Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
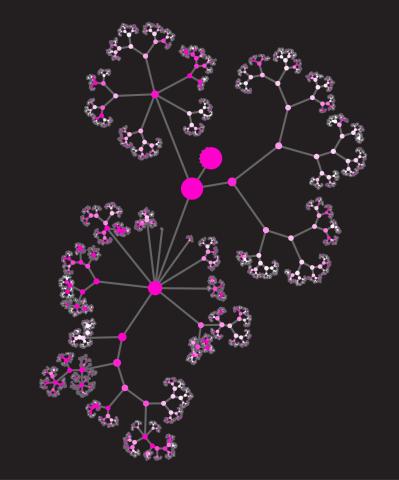
3436: Network diagram of genes, cellular components and processes (unlabeled)
3436: Network diagram of genes, cellular components and processes (unlabeled)
This image shows the hierarchical ontology of genes, cellular components and processes derived from large genomic datasets. From Dutkowski et al. A gene ontology inferred from molecular networks Nat Biotechnol. 2013 Jan;31(1):38-45. Related to 3437.
Janusz Dutkowski and Trey Ideker
View Media
2635: Mitochondria and endoplasmic reticulum
2635: Mitochondria and endoplasmic reticulum
A computer model shows how the endoplasmic reticulum is close to and almost wraps around mitochondria in the cell. The endoplasmic reticulum is lime green and the mitochondria are yellow. This image relates to a July 27, 2009 article in Computing Life.
Bridget Wilson, University of New Mexico
View Media

3265: Microfluidic chip
3265: Microfluidic chip
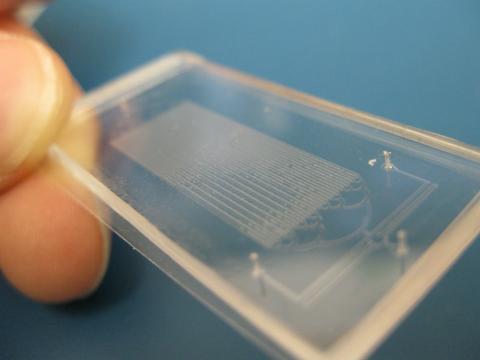
Microfluidic chips have many uses in biology labs. The one shown here was used by bioengineers to study bacteria, allowing the researchers to synchronize their fluorescing so they would blink in unison. Related to images 3266 and 3268. From a UC San Diego news release, "Researchers create living 'neon signs' composed of millions of glowing bacteria."
Jeff Hasty Lab, UC San Diego
View Media
3618: Hair cells: the sound-sensing cells in the ear
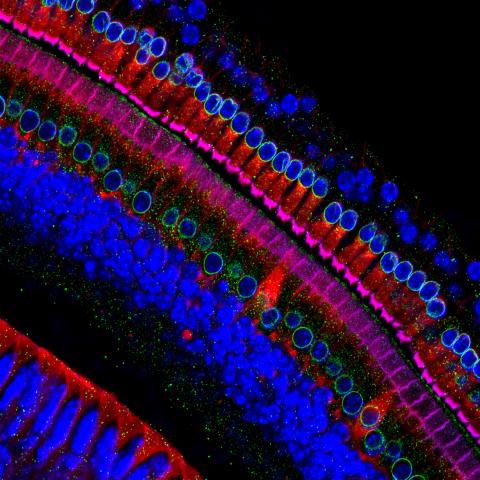
3618: Hair cells: the sound-sensing cells in the ear
These cells get their name from the hairlike structures that extend from them into the fluid-filled tube of the inner ear. When sound reaches the ear, the hairs bend and the cells convert this movement into signals that are relayed to the brain. When we pump up the music in our cars or join tens of thousands of cheering fans at a football stadium, the noise can make the hairs bend so far that they actually break, resulting in long-term hearing loss.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Henning Horn, Brian Burke, and Colin Stewart, Institute of Medical Biology, Agency for Science, Technology, and Research, Singapore
View Media
2541: Nucleotides make up DNA
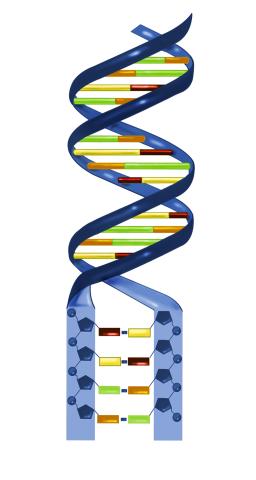
2541: Nucleotides make up DNA
DNA consists of two long, twisted chains made up of nucleotides. Each nucleotide contains one base, one phosphate molecule, and the sugar molecule deoxyribose. The bases in DNA nucleotides are adenine, thymine, cytosine, and guanine. See image 2542 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

3689: Computer sketch of bird-and-flower DNA origami
3689: Computer sketch of bird-and-flower DNA origami
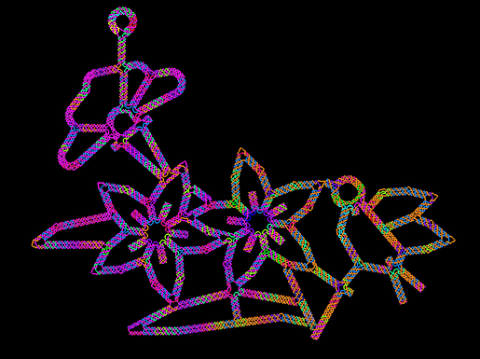
A computer-generated sketch of a DNA origami folded into a flower-and-bird structure. See also related image 3690.
Hao Yan, Arizona State University
View Media
3648: Symmetrically and asymmetrically elongating cells
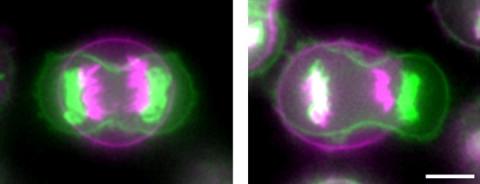
3648: Symmetrically and asymmetrically elongating cells
Merged fluorescent images of symmetrically (left) or asymmetrically (right) elongating HeLa cells at the end of early anaphase (magenta) and late anaphase (green). Chromosomes and cortical actin are visualized by expressing mCherry-histone H2B and Lifeact-mCherry. Scale bar, 10µm. See the PubMed abstract of this research.
