Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
3688: Brain cells in the hippocampus
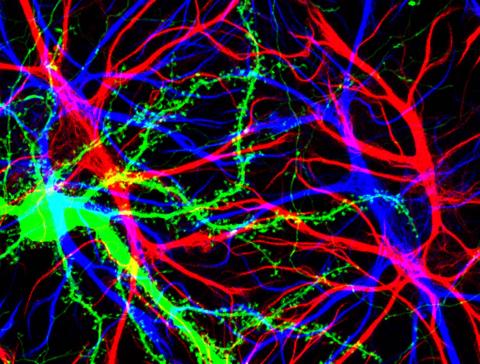
3688: Brain cells in the hippocampus
Hippocampal cells in culture with a neuron in green, showing hundreds of the small protrusions known as dendritic spines. The dendrites of other neurons are labeled in blue, and adjacent glial cells are shown in red.
Shelley Halpain, UC San Diego
View Media
3427: Antitoxin GhoS (Illustration 1)
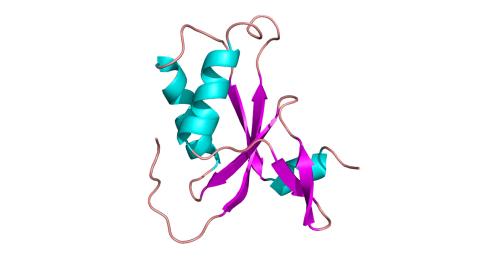
3427: Antitoxin GhoS (Illustration 1)
Structure of the bacterial antitoxin protein GhoS. GhoS inhibits the production of a bacterial toxin, GhoT, which can contribute to antibiotic resistance. GhoS is the first known bacterial antitoxin that works by cleaving the messenger RNA that carries the instructions for making the toxin. More information can be found in the paper: Wang X, Lord DM, Cheng HY, Osbourne DO, Hong SH, Sanchez-Torres V, Quiroga C, Zheng K, Herrmann T, Peti W, Benedik MJ, Page R, Wood TK. A new type V toxin-antitoxin system where mRNA for toxin GhoT is cleaved by antitoxin GhoS. Nat Chem Biol. 2012 Oct;8(10):855-61. Related to 3428.
Rebecca Page and Wolfgang Peti, Brown University and Thomas K. Wood, Pennsylvania State University
View Media
6350: Aldolase
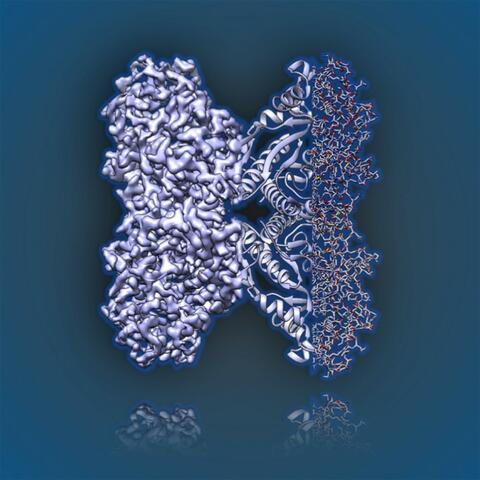
6350: Aldolase
2.5Å resolution reconstruction of rabbit muscle aldolase collected on a FEI/Thermo Fisher Titan Krios with energy filter and image corrector.
