Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
6993: RNA polymerase
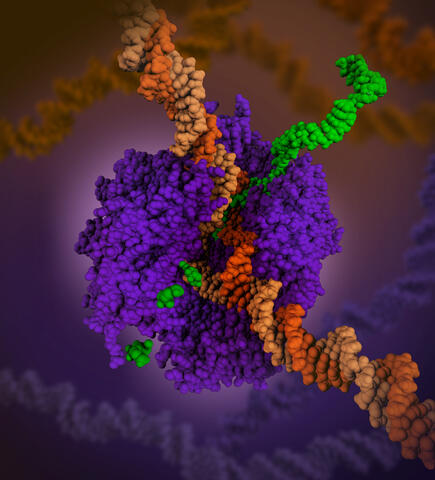
6993: RNA polymerase
RNA polymerase (purple) is a complex enzyme at the heart of transcription. During this process, the enzyme unwinds the DNA double helix and uses one strand (darker orange) as a template to create the single-stranded messenger RNA (green), later used by ribosomes for protein synthesis.
From the RNA polymerase II elongation complex of Saccharomyces cerevisiae (PDB entry 1I6H) as seen in PDB-101's What is a Protein? video.
From the RNA polymerase II elongation complex of Saccharomyces cerevisiae (PDB entry 1I6H) as seen in PDB-101's What is a Protein? video.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
2328: Neural tube development
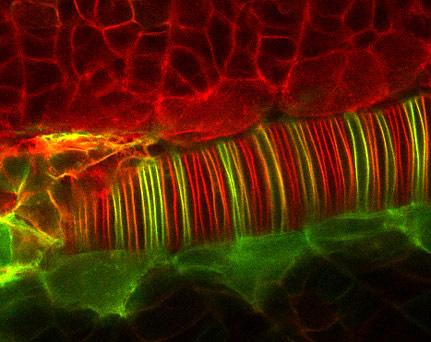
2328: Neural tube development
Proteins in the neural tissues of this zebrafish embryo direct cells to line up and form the neural tube, which will become the spinal cord and brain. Studies of zebrafish embryonic development may help pinpoint the underlying cause of common neural tube defects--such as spina bifida--which occur in about 1 in 1,000 newborn children.
Alexander Schier, Harvard University
View Media
2352: Human aspartoacylase
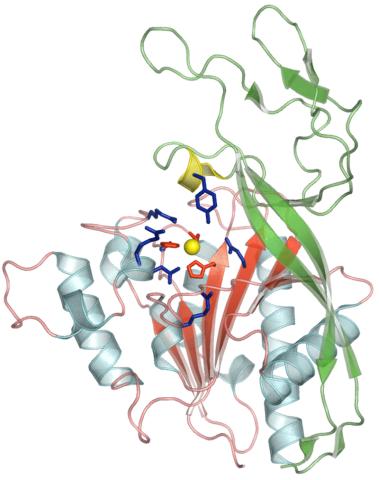
2352: Human aspartoacylase
Model of aspartoacylase, a human enzyme involved in brain metabolism.
Center for Eukaryotic Structural Genomics, PSI
View Media
3647: Epithelial cells
3647: Epithelial cells
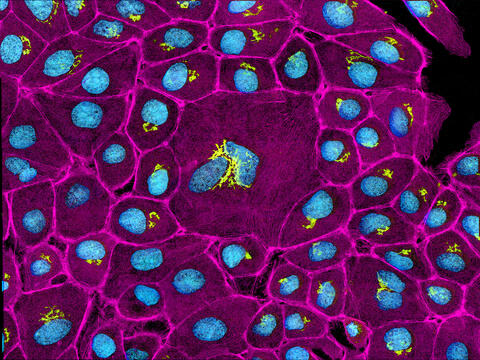
This image mostly shows normal cultured epithelial cells expressing green fluorescent protein targeted to the Golgi apparatus (yellow-green) and stained for actin (magenta) and DNA (cyan). The middle cell is an abnormal large multinucleated cell. All the cells in this image have a Golgi but not all are expressing the targeted recombinant fluorescent protein.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
6927: Axolotl showing nervous system
6927: Axolotl showing nervous system
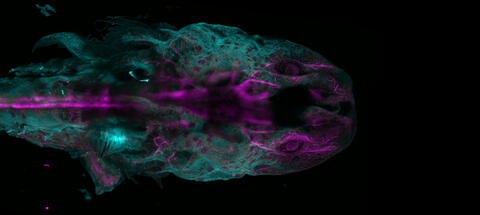
The head of an axolotl—a type of salamander—that has been genetically modified so that its developing nervous system glows purple and its Schwann cell nuclei appear light blue. Schwann cells insulate and provide nutrients to peripheral nerve cells. Researchers often study axolotls for their extensive regenerative abilities. They can regrow tails, limbs, spinal cords, brains, and more. The researcher who took this image focuses on the role of the peripheral nervous system during limb regeneration.
This image was captured using a light sheet microscope.
Related to images 6928 and 6932.
This image was captured using a light sheet microscope.
Related to images 6928 and 6932.
Prayag Murawala, MDI Biological Laboratory and Hannover Medical School.