Tomomi Kiyomitsu and Iain M. Cheeseman, Whitehead Institute for Biomedical Research
View Media
2494: VDAC-1 (3)
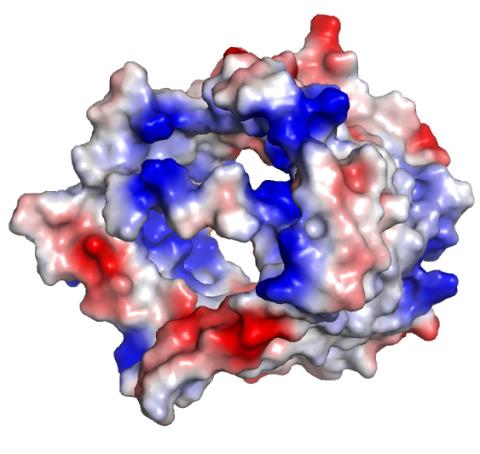
2494: VDAC-1 (3)
The structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to images 2491, 2495, and 2488.
Related to images 2491, 2495, and 2488.
Gerhard Wagner, Harvard Medical School
View Media
5762: Panorama view of golden mitochondria
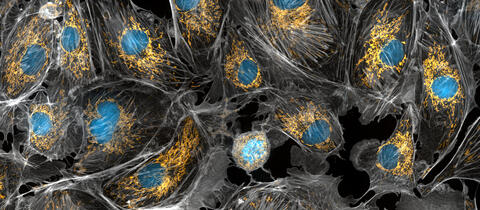
5762: Panorama view of golden mitochondria
Mitochondria are the powerhouses of the cells, generating the energy the cells need to do their tasks and to stay alive. Researchers have studied mitochondria for some time because when these cell organelles don't work as well as they should, several diseases develop. In this photograph of cow cells taken with a microscope, the mitochondria were stained in bright yellow to visualize them in the cell. The large blue dots are the cell nuclei and the gray web is the cytoskeleton of the cells.
Torsten Wittmann, University of California, San Francisco
View Media
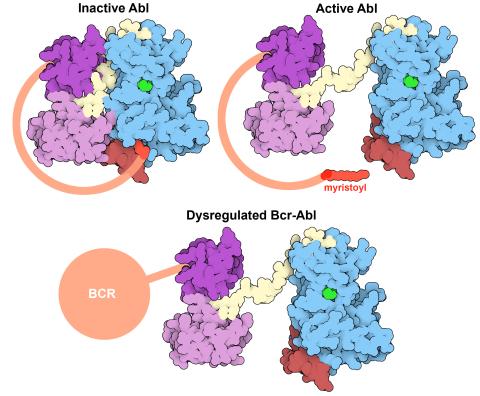
7004: Protein kinases as cancer chemotherapy targets
7004: Protein kinases as cancer chemotherapy targets
Protein kinases—enzymes that add phosphate groups to molecules—are cancer chemotherapy targets because they play significant roles in almost all aspects of cell function, are tightly regulated, and contribute to the development of cancer and other diseases if any alterations to their regulation occur. Genetic abnormalities affecting the c-Abl tyrosine kinase are linked to chronic myelogenous leukemia, a cancer of immature cells in the bone marrow. In the noncancerous form of the protein, binding of a myristoyl group to the kinase domain inhibits the activity of the protein until it is needed (top left shows the inactive form, top right shows the open and active form). The cancerous variant of the protein, called Bcr-Abl, lacks this autoinhibitory myristoyl group and is continually active (bottom). ATP is shown in green bound in the active site of the kinase.
Find these in the RCSB Protein Data Bank: c-Abl tyrosine kinase and regulatory domains (PDB entry 1OPL) and F-actin binding domain (PDB entry 1ZZP).
Find these in the RCSB Protein Data Bank: c-Abl tyrosine kinase and regulatory domains (PDB entry 1OPL) and F-actin binding domain (PDB entry 1ZZP).
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media

6965: Dividing cell
6965: Dividing cell
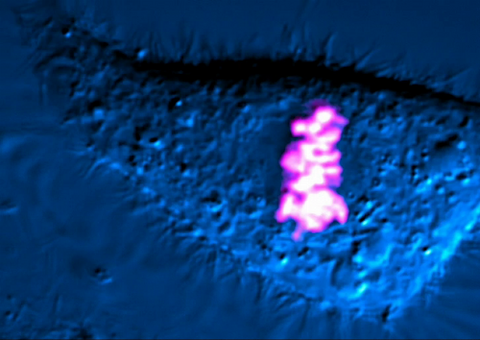
As this cell was undergoing cell division, it was imaged with two microscopy techniques: differential interference contrast (DIC) and confocal. The DIC view appears in blue and shows the entire cell. The confocal view appears in pink and shows the chromosomes.
Dylan T. Burnette, Vanderbilt University School of Medicine.