National Resource for Automated Molecular Microscopy http://nramm.nysbc.org/nramm-images/ Source: Bridget Carragher
View Media
6897: Zebrafish embryo
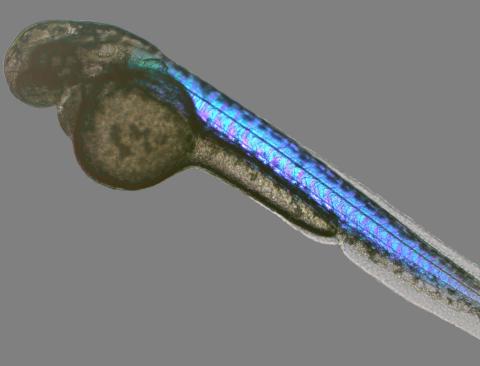
6897: Zebrafish embryo
A zebrafish embryo showing its natural colors. Zebrafish have see-through eggs and embryos, making them ideal research organisms for studying the earliest stages of development. This image was taken in transmitted light under a polychromatic polarizing microscope.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media
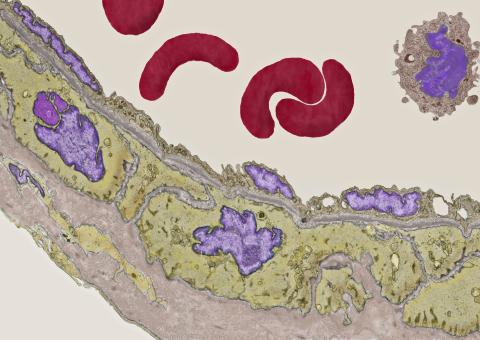
3738: Transmission electron microscopy of coronary artery wall with elastin-rich ECM pseudocolored in light brown
3738: Transmission electron microscopy of coronary artery wall with elastin-rich ECM pseudocolored in light brown
Elastin is a fibrous protein in the extracellular matrix (ECM). It is abundant in artery walls like the one shown here. As its name indicates, elastin confers elasticity. Elastin fibers are at least five times stretchier than rubber bands of the same size. Tissues that expand, such as blood vessels and lungs, need to be both strong and elastic, so they contain both collagen (another ECM protein) and elastin. In this photo, the elastin-rich ECM is colored grayish brown and is most visible at the bottom of the photo. The curved red structures near the top of the image are red blood cells.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
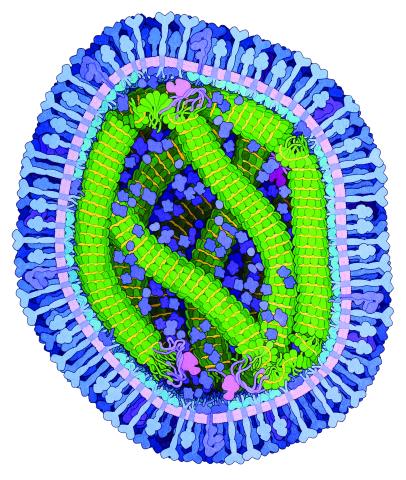
6995: Measles virus
6995: Measles virus
A cross section of the measles virus in which six proteins work together to infect cells. The measles virus is extremely infectious; 9 out of 10 people exposed will contract the disease. Fortunately, an effective vaccine protects against infection.
For a zoomed-in look at the six important proteins, see Measles Virus Proteins.
For a zoomed-in look at the six important proteins, see Measles Virus Proteins.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
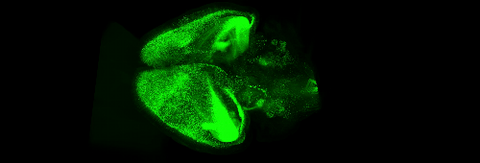
6931: Mouse brain 3
6931: Mouse brain 3
Various views of a mouse brain that was genetically modified so that subpopulations of its neurons glow. Researchers often study mice because they share many genes with people and can shed light on biological processes, development, and diseases in humans.
This video was captured using a light sheet microscope.
Related to images 6929 and 6930.
This video was captured using a light sheet microscope.
Related to images 6929 and 6930.
Prayag Murawala, MDI Biological Laboratory and Hannover Medical School.
View Media
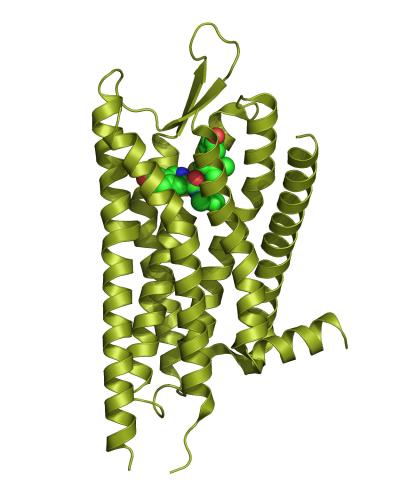
3359: Kappa opioid receptor
3359: Kappa opioid receptor
The receptor is shown bound to an antagonist, JDTic.