View Media
3290: Three neurons and human ES cells
3290: Three neurons and human ES cells
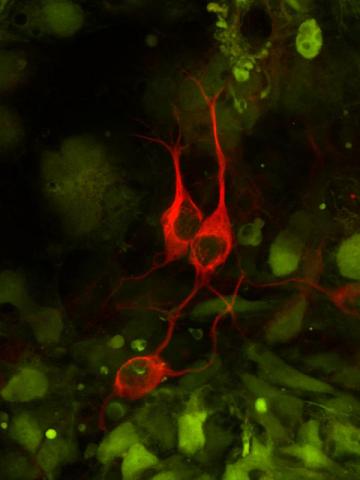
The three neurons (red) visible in this image were derived from human embryonic stem cells. Undifferentiated stem cells are green here. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Anirvan Ghosh lab, University of California, San Diego, via CIRM
View Media
5751: Genetically identical mycobacteria respond differently to antibiotic 1
5751: Genetically identical mycobacteria respond differently to antibiotic 1
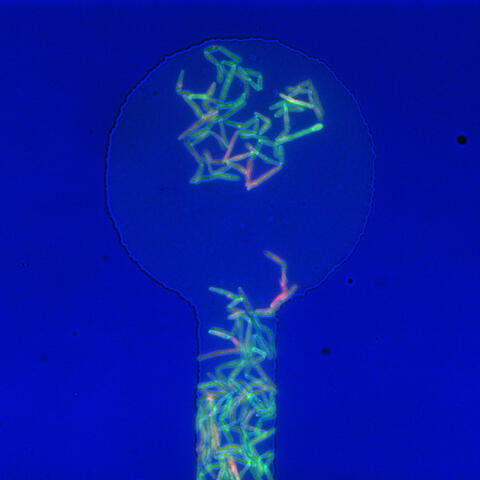
Antibiotic resistance in microbes is a serious health concern. So researchers have turned their attention to how bacteria undo the action of some antibiotics. Here, scientists set out to find the conditions that help individual bacterial cells survive in the presence of the antibiotic rifampicin. The research team used Mycobacterium smegmatis, a more harmless relative of Mycobacterium tuberculosis, which infects the lung and other organs and causes serious disease.
In this image, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to video 5752.
In this image, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to video 5752.
Bree Aldridge, Tufts University
View Media
1018: Lily mitosis 12
1018: Lily mitosis 12
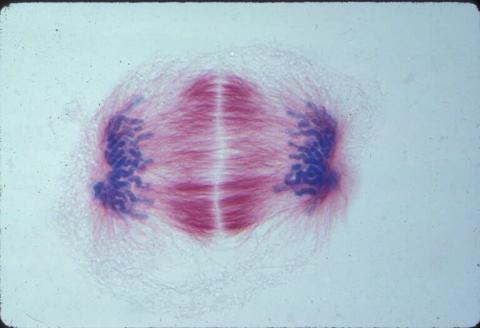
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible near the end of a round of mitosis.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1019, and 1021.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media

3772: The Proteasome: The Cell's Trash Processor in Action
3772: The Proteasome: The Cell's Trash Processor in Action
Our cells are constantly removing and recycling molecular waste. This video shows one way cells process their trash.
View Media

5810: Tongue 1
5810: Tongue 1
Microscopy image of tongue. One in a series of two, see image 5811
National Center for Microscopy and Imaging Research (NCMIR)
View Media
3314: Human opioid receptor structure superimposed on poppy
3314: Human opioid receptor structure superimposed on poppy
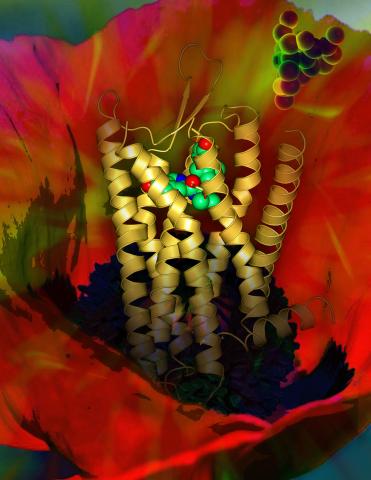
Opioid receptors on the surfaces of brain cells are involved in pleasure, pain, addiction, depression, psychosis, and other conditions. The receptors bind to both innate opioids and drugs ranging from hospital anesthetics to opium. Researchers at The Scripps Research Institute, supported by the NIGMS Protein Structure Initiative, determined the first three-dimensional structure of a human opioid receptor, a kappa-opioid receptor. In this illustration, the submicroscopic receptor structure is shown while bound to an agonist (or activator). The structure is superimposed on a poppy flower, the source of opium.
Raymond Stevens, The Scripps Research Institute
View Media
3365: Chemokine CXCR4 receptor
3365: Chemokine CXCR4 receptor
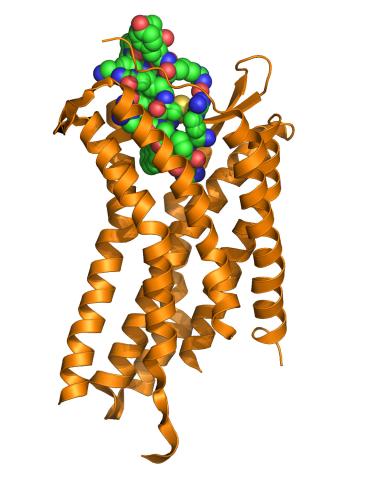
The receptor is shown bound to a small molecule peptide called CVX15.
Raymond Stevens, The Scripps Research Institute
View Media

2715: Glow-in-the-dark salamanders
2715: Glow-in-the-dark salamanders
These six-month-old axolotls, a kind of salamander, glow green and blue under ultraviolet light. That's because they were genetically modified to make harmless green fluorescent protein, or GFP. Like X-ray vision, GFP lets you see inside the axolotls as they hang out in their aquarium. GFP not only can reveal internal structures in living organisms, but it also can light up specific cells and even proteins within a cell. That allows scientists to identify and track things like cancer cells.