View Media
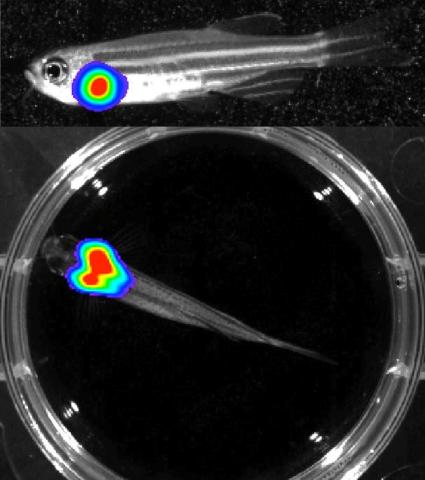
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression. This is the lateral and overhead (Bottom) view.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
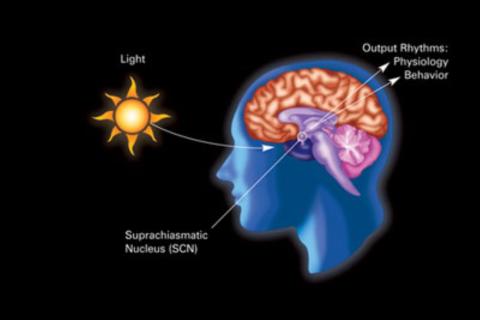
2841: Circadian rhythm
2841: Circadian rhythm
The human body keeps time with a master clock called the suprachiasmatic nucleus or SCN. Situated inside the brain, it's a tiny sliver of tissue about the size of a grain of rice, located behind the eyes. It sits quite close to the optic nerve, which controls vision, and this means that the SCN "clock" can keep track of day and night. The SCN helps control sleep by coordinating the actions of billions of miniature "clocks" throughout the body. These aren't actually clocks, but rather are ensembles of genes inside clusters of cells that switch on and off in a regular, 24-hour cycle in our physiological day.
Crabtree + Company
View Media
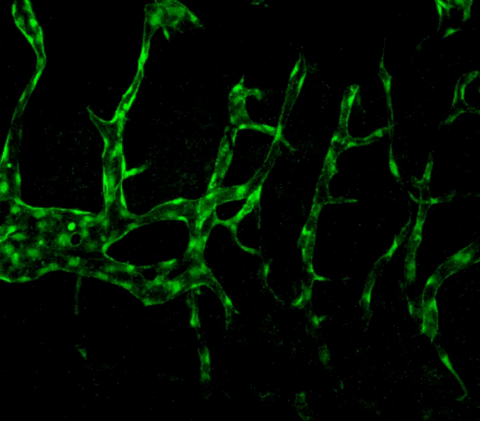
3404: Normal vascular development in frog embryos
3404: Normal vascular development in frog embryos
Hye Ji Cha, University of Texas at Austin
View Media

2397: Bovine milk alpha-lactalbumin (1)
2397: Bovine milk alpha-lactalbumin (1)
A crystal of bovine milk alpha-lactalbumin protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
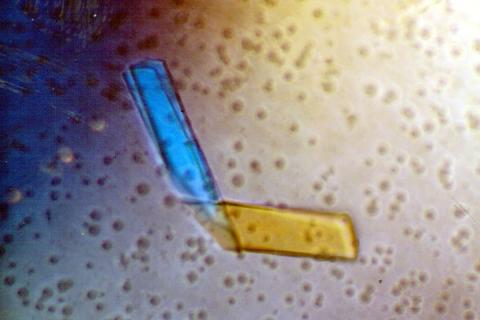
2399: Bence Jones protein MLE
2399: Bence Jones protein MLE
A crystal of Bence Jones protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
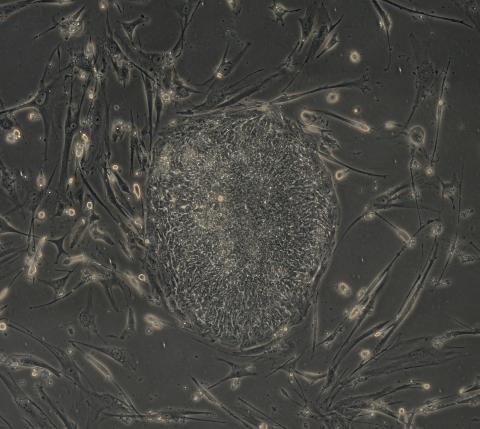
2605: Induced stem cells from adult skin 03
2605: Induced stem cells from adult skin 03
The human skin cells pictured contain genetic modifications that make them pluripotent, essentially equivalent to embryonic stem cells. A scientific team from the University of Wisconsin-Madison including researchers Junying Yu, James Thomson, and their colleagues produced the transformation by introducing a set of four genes into human fibroblasts, skin cells that are easy to obtain and grow in culture.
James Thomson, University of Wisconsin-Madison
View Media
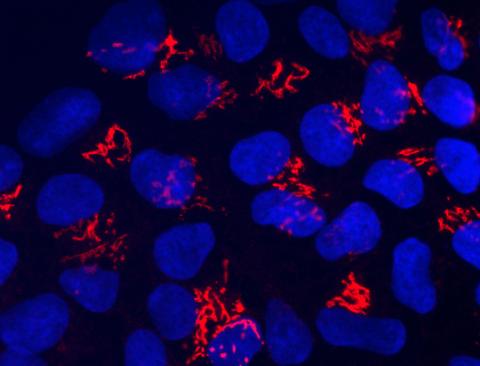
3341: Suicidal Stem Cells
3341: Suicidal Stem Cells
Embryonic stem cells store pre-activated Bax (red) in the Golgi, near the nucleus (blue). Featured in the June 21, 2012, issue of Biomedical Beat.