Raymond Stevens, The Scripps Research Institute
View Media
6591: Cell-like compartments from frog eggs 4
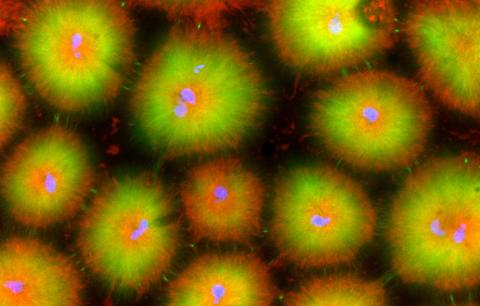
6591: Cell-like compartments from frog eggs 4
Cell-like compartments that spontaneously emerged from scrambled frog eggs, with nuclei (blue) from frog sperm. Endoplasmic reticulum (red) and microtubules (green) are also visible. Image created using confocal microscopy.
For more photos of cell-like compartments from frog eggs view: 6584, 6585, 6586, 6592, and 6593.
For videos of cell-like compartments from frog eggs view: 6587, 6588, 6589, and 6590.
Xianrui Cheng, Stanford University School of Medicine.
View Media
6353: ATP Synthase
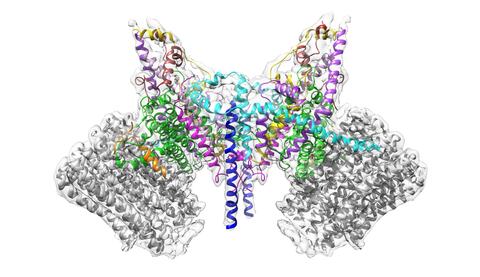
6353: ATP Synthase
Atomic model of the membrane region of the mitochondrial ATP synthase built into a cryo-EM map at 3.6 Å resolution. ATP synthase is the primary producer of ATP in aerobic cells. Drugs that inhibit the bacterial ATP synthase, but not the human mitochondrial enzyme, can serve as antibiotics. This therapeutic approach was successfully demonstrated with the bedaquiline, an ATP synthase inhibitor now used in the treatment of extensively drug resistant tuberculosis.
More information about this structure can be found in the Science paper ”Atomic model for the dimeric F0 region of mitochondrial ATP synthase” by Guo et. al.
More information about this structure can be found in the Science paper ”Atomic model for the dimeric F0 region of mitochondrial ATP synthase” by Guo et. al.
Bridget Carragher, <a href="http://nramm.nysbc.org/">NRAMM National Resource for Automated Molecular Microscopy</a>
View Media
6593: Cell-like compartments from frog eggs 6
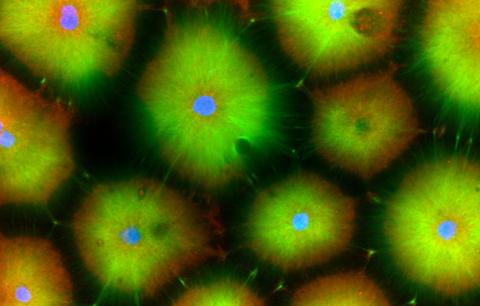
6593: Cell-like compartments from frog eggs 6
Cell-like compartments that spontaneously emerged from scrambled frog eggs, with nuclei (blue) from frog sperm. Endoplasmic reticulum (red) and microtubules (green) are also visible. Image created using confocal microscopy.
For more photos of cell-like compartments from frog eggs view: 6584, 6585, 6586, 6591, 6592.
For videos of cell-like compartments from frog eggs view: 6587, 6588, 6589, and 6590.
Xianrui Cheng, Stanford University School of Medicine.