View Media
2532: Drugs enter skin (with labels)
2532: Drugs enter skin (with labels)
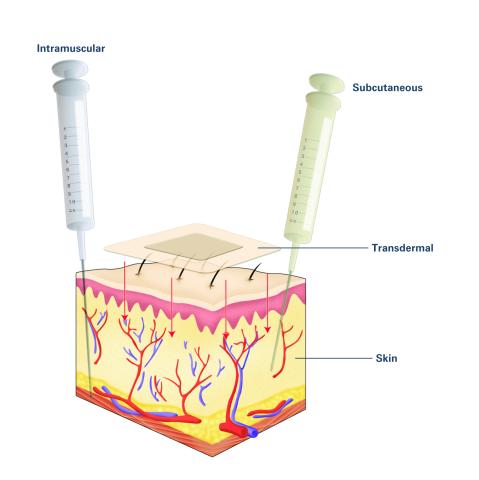
Drugs enter different layers of skin via intramuscular, subcutaneous, or transdermal delivery methods. See image 2531 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
3791: Nucleolus subcompartments spontaneously self-assemble 2
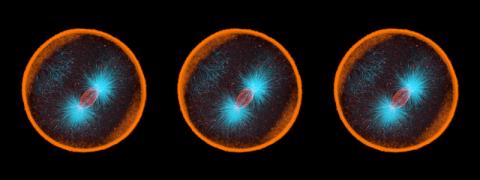
3791: Nucleolus subcompartments spontaneously self-assemble 2
The nucleolus is a small but very important protein complex located in the cell's nucleus. It forms on the chromosomes at the location where the genes for the RNAs are that make up the structure of the ribosome, the indispensable cellular machine that makes proteins from messenger RNAs.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This video of nucleoli in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows how each of the compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue) spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, image 3792 and image 3793.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This video of nucleoli in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows how each of the compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue) spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, image 3792 and image 3793.
Nilesh Vaidya, Princeton University
View Media
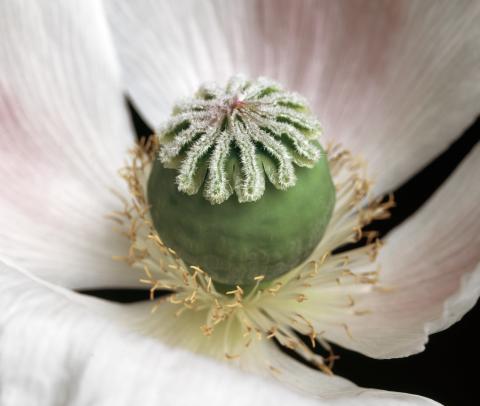

3423: White Poppy (cropped)
3423: White Poppy (cropped)
A cropped image of a white poppy. View poppy uncropped here 3424.
Judy Coyle, Donald Danforth Plant Science Center
View Media
2309: Cellular polarity
2309: Cellular polarity
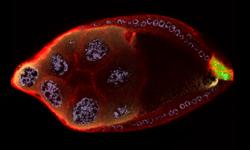
As an egg cell develops, a process called polarization controls what parts ultimately become the embryo's head and tail. This picture shows an egg of the fruit fly Drosophila. Red and green mark two types of signaling proteins involved in polarization. Disrupting these signals can scramble the body plan of the embryo, leading to severe developmental disorders.
Wu-Min Deng, Florida State University
View Media
6984: Fruit fly starvation leads to adipokine accumulation
6984: Fruit fly starvation leads to adipokine accumulation
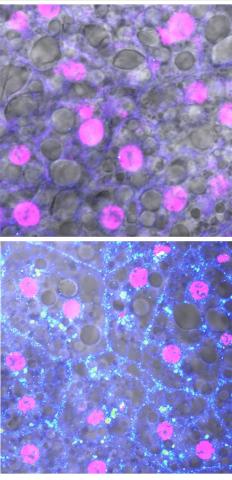
Adult Drosophila abdominal fat tissue showing cell nuclei labelled in magenta. The upper panel is from well-fed flies, and the lower panel is from flies that have been deprived of food for 4 hours. Starvation results in the accumulation of a key adipokine—a fat hormone (blue-green dots).
Related to images 6982, 6983, and 6985.
Related to images 6982, 6983, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media
6805: Staphylococcus aureus aggregating upon contact with synovial fluid
6805: Staphylococcus aureus aggregating upon contact with synovial fluid
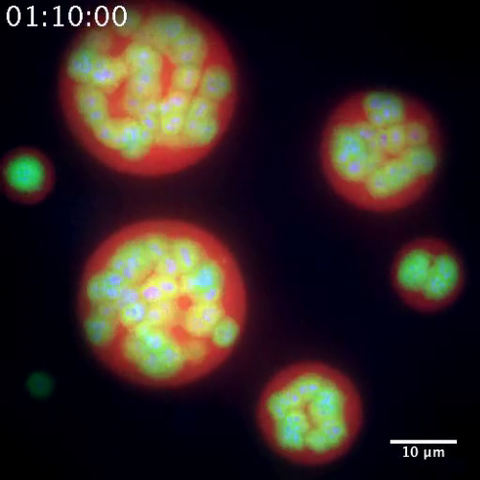
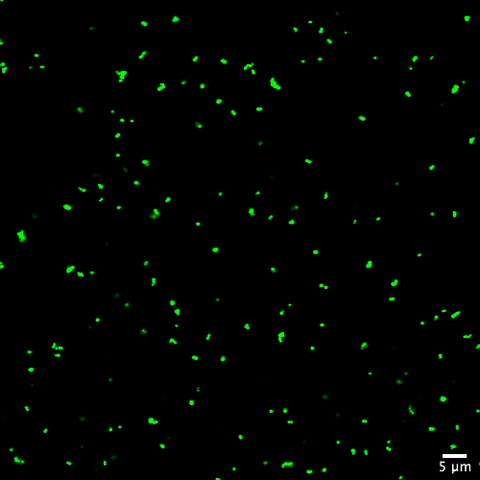
Staphylococcus aureus bacteria (green) grouping together upon contact with synovial fluid—a viscous substance found in joints. The formation of groups can help protect the bacteria from immune system defenses and from antibiotics, increasing the likelihood of an infection. This video is a 1-hour time lapse and was captured using a confocal laser scanning microscope.