Mohanish Deshmukh
View Media

6344: Drosophila
6344: Drosophila
Two adult fruit flies (Drosophila)
Dr. Vicki Losick, MDI Biological Laboratory, www.mdibl.org
View Media
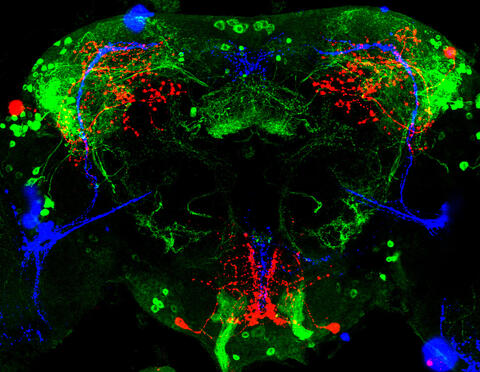
3754: Circadian rhythm neurons in the fruit fly brain
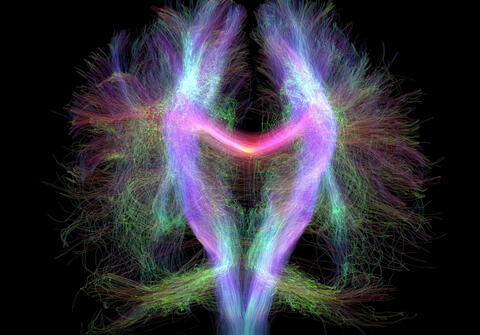
3754: Circadian rhythm neurons in the fruit fly brain
Some nerve cells (neurons) in the brain keep track of the daily cycle. This time-keeping mechanism, called the circadian clock, is found in all animals including us. The circadian clock controls our daily activities such as sleep and wakefulness. Researchers are interested in finding the neuron circuits involved in this time keeping and how the information about daily time in the brain is relayed to the rest of the body. In this image of a brain of the fruit fly Drosophila the time-of-day information flowing through the brain has been visualized by staining the neurons involved: clock neurons (shown in blue) function as "pacemakers" by communicating with neurons that produce a short protein called leucokinin (LK) (red), which, in turn, relays the time signal to other neurons, called LK-R neurons (green). This signaling cascade set in motion by the pacemaker neurons helps synchronize the fly's daily activity with the 24-hour cycle. To learn more about what scientists have found out about circadian pacemaker neurons in the fruit fly see this news release by New York University. This work was featured in the Biomedical Beat blog post Cool Image: A Circadian Circuit.
Justin Blau, New York University
View Media

2683: GFP sperm
2683: GFP sperm
Fruit fly sperm cells glow bright green when they express the gene for green fluorescent protein (GFP).
View Media
3284: Neurons from human ES cells
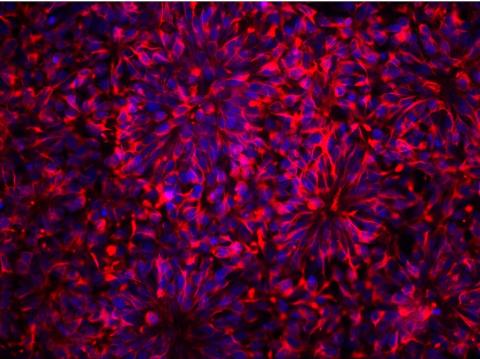
3284: Neurons from human ES cells
These neural precursor cells were derived from human embryonic stem cells. The neural cell bodies are stained red, and the nuclei are blue. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Xianmin Zeng lab, Buck Institute for Age Research, via CIRM
View Media
6661: Zebrafish embryo showing vasculature
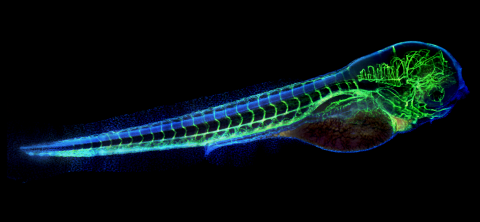
6661: Zebrafish embryo showing vasculature
A zebrafish embryo. The blue areas are cell bodies, the green lines are blood vessels, and the red glow is blood. This image was created by stitching together five individual images captured with a hyperspectral multipoint confocal fluorescence microscope that was developed at the Eliceiri Lab.
Kevin Eliceiri, University of Wisconsin-Madison.
View Media
3292: Centrioles anchor cilia in planaria
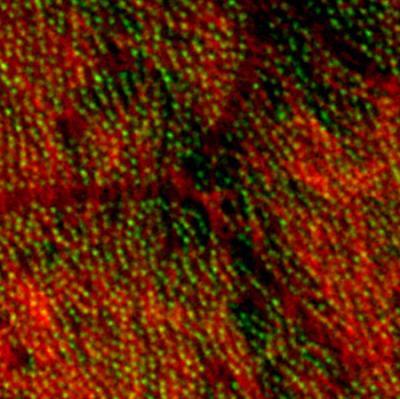
3292: Centrioles anchor cilia in planaria
Centrioles (green) anchor cilia (red), which project on the surface of pharynx cells of the freshwater planarian Schmidtea mediterranea. Centrioles require cellular structures called centrosomes for assembly in other animal species, but this flatworm known for its regenerative ability was unexpectedly found to lack centrosomes. From a Stowers University news release.
Juliette Azimzadeh, University of California, San Francisco
View Media
3396: Myelinated axons 1
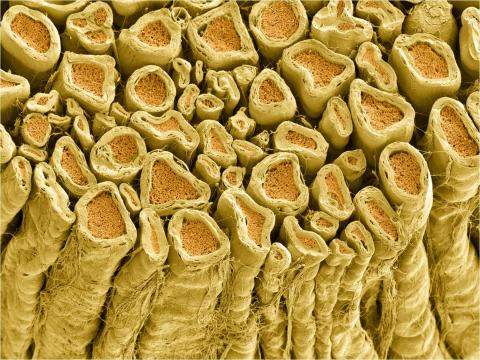
3396: Myelinated axons 1
Myelinated axons in a rat spinal root. Myelin is a type of fat that forms a sheath around and thus insulates the axon to protect it from losing the electrical current needed to transmit signals along the axon. The axoplasm inside the axon is shown in pink. Related to 3397.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
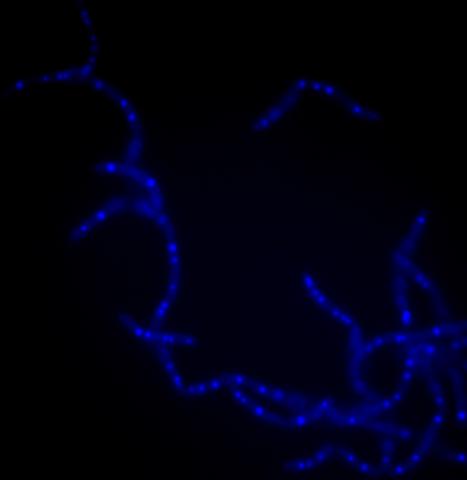
3525: Bacillus anthracis being killed
3525: Bacillus anthracis being killed
Bacillus anthracis (anthrax) cells being killed by a fluorescent trans-translation inhibitor, which disrupts bacterial protein synthesis. The inhibitor is naturally fluorescent and looks blue when it is excited by ultraviolet light in the microscope. This is a color version of Image 3481.