View Media

3658: Electrostatic map of human spermine synthase
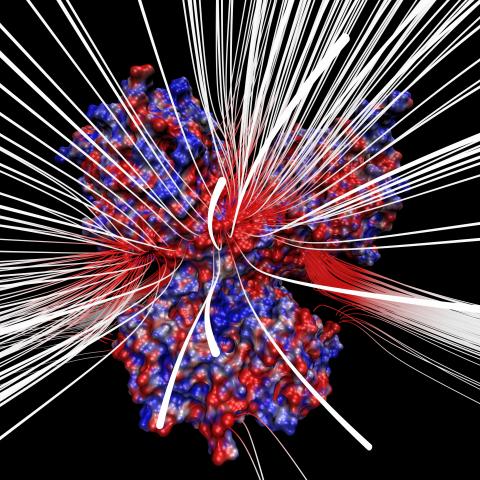
3658: Electrostatic map of human spermine synthase
From PDB entry 3c6k, Crystal structure of human spermine synthase in complex with spermidine and 5-methylthioadenosine.
Emil Alexov, Clemson University
View Media
3590: Fruit fly spermatids
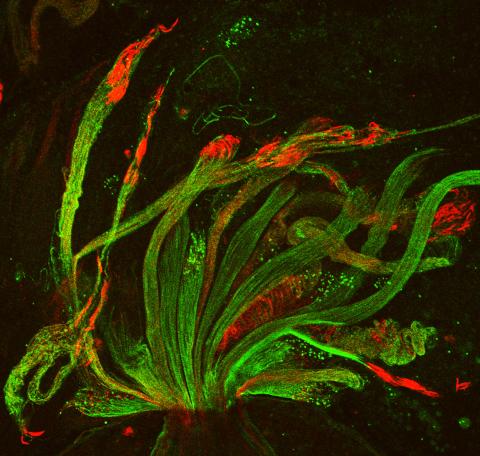
3590: Fruit fly spermatids
Developing spermatids (precursors of mature sperm cells) begin as small, round cells and mature into long-tailed, tadpole-shaped ones. In the sperm cell's head is the cell nucleus; in its tail is the power to outswim thousands of competitors to fertilize an egg. As seen in this microscopy image, fruit fly spermatids start out as groups of interconnected cells. A small lipid molecule called PIP2 helps spermatids tell their heads from their tails. Here, PIP2 (red) marks the nuclei and a cell skeleton-building protein called tubulin (green) marks the tails. When PIP2 levels are too low, some spermatids get mixed up and grow with their heads at the wrong end. Because sperm development is similar across species, studies in fruit flies could help researchers understand male infertility in humans.
Lacramioara Fabian, The Hospital for Sick Children, Toronto, Canada
View Media
2382: PanB from M. tuberculosis (2)
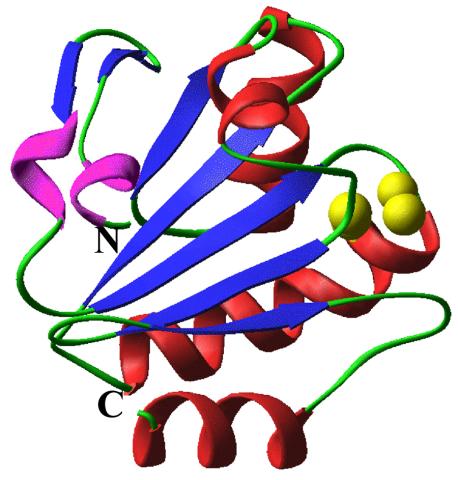
2382: PanB from M. tuberculosis (2)
Model of an enzyme, PanB, from Mycobacterium tuberculosis, the bacterium that causes most cases of tuberculosis. This enzyme is an attractive drug target.
Mycobacterium Tuberculosis Center, PSI-1
View Media
6547: Cell Nucleus and Lipid Droplets
6547: Cell Nucleus and Lipid Droplets
A cell nucleus (blue) surrounded by lipid droplets (yellow). Exogenously expressed, S-tagged UBXD8 (green) recruits endogenous p97/VCP (red) to the surface of lipid droplets in oleate-treated HeLa cells. Nucleus stained with DAPI.