More information about the research that produced this video can be found in the Journal of Bacteriology paper "In Vitro Staphylococcal Aggregate Morphology and Protection from Antibiotics Are Dependent on Distinct Mechanisms Arising from Postsurgical Joint Components and Fluid Motion" by Staats et al.
Related to images 6803 and 6804.
More information about the research that produced this video can be found in the Journal of Bacteriology paper "In Vitro Staphylococcal Aggregate Morphology and Protection from Antibiotics Are Dependent on Distinct Mechanisms Arising from Postsurgical Joint Components and Fluid Motion" by Staats et al.
Related to images 6803 and 6804.
Paul Stoodley, The Ohio State University.
View Media
1158: Bacteria shapes
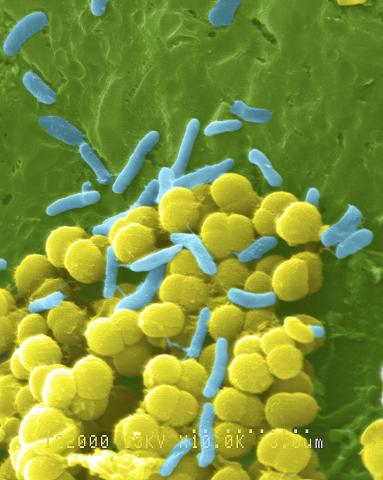
1158: Bacteria shapes
A colorized scanning electron micrograph of bacteria. Scanning electron microscopes allow scientists to see the three-dimensional surface of their samples.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
1016: Lily mitosis 06
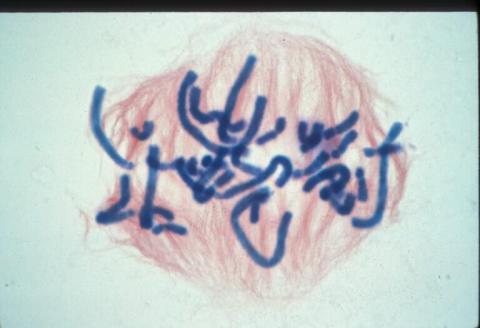
1016: Lily mitosis 06
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and are starting to line up.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1017, 1018, 1019, and 1021.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1017, 1018, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
3573: Myotonic dystrophy type 2 genetic defect
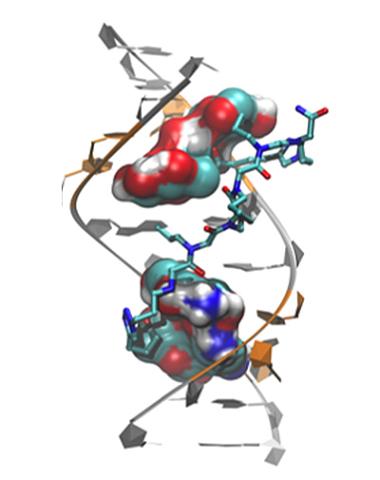
3573: Myotonic dystrophy type 2 genetic defect
Scientists revealed a detailed image of the genetic defect that causes myotonic dystrophy type 2 and used that information to design drug candidates to counteract the disease.
Matthew Disney, Scripps Research Institute and Ilyas Yildirim, Northwestern University
View Media
3273: Heart muscle with reprogrammed skin cells
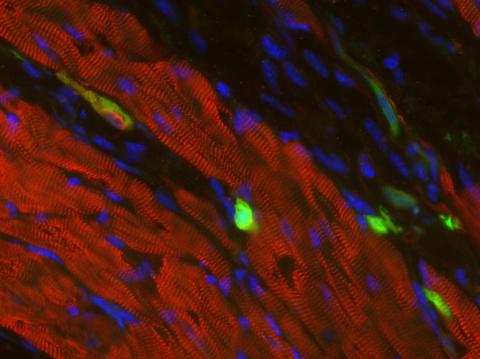
3273: Heart muscle with reprogrammed skin cells
Skins cells were reprogrammed into heart muscle cells. The cells highlighted in green are remaining skin cells. Red indicates a protein that is unique to heart muscle. The technique used to reprogram the skin cells into heart cells could one day be used to mend heart muscle damaged by disease or heart attack. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Deepak Srivastava, Gladstone Institute of Cardiovascular Disease, via CIRM
View Media
1284: Ion channels
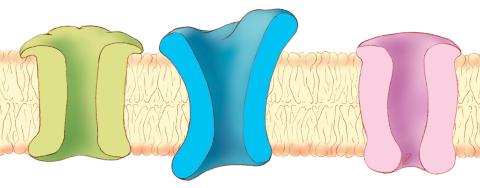
1284: Ion channels
The body uses a variety of ion channels to transport small molecules across cell membranes.