Kenneth Keiler, Penn State University
View Media
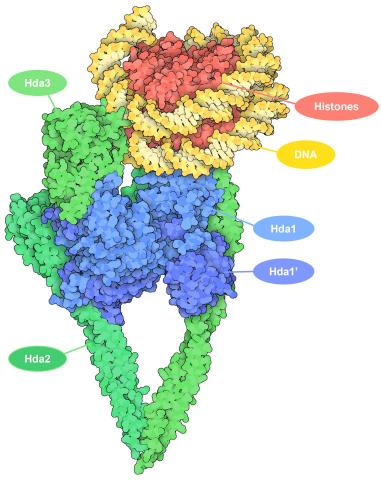
7001: Histone deacetylases
7001: Histone deacetylases
The human genome contains much of the information needed for every cell in the body to function. However, different types of cells often need different types of information. Access to DNA is controlled, in part, by how tightly it’s wrapped around proteins called histones to form nucleosomes. The complex shown here, from yeast cells (PDB entry 6Z6P), includes several histone deacetylase (HDAC) enzymes (green and blue) bound to a nucleosome (histone proteins in red; DNA in yellow). The yeast HDAC enzymes are similar to the human enzymes. Two enzymes form a V-shaped clamp (green) that holds the other others, a dimer of the Hda1 enzymes (blue). In this assembly, Hda1 is activated and positioned to remove acetyl groups from histone tails.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
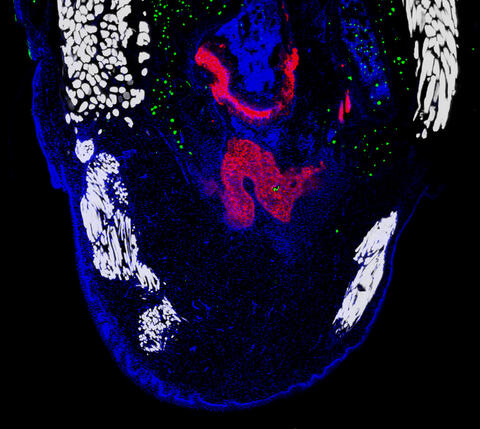
6968: Regenerating lizard tail
6968: Regenerating lizard tail
The interior of a regenerating lizard tail 14 days after the original tail was amputated. Cell nuclei (blue), proliferating cells (green), cartilage (red), and muscle (white) have been visualized with immunofluorescence staining.
Thomas Lozito, University of Southern California.
View Media

6777: Human endoplasmic reticulum membrane protein complex
6777: Human endoplasmic reticulum membrane protein complex
A 3D model of the human endoplasmic reticulum membrane protein complex (EMC) that identifies its nine essential subunits. The EMC plays an important role in making membrane proteins, which are essential for all cellular processes. This is the first atomic-level depiction of the EMC. Its structure was obtained using single-particle cryo-electron microscopy.
Rebecca Voorhees, California Institute of Technology.
View Media
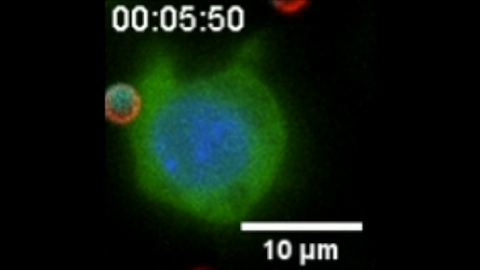
6801: “Two-faced” Janus particle activating a macrophage
6801: “Two-faced” Janus particle activating a macrophage
A macrophage—a type of immune cell that engulfs invaders—“eats” and is activated by a “two-faced” Janus particle. The particle is called “two-faced” because each of its two hemispheres is coated with a different type of molecule, shown here in red and cyan. During macrophage activation, a transcription factor tagged with a green fluorescence protein (NF-κB) gradually moves from the cell’s cytoplasm into its nucleus and causes DNA transcription. The distribution of molecules on “two-faced” Janus particles can be altered to control the activation of immune cells. Details on this “geometric manipulation” strategy can be found in the Proceedings of the National Academy of Sciences paper "Geometrical reorganization of Dectin-1 and TLR2 on single phagosomes alters their synergistic immune signaling" by Li et al. and the Scientific Reports paper "Spatial organization of FcγR and TLR2/1 on phagosome membranes differentially regulates their synergistic and inhibitory receptor crosstalk" by Li et al. This video was captured using epi-fluorescence microscopy.
Related to video 6800.
Related to video 6800.
Yan Yu, Indiana University, Bloomington.
View Media

2395: Fungal lipase (1)
2395: Fungal lipase (1)
Crystals of fungal lipase protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
3748: Cryo-electron microscopy of the dengue virus showing protective membrane and membrane proteins
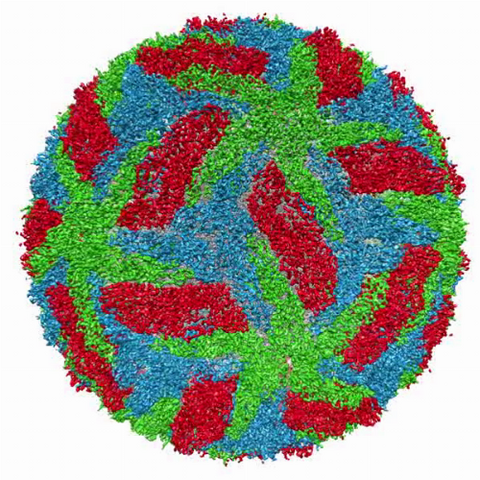
3748: Cryo-electron microscopy of the dengue virus showing protective membrane and membrane proteins
Dengue virus is a mosquito-borne illness that infects millions of people in the tropics and subtropics each year. Like many viruses, dengue is enclosed by a protective membrane. The proteins that span this membrane play an important role in the life cycle of the virus. Scientists used cryo-EM to determine the structure of a dengue virus at a 3.5-angstrom resolution to reveal how the membrane proteins undergo major structural changes as the virus matures and infects a host. For more on cryo-EM see the blog post Cryo-Electron Microscopy Reveals Molecules in Ever Greater Detail. Related to image 3756.
