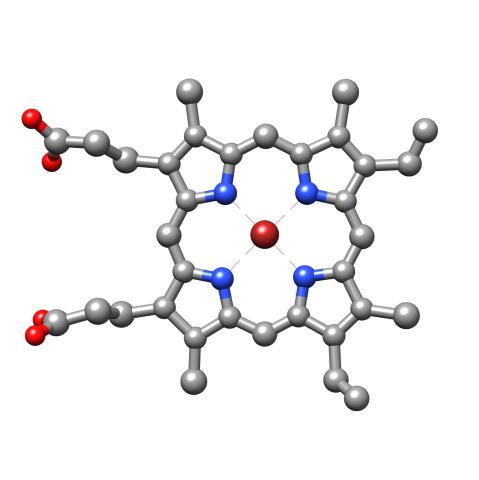
3539: Structure of heme, top view
Molecular model of the struture of heme. Heme is a small, flat molecule with an iron ion (dark red) at its center. Heme is an essential component of hemoglobin, the protein in blood that carries oxygen throughout our bodies. This image first appeared in the September 2013 issue of Findings Magazine. View side view of heme here 3540.
Rachel Kramer Green, RCSB Protein Data Bank
View Media
6848: Himastatin
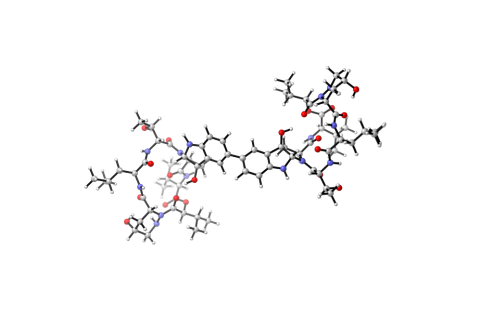
6848: Himastatin
A model of the molecule himastatin, which was first isolated from the bacterium Streptomyces himastatinicus. Himastatin shows antibiotic activity. The researchers who created this image developed a new, more concise way to synthesize himastatin so it can be studied more easily.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6850 and video 6851.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6850 and video 6851.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
3481: Bacillus anthracis being killed
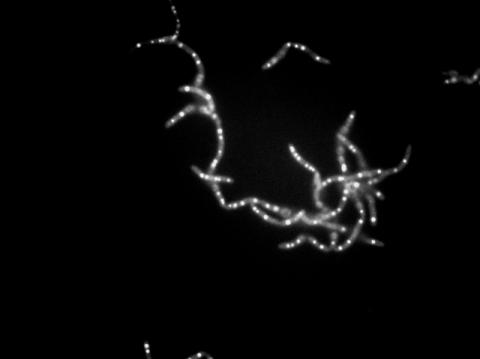
3481: Bacillus anthracis being killed
Bacillus anthracis (anthrax) cells being killed by a fluorescent trans-translation inhibitor, which disrupts bacterial protein synthesis. The inhibitor is naturally fluorescent and looks blue when it is excited by ultraviolet light in the microscope. This is a black-and-white version of Image 3525.
John Alumasa, Keiler Laboratory, Pennsylvania State University
View Media
2605: Induced stem cells from adult skin 03
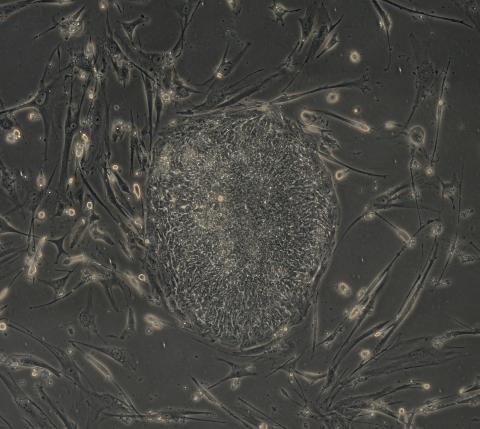
2605: Induced stem cells from adult skin 03
The human skin cells pictured contain genetic modifications that make them pluripotent, essentially equivalent to embryonic stem cells. A scientific team from the University of Wisconsin-Madison including researchers Junying Yu, James Thomson, and their colleagues produced the transformation by introducing a set of four genes into human fibroblasts, skin cells that are easy to obtain and grow in culture.