Judith Stoffer
View Media
1191: Mouse sperm sections
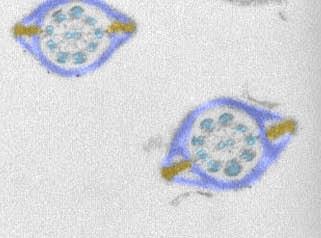
1191: Mouse sperm sections
This transmission electron micrograph shows sections of mouse sperm tails, or flagella.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
2505: Influenza virus attaches to host membrane (with labels)
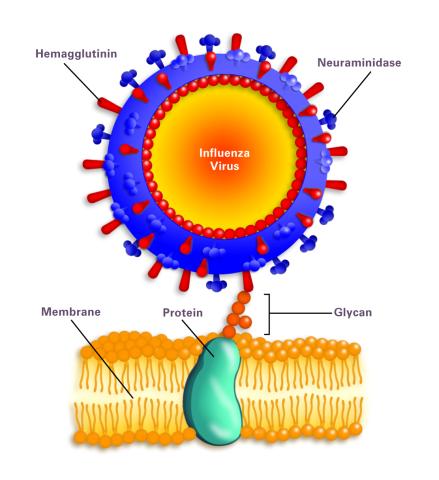
2505: Influenza virus attaches to host membrane (with labels)
Influenza A infects a host cell when hemagglutinin grips onto glycans on its surface. Neuraminidase, an enzyme that chews sugars, helps newly made virus particles detach so they can infect other cells. Related to 213.
Crabtree + Company
View Media

2412: Pig alpha amylase
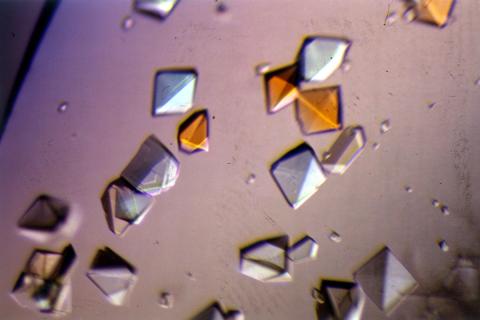
2412: Pig alpha amylase
Crystals of porcine alpha amylase protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
6982: Insulin production and fat sensing in fruit flies
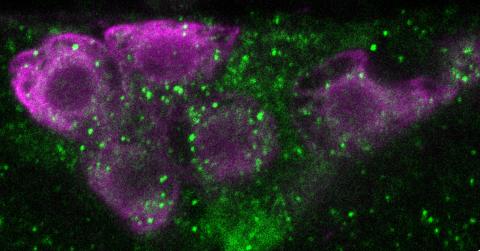
6982: Insulin production and fat sensing in fruit flies
Fourteen neurons (magenta) in the adult Drosophila brain produce insulin, and fat tissue sends packets of lipids to the brain via the lipoprotein carriers (green). This image was captured using a confocal microscope and shows a maximum intensity projection of many slices.
Related to images 6983, 6984, and 6985.
Related to images 6983, 6984, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media
3792: Nucleolus subcompartments spontaneously self-assemble 3
3792: Nucleolus subcompartments spontaneously self-assemble 3
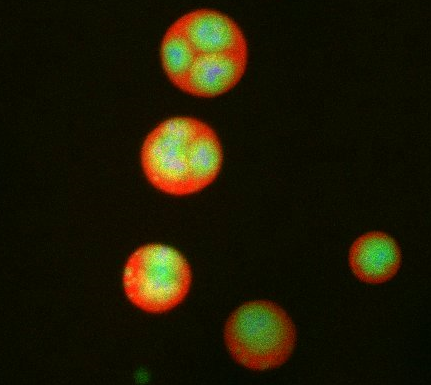
What looks a little like distant planets with some mysterious surface features are actually assemblies of proteins normally found in the cell's nucleolus, a small but very important protein complex located in the cell's nucleus. It forms on the chromosomes at the location where the genes for the RNAs are that make up the structure of the ribosome, the indispensable cellular machine that makes proteins from messenger RNAs.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This photo of nucleolus proteins in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows each of the nucleolus compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue). The researchers have found that these compartments spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, video 3791 and image 3793.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This photo of nucleolus proteins in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows each of the nucleolus compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue). The researchers have found that these compartments spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, video 3791 and image 3793.
Nilesh Vaidya, Princeton University
View Media
6928: Axolotls showing nervous system components
6928: Axolotls showing nervous system components
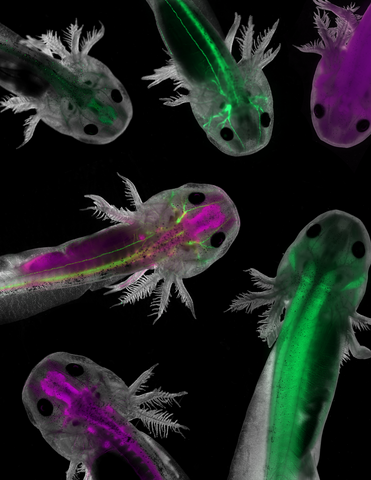
Axolotls—a type of salamander—that have been genetically modified so that various parts of their nervous systems glow purple and green. Researchers often study axolotls for their extensive regenerative abilities. They can regrow tails, limbs, spinal cords, brains, and more. The researcher who took this image focuses on the role of the peripheral nervous system during limb regeneration.