Hong Zhou, UCLA
View Media
6792: Yeast cells with nuclei and contractile rings
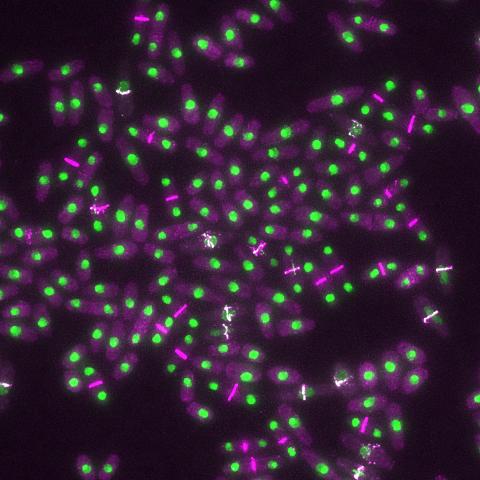
6792: Yeast cells with nuclei and contractile rings
Yeast cells with nuclei shown in green and contractile rings shown in magenta. Nuclei store DNA, and contractile rings help cells divide. This image was captured using wide-field microscopy with deconvolution.
Related to images 6791, 6793, 6794, 6797, 6798, and videos 6795 and 6796.
Related to images 6791, 6793, 6794, 6797, 6798, and videos 6795 and 6796.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media

6902: Arachnoidiscus diatom
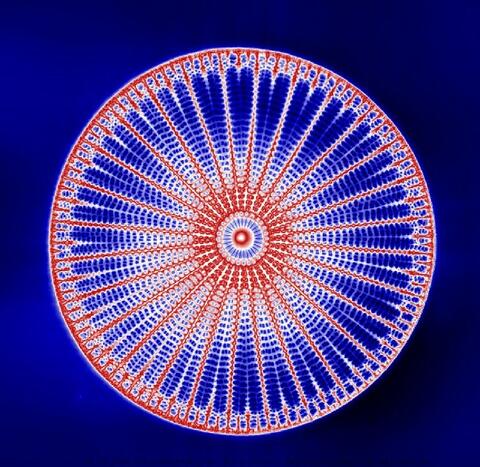
6902: Arachnoidiscus diatom
An Arachnoidiscus diatom with a diameter of 190µm. Diatoms are microscopic algae that have cell walls made of silica, which is the strongest known biological material relative to its density. In Arachnoidiscus, the cell wall is a radially symmetric pillbox-like shell composed of overlapping halves that contain intricate and delicate patterns. Sometimes, Arachnoidiscus is called “a wheel of glass.”
This image was taken with the orientation-independent differential interference contrast microscope.
This image was taken with the orientation-independent differential interference contrast microscope.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media

2423: Protein map
2423: Protein map
Network diagram showing a map of protein-protein interactions in a yeast (Saccharomyces cerevisiae) cell. This cluster includes 78 percent of the proteins in the yeast proteome. The color of a node represents the phenotypic effect of removing the corresponding protein (red, lethal; green, nonlethal; orange, slow growth; yellow, unknown).
Hawoong Jeong, KAIST, Korea
View Media
3646: Cells lining the trachea
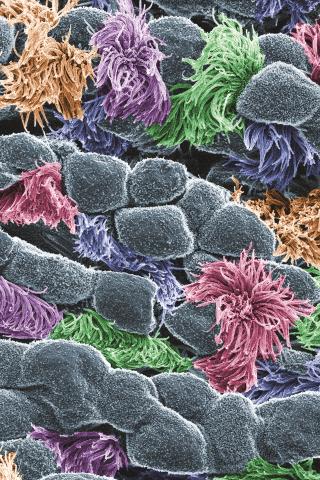
3646: Cells lining the trachea
In this image, viewed with a ZEISS ORION NanoFab microscope, the community of cells lining a mouse airway is magnified more than 10,000 times. This collection of cells, known as the mucociliary escalator, is also found in humans. It is our first line of defense against inhaled bacteria, allergens, pollutants, and debris. Malfunctions in the system can cause or aggravate lung infections and conditions such as asthma and chronic obstructive pulmonary disease. The cells shown in gray secrete mucus, which traps inhaled particles. The colored cells sweep the mucus layer out of the lungs.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Eva Mutunga and Kate Klein, University of the District of Columbia and National Institute of Standards and Technology
View Media
5759: TEM cross-section of C. elegans (roundworm)
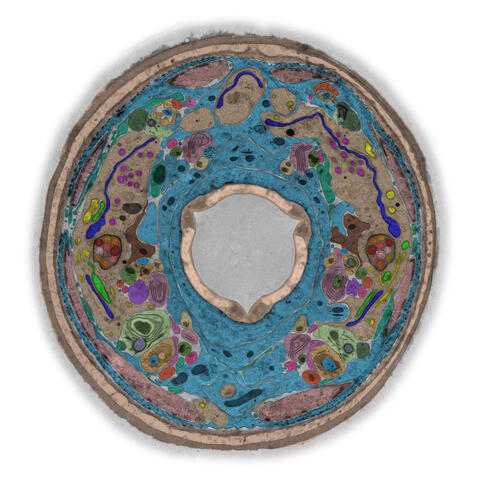
5759: TEM cross-section of C. elegans (roundworm)
The worm Caenorhabditis elegans is a popular laboratory animal because its small size and fairly simple body make it easy to study. Scientists use this small worm to answer many research questions in developmental biology, neurobiology, and genetics. This image, which was taken with transmission electron microscopy (TEM), shows a cross-section through C. elegans, revealing various internal structures.