James Olzmann, University of California, Berkeley
View Media

2579: Bottles of warfarin
2579: Bottles of warfarin
In 2007, the FDA modified warfarin's label to indicate that genetic makeup may affect patient response to the drug. The widely used blood thinner is sold under the brand name Coumadin®. Scientists involved in the NIH Pharmacogenetics Research Network are investigating whether genetic information can be used to improve optimal dosage prediction for patients.
Alisa Machalek, NIGMS/NIH
View Media
3481: Bacillus anthracis being killed
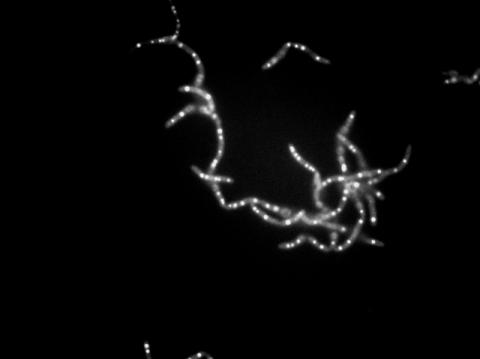
3481: Bacillus anthracis being killed
Bacillus anthracis (anthrax) cells being killed by a fluorescent trans-translation inhibitor, which disrupts bacterial protein synthesis. The inhibitor is naturally fluorescent and looks blue when it is excited by ultraviolet light in the microscope. This is a black-and-white version of Image 3525.
John Alumasa, Keiler Laboratory, Pennsylvania State University
View Media
2341: Aminopeptidase N from N. meningitidis
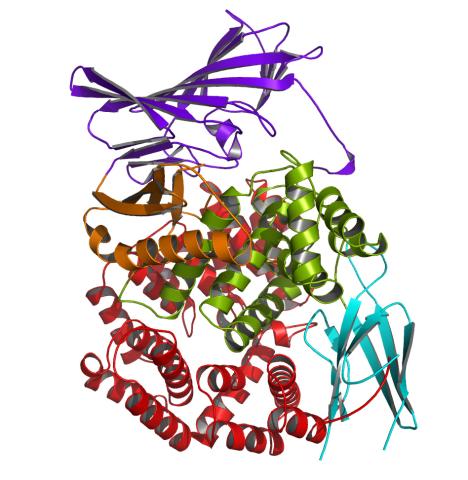
2341: Aminopeptidase N from N. meningitidis
Model of the enzyme aminopeptidase N from the human pathogen Neisseria meningitidis, which can cause meningitis epidemics. The structure provides insight on the active site of this important molecule.
Midwest Center for Structural Genomics, PSI
View Media
3403: Disrupted vascular development in frog embryos
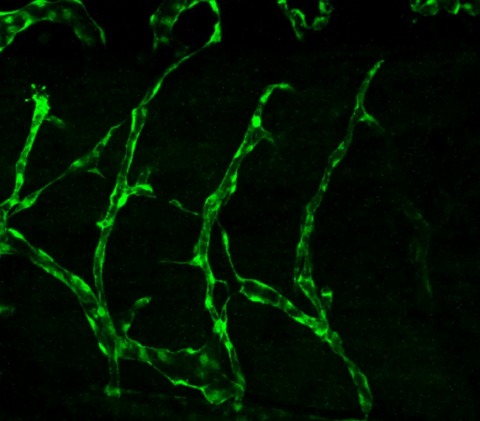
3403: Disrupted vascular development in frog embryos
Disassembly of vasculature in kdr:GFP frogs following addition of 250 µM TBZ. Related to images 3404 and 3505.