This image was captured using a stereo microscope.
Related to images 6927 and 6932.
This image was captured using a stereo microscope.
Related to images 6927 and 6932.
Prayag Murawala, MDI Biological Laboratory and Hannover Medical School.
View Media
2526: Activation energy (with labels)
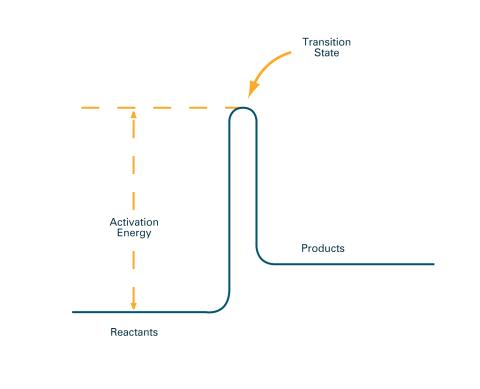
2526: Activation energy (with labels)
To become products, reactants must overcome an energy hill. See image 2525 for an unlabeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
5764: Host infection stimulates antibiotic resistance
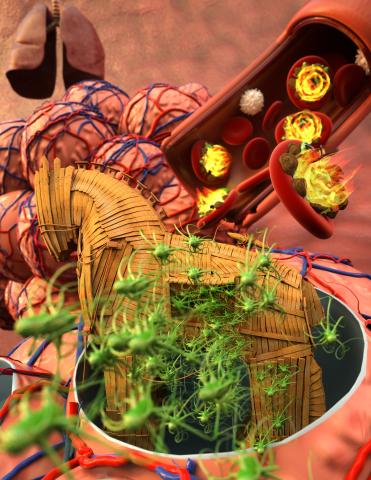
5764: Host infection stimulates antibiotic resistance
This illustration shows pathogenic bacteria behave like a Trojan horse: switching from antibiotic susceptibility to resistance during infection. Salmonella are vulnerable to antibiotics while circulating in the blood (depicted by fire on red blood cell) but are highly resistant when residing within host macrophages. This leads to treatment failure with the emergence of drug-resistant bacteria.
This image was chosen as a winner of the 2016 NIH-funded research image call, and the research was funded in part by NIGMS.
View Media
This image was chosen as a winner of the 2016 NIH-funded research image call, and the research was funded in part by NIGMS.
5883: Beta-galactosidase montage showing cryo-EM improvement--gradient background
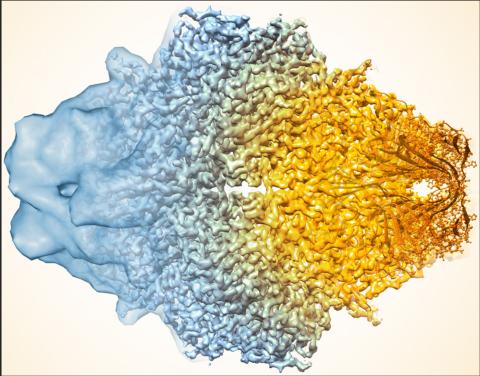
5883: Beta-galactosidase montage showing cryo-EM improvement--gradient background
Composite image of beta-galactosidase showing how cryo-EM’s resolution has improved dramatically in recent years. Older images to the left, more recent to the right. Related to image 5882. NIH Director Francis Collins featured this on his blog on January 14, 2016.
Veronica Falconieri, Sriram Subramaniam Lab, National Cancer Institute
View Media
2514: Life of an AIDS virus (with labels)
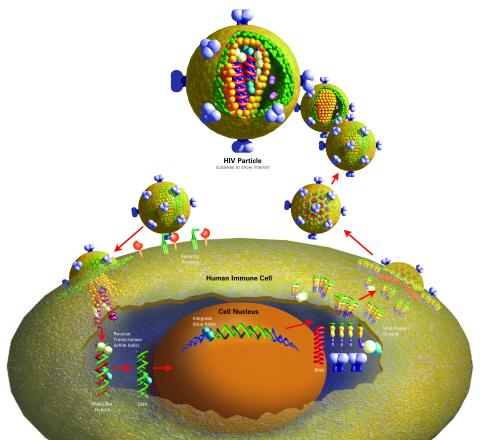
2514: Life of an AIDS virus (with labels)
HIV is a retrovirus, a type of virus that carries its genetic material not as DNA but as RNA. Long before anyone had heard of HIV, researchers in labs all over the world studied retroviruses, tracing out their life cycle and identifying the key proteins the viruses use to infect cells. When HIV was identified as a retrovirus, these studies gave AIDS researchers an immediate jump-start. The previously identified viral proteins became initial drug targets. See images 2513 and 2515 for other versions of this illustration. Featured in The Structures of Life.