The image is from a figure in an article published in the journal eLife. There is an annotated version of this graphic at 5760.
The image is from a figure in an article published in the journal eLife. There is an annotated version of this graphic at 5760.
Piali Sengupta, Brandeis University
View Media

6756: Honeybees marked with paint
6756: Honeybees marked with paint
Researchers doing behavioral experiments with honeybees sometimes use paint or enamel to give individual bees distinguishing marks. The elaborate social structure and impressive learning and navigation abilities of bees make them good models for behavioral and neurobiological research. Since the sequencing of the honeybee genome, published in 2006, bees have been used increasingly for research into the molecular basis for social interaction and other complex behaviors.
Gene Robinson, University of Illinois at Urbana-Champaign.
View Media
2374: Protein from Methanobacterium thermoautotrophicam
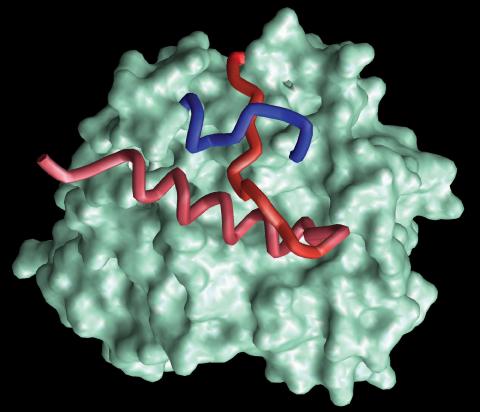
2374: Protein from Methanobacterium thermoautotrophicam
A knotted protein from an archaebacterium called Methanobacterium thermoautotrophicam. This organism breaks down waste products and produces methane gas. Protein folding theory previously held that forming a knot was beyond the ability of a protein, but this structure, determined at Argonne's Structural Biology Center, proves differently. Researchers theorize that this knot stabilizes the amino acid subunits of the protein.
Midwest Center For Structural Genomics, PSI
View Media
2565: Recombinant DNA (with labels)
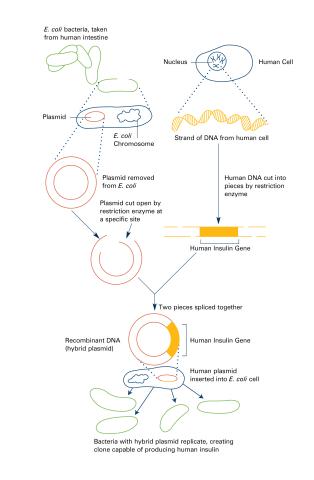
2565: Recombinant DNA (with labels)
To splice a human gene (in this case, the one for insulin) into a plasmid, scientists take the plasmid out of an E. coli bacterium, cut the plasmid with a restriction enzyme, and splice in insulin-making human DNA. The resulting hybrid plasmid can be inserted into another E. coli bacterium, where it multiplies along with the bacterium. There, it can produce large quantities of insulin. See image 2564 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
2428: Colorful cells
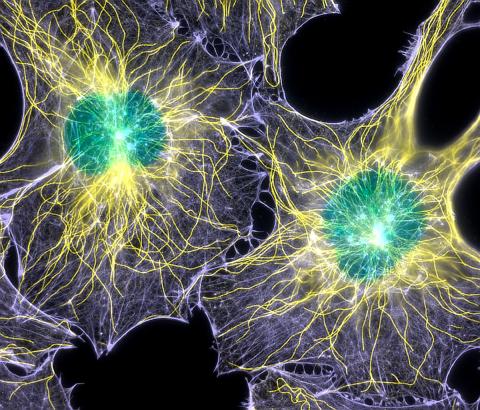
2428: Colorful cells
Actin (purple), microtubules (yellow), and nuclei (green) are labeled in these cells by immunofluorescence. This image won first place in the Nikon 2003 Small World photo competition.
Torsten Wittmann, Scripps Research Institute
View Media
1307: Cisternae maturation model
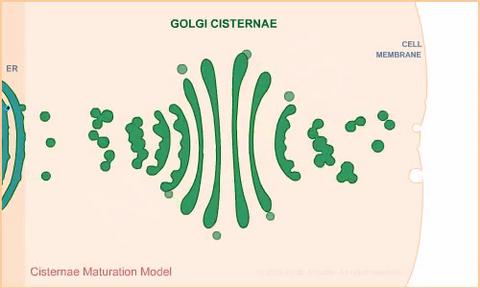
1307: Cisternae maturation model
Animation for the cisternae maturation model of Golgi transport.
Judith Stoffer
View Media
2563: Epigenetic code (with labels)
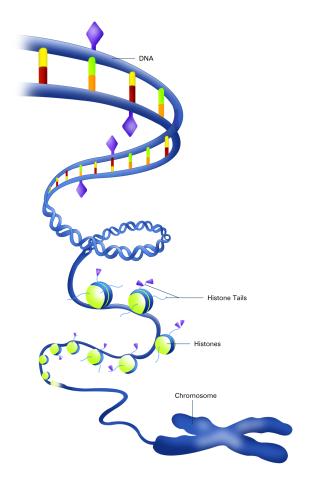
2563: Epigenetic code (with labels)
The "epigenetic code" controls gene activity with chemical tags that mark DNA (purple diamonds) and the "tails" of histone proteins (purple triangles). These markings help determine whether genes will be transcribed by RNA polymerase. Genes hidden from access to RNA polymerase are not expressed. See image 2562 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

6356: H1N1 Influenza Virus
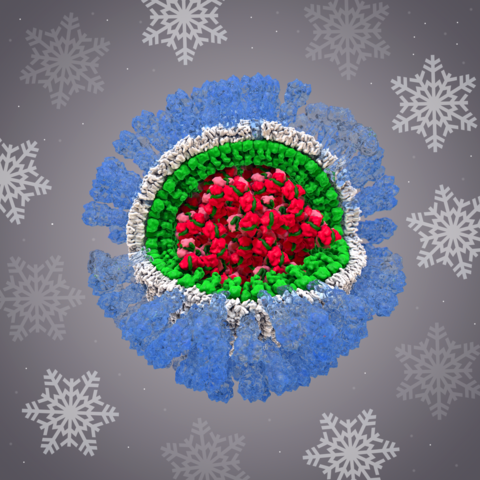
6356: H1N1 Influenza Virus
Related to image 6355.