James Thomson, University of Wisconsin-Madison
View Media
2494: VDAC-1 (3)
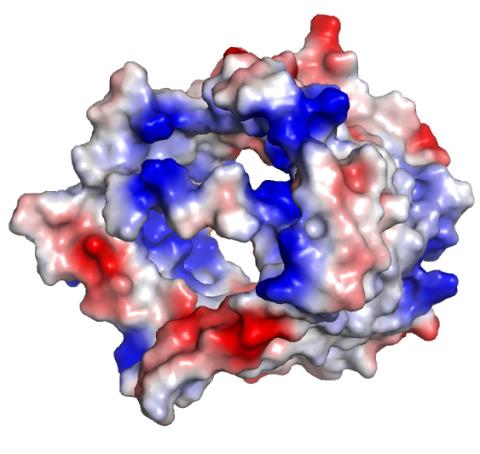
2494: VDAC-1 (3)
The structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to images 2491, 2495, and 2488.
Related to images 2491, 2495, and 2488.
Gerhard Wagner, Harvard Medical School
View Media
3483: Chang Shan
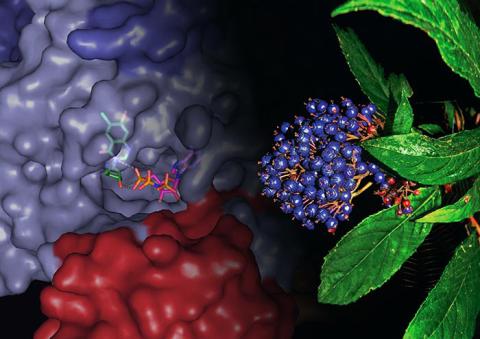
3483: Chang Shan
For thousands of years, Chinese herbalists have treated malaria using Chang Shan, a root extract from a type of hydrangea that grows in Tibet and Nepal. Recent studies have suggested Chang Shan can also reduce scar formation, treat multiple sclerosis and even slow cancer progression.
Paul Schimmel Lab, Scripps Research Institute
View Media
2782: Disease-susceptible Arabidopsis leaf
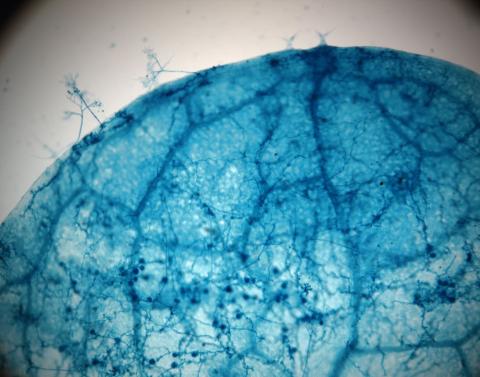
2782: Disease-susceptible Arabidopsis leaf
This is a magnified view of an Arabidopsis thaliana leaf after several days of infection with the pathogen Hyaloperonospora arabidopsidis. The pathogen's blue hyphae grow throughout the leaf. On the leaf's edges, stalk-like structures called sporangiophores are beginning to mature and will release the pathogen's spores. Inside the leaf, the large, deep blue spots are structures called oopsorangia, also full of spores. Compare this response to that shown in Image 2781. Jeff Dangl has been funded by NIGMS to study the interactions between pathogens and hosts that allow or suppress infection.
Jeff Dangl, University of North Carolina, Chapel Hill
View Media

1334: Aging book of life
1334: Aging book of life
Damage to each person's genome, often called the "Book of Life," accumulates with time. Such DNA mutations arise from errors in the DNA copying process, as well as from external sources, such as sunlight and cigarette smoke. DNA mutations are known to cause cancer and also may contribute to cellular aging.
Judith Stoffer
View Media
2432: ARTS triggers apoptosis
2432: ARTS triggers apoptosis
Cell showing overproduction of the ARTS protein (red). ARTS triggers apoptosis, as shown by the activation of caspase-3 (green) a key tool in the cell's destruction. The nucleus is shown in blue. Image is featured in October 2015 Biomedical Beat blog post Cool Images: A Halloween-Inspired Cell Collection.
Hermann Steller, Rockefeller University
View Media

2543: DNA replication illustration
2543: DNA replication illustration
During DNA replication, each strand of the original molecule acts as a template for the synthesis of a new, complementary DNA strand. See image 2544 for a labeled version of this illustration.
Crabtree + Company
View Media