Crabtree + Company
View Media
3677: Human skeletal muscle
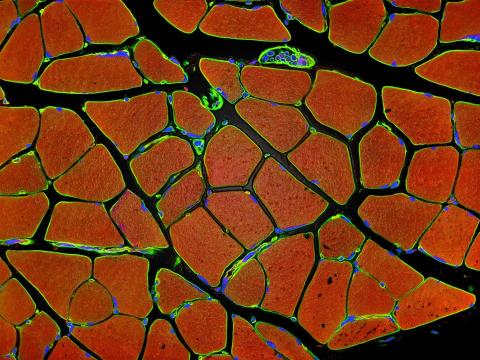
3677: Human skeletal muscle
Cross section of human skeletal muscle. Image taken with a confocal fluorescent light microscope.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
2550: Introns
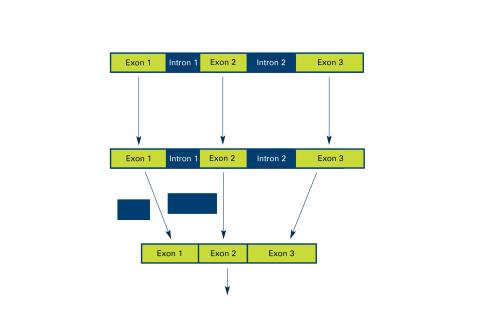
2550: Introns
Genes are often interrupted by stretches of DNA (introns, blue) that do not contain instructions for making a protein. The DNA segments that do contain protein-making instructions are known as exons (green). See image 2551 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
2716: Mycobacterium tuberculosis
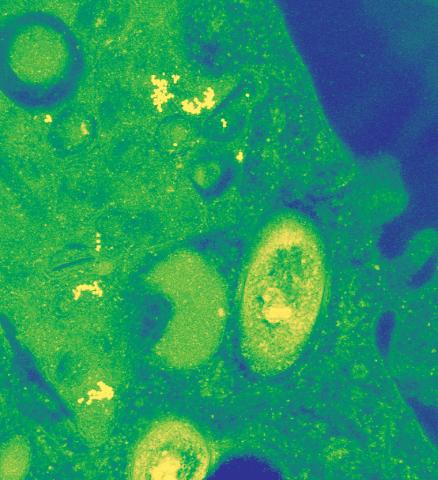
2716: Mycobacterium tuberculosis
Mycobacterium tuberculosis, the bacterium that causes tuberculosis, has infected one-quarter of the world's population and causes more than one million deaths each year, according to the World Health Organization.
Reuben Peters, Iowa State University
View Media
3278: Induced pluripotent stem cells from skin
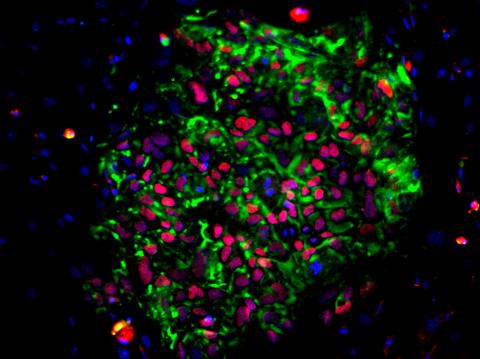
3278: Induced pluripotent stem cells from skin
These induced pluripotent stem cells (iPS cells) were derived from a woman's skin. Green and red indicate proteins found in reprogrammed cells but not in skin cells (TRA1-62 and NANOG). These cells can then develop into different cell types. Image and caption information courtesy of the California Institute for Regenerative Medicine. Related to image 3279.
Kathrin Plath lab, University of California, Los Angeles, via CIRM
View Media
2757: Draper, shown in the fatbody of a Drosophila melanogaster larva
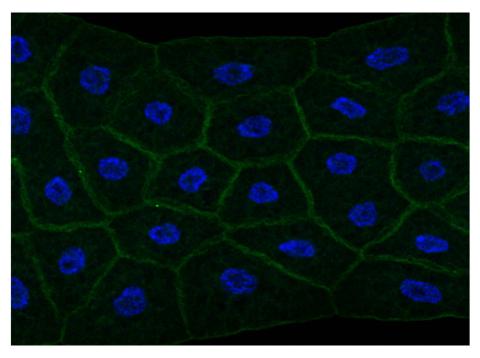
2757: Draper, shown in the fatbody of a Drosophila melanogaster larva
The fly fatbody is a nutrient storage and mobilization organ akin to the mammalian liver. The engulfment receptor Draper (green) is located at the cell surface of fatbody cells. The cell nuclei are shown in blue.
Christina McPhee and Eric Baehrecke, University of Massachusetts Medical School
View Media
3525: Bacillus anthracis being killed
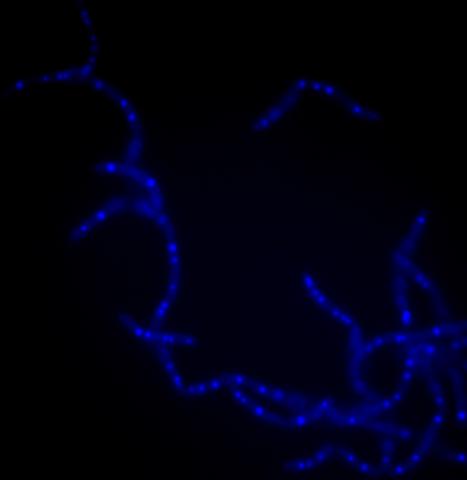
3525: Bacillus anthracis being killed
Bacillus anthracis (anthrax) cells being killed by a fluorescent trans-translation inhibitor, which disrupts bacterial protein synthesis. The inhibitor is naturally fluorescent and looks blue when it is excited by ultraviolet light in the microscope. This is a color version of Image 3481.