Dr. Rommie Amaro, University of California, San Diego
View Media
2347: Cysteine dioxygenase from mouse
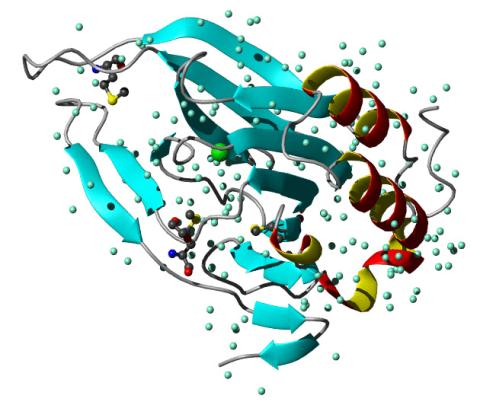
2347: Cysteine dioxygenase from mouse
Model of the mammalian iron enzyme cysteine dioxygenase from a mouse.
Center for Eukaryotic Structural Genomics, PSI
View Media
3289: Smooth muscle from mouse stem cells
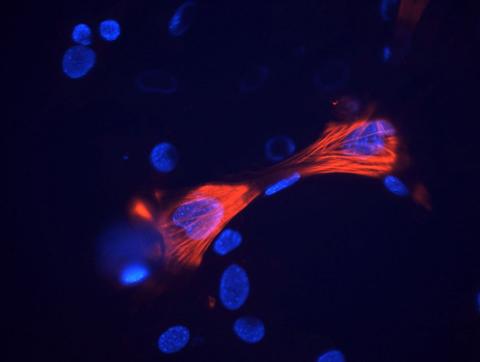
3289: Smooth muscle from mouse stem cells
These smooth muscle cells were derived from mouse neural crest stem cells. Red indicates smooth muscle proteins, blue indicates nuclei. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Deepak Srivastava, Gladstone Institutes, via CIRM
View Media
3264: Peripheral nerve cell derived from ES cells
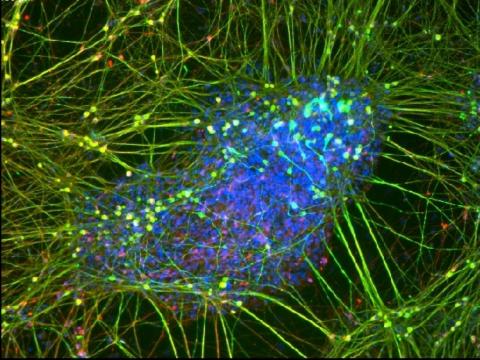
3264: Peripheral nerve cell derived from ES cells
A peripheral nerve cell made from human embryonic stem cell-derived neural crest stem cells. The nucleus is shown in blue, and nerve cell proteins peripherin and beta-tubulin (Tuj1) are shown in green and red, respectively. Related to image 3263.
Stephen Dalton, University of Georgia
View Media
3792: Nucleolus subcompartments spontaneously self-assemble 3
3792: Nucleolus subcompartments spontaneously self-assemble 3
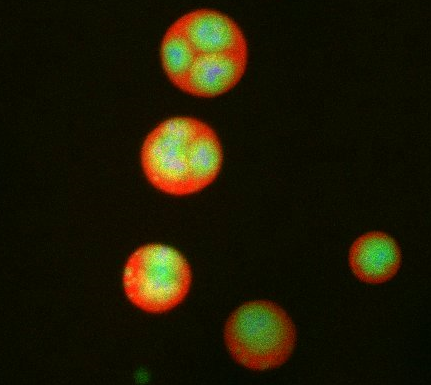
What looks a little like distant planets with some mysterious surface features are actually assemblies of proteins normally found in the cell's nucleolus, a small but very important protein complex located in the cell's nucleus. It forms on the chromosomes at the location where the genes for the RNAs are that make up the structure of the ribosome, the indispensable cellular machine that makes proteins from messenger RNAs.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This photo of nucleolus proteins in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows each of the nucleolus compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue). The researchers have found that these compartments spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, video 3791 and image 3793.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This photo of nucleolus proteins in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows each of the nucleolus compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue). The researchers have found that these compartments spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, video 3791 and image 3793.
Nilesh Vaidya, Princeton University
View Media
6570: Stress Response in Cells
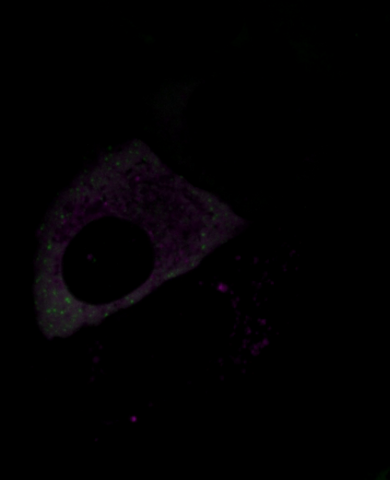
6570: Stress Response in Cells
Two highly stressed osteosarcoma cells are shown with a set of green droplet-like structures followed by a second set of magenta droplets. These droplets are composed of fluorescently labeled stress-response proteins, either G3BP or UBQLN2 (Ubiquilin-2). Each protein is undergoing a fascinating process, called phase separation, in which a non-membrane bound compartment of the cytoplasm emerges with a distinct environment from the surrounding cytoplasm. Subsequently, the proteins fuse with like proteins to form larger droplets, in much the same way that raindrops merge on a car’s windshield.
Julia F. Riley and Carlos A. Castañeda, Syracuse University
View Media