Hye Ji Cha, University of Texas at Austin
View Media
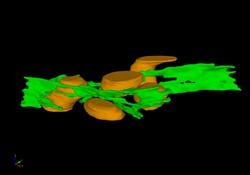
2635: Mitochondria and endoplasmic reticulum
2635: Mitochondria and endoplasmic reticulum
A computer model shows how the endoplasmic reticulum is close to and almost wraps around mitochondria in the cell. The endoplasmic reticulum is lime green and the mitochondria are yellow. This image relates to a July 27, 2009 article in Computing Life.
Bridget Wilson, University of New Mexico
View Media
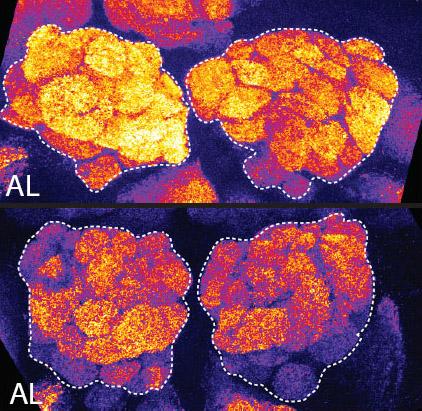
2596: Sleep and the fly brain
2596: Sleep and the fly brain
In the top snapshots, the brain of a sleep-deprived fruit fly glows orange, marking high concentrations of a synaptic protein called Bruchpilot (BRP) involved in communication between neurons. The color particularly lights up brain areas associated with learning. By contrast, the bottom images from a well-rested fly show lower levels of the protein. These pictures illustrate the results of an April 2009 study showing that sleep reduces the protein's levels, suggesting that such "downscaling" resets the brain to normal levels of synaptic activity and makes it ready to learn after a restful night.
Chiara Cirelli, University of Wisconsin-Madison
View Media

2543: DNA replication illustration
2543: DNA replication illustration
During DNA replication, each strand of the original molecule acts as a template for the synthesis of a new, complementary DNA strand. See image 2544 for a labeled version of this illustration.
Crabtree + Company
View Media
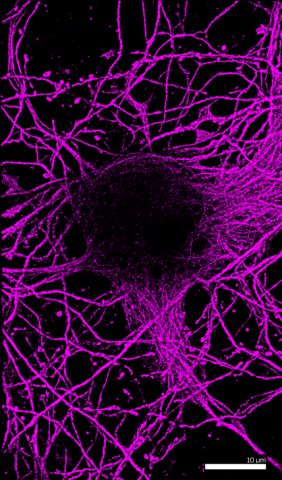
6890: Microtubules in hippocampal neurons
6890: Microtubules in hippocampal neurons
Microtubules (magenta) in neurons of the hippocampus, a part of the brain involved in learning and memory. Microtubules are strong, hollow fibers that provide structural support to cells. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6889, 6891, and 6892.
Related to images 6889, 6891, and 6892.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media
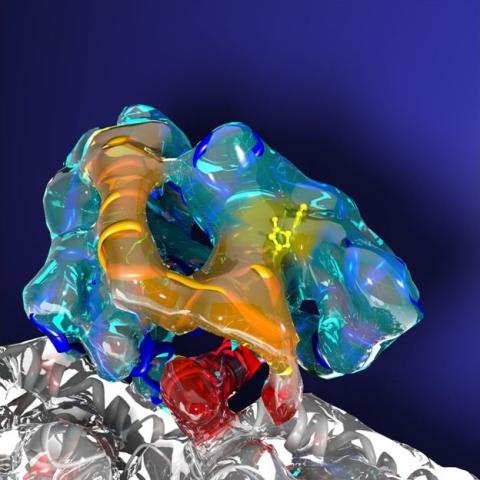
3491: Kinesin moves cellular cargo
3491: Kinesin moves cellular cargo
A protein called kinesin (blue) is in charge of moving cargo around inside cells and helping them divide. It's powered by biological fuel called ATP (bright yellow) as it scoots along tube-like cellular tracks called microtubules (gray).
Charles Sindelar, Yale University
View Media
6553: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 48 hours (photo 1)
6553: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 48 hours (photo 1)
Floral pattern emerging as two bacterial species, motile Acinetobacter baylyi (red) and non-motile Escherichia coli (green), are grown together for 48 hours on 1% agar surface from a small inoculum in the center of a Petri dish.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6550 for a video of this process.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6550 for a video of this process.