Kenneth Keiler, Penn State University
View Media
2567: Haplotypes (with labels)
2567: Haplotypes (with labels)
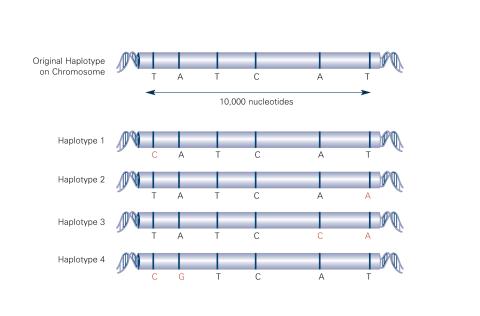
Haplotypes are combinations of gene variants that are likely to be inherited together within the same chromosomal region. In this example, an original haplotype (top) evolved over time to create three newer haplotypes that each differ by a few nucleotides (red). See image 2566 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
3725: Fluorescent microscopy of kidney tissue--close-up
3725: Fluorescent microscopy of kidney tissue--close-up
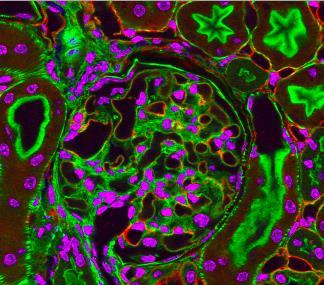
This photograph of kidney tissue, taken using fluorescent light microscopy, shows a close-up view of part of image 3723. Kidneys filter the blood, removing waste and excessive fluid, which is excreted in urine. The filtration system is made up of components that include glomeruli (for example, the round structure taking up much of the image's center is a glomerulus) and tubules (seen in cross-section here with their inner lining stained green). Related to image 3675 .
Tom Deerinck , National Center for Microscopy and Imaging Research
View Media
1019: Lily mitosis 13
1019: Lily mitosis 13
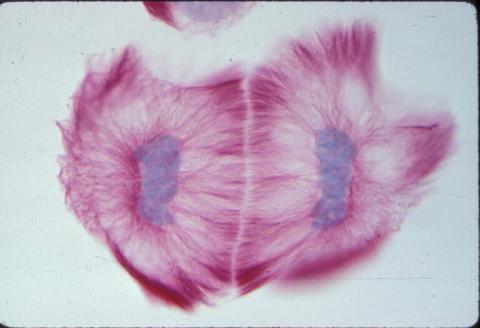
A light microscope image of cells from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, two cells have formed after a round of mitosis.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1018, and 1021.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1018, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
3405: Disrupted and restored vasculature development in frog embryos
3405: Disrupted and restored vasculature development in frog embryos
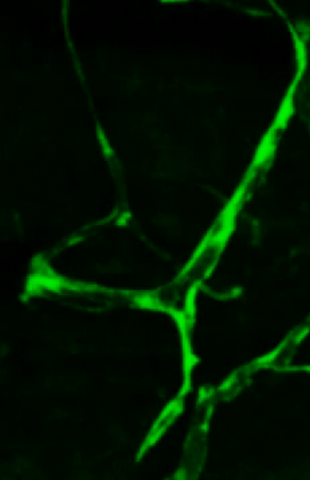
Disassembly of vasculature and reassembly after addition and then washout of 250 µM TBZ in kdr:GFP frogs. Related to images 3403 and 3404.
Hye Ji Cha, University of Texas at Austin
View Media
1332: Mitosis - telophase
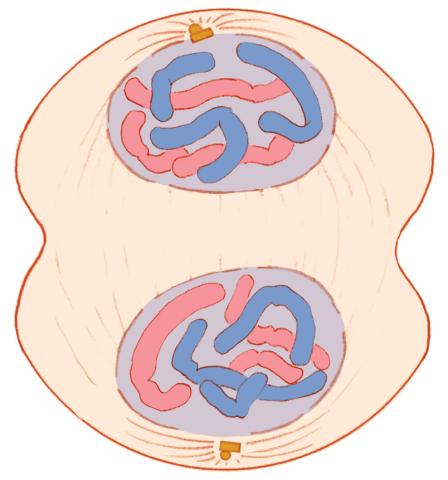
1332: Mitosis - telophase
Telophase during mitosis: Nuclear membranes form around each of the two sets of chromosomes, the chromosomes begin to spread out, and the spindle begins to break down. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
2693: Fruit fly in the pink
2693: Fruit fly in the pink
Fruit flies are a common model organism for basic medical research.
Crabtree + Company
View Media
1085: Natcher Building 05
1085: Natcher Building 05
NIGMS staff are located in the Natcher Building on the NIH campus.
Alisa Machalek, National Institute of General Medical Sciences
View Media

1120: Superconducting magnet
1120: Superconducting magnet
Superconducting magnet for NMR research, from the February 2003 profile of Dorothee Kern in Findings.
Mike Lovett
View Media
6994: Respiratory droplet
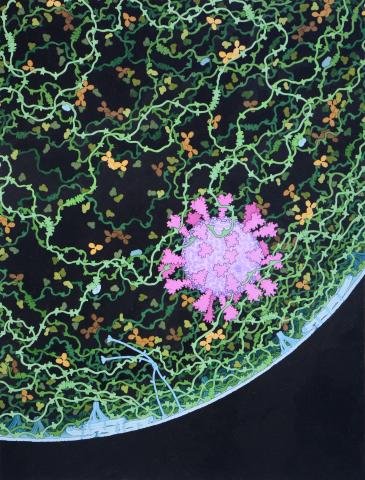
6994: Respiratory droplet
This painting shows a cross section of a small respiratory droplet, like the ones that are thought to transmit SARS-CoV-2, the virus that causes COVID-19. The virus is shown in pink, and the droplet is also filled with molecules that are present in the respiratory tract, including mucins (green), pulmonary surfactant proteins and lipids (blue), and antibodies (tan).
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media