L. Xiong et al, eLife 2020;9: e48885
View Media
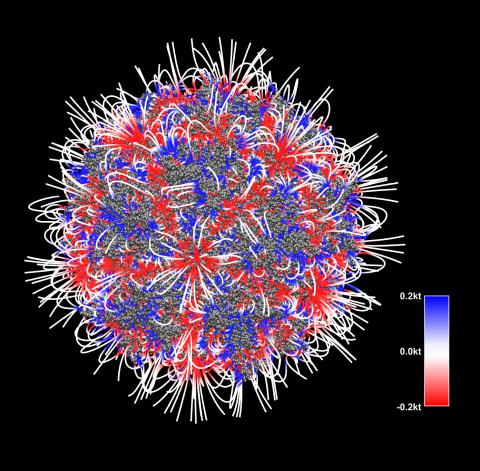
3375: Electrostatic map of the adeno-associated virus with scale
3375: Electrostatic map of the adeno-associated virus with scale
The new highly efficient parallelized DelPhi software was used to calculate the potential map distribution of an entire virus, the adeno-associated virus, which is made up of more than 484,000 atoms. Despite the relatively large dimension of this biological system, resulting in 815x815x815 mesh points, the parallelized DelPhi, utilizing 100 CPUs, completed the calculations within less than three minutes. Related to image 3374.
Emil Alexov, Clemson University
View Media

1287: Mitochondria
1287: Mitochondria
Bean-shaped mitochondria are cells' power plants. These organelles have their own DNA and replicate independently. The highly folded inner membranes are the site of energy generation.
Judith Stoffer
View Media
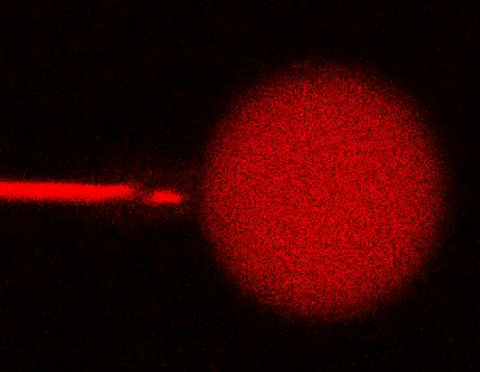
3583: Bee venom toxin destroying a cell
3583: Bee venom toxin destroying a cell
This video condenses 6.5 minutes into less than a minute to show how the toxin in bee venom, called melittin, destroys an animal or bacterial cell. What looks like a red balloon is an artificial cell filled with red dye. Melittin molecules are colored green and float on the cell's surface like twigs on a pond. As melittin accumulates on the cell's membrane, the membrane expands to accommodate it. In the video, the membrane stretches into a column on the left. When melittin levels reach a critical threshold, countless pinhole leaks burst open in the membrane. The cell's vital fluids (red dye in the video) leak out through these pores. Within minutes, the cell collapses.
Huey Huang, Rice University
View Media
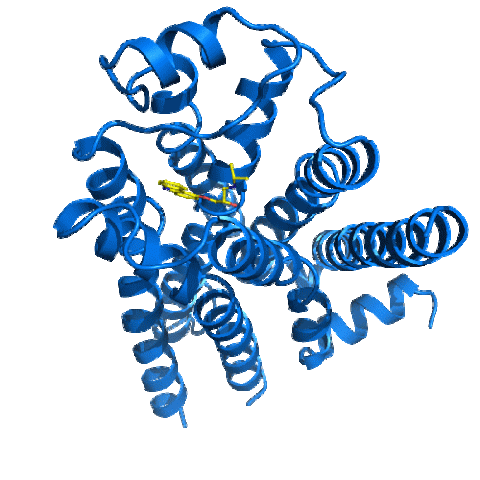
2337: Beta2-adrenergic receptor protein
2337: Beta2-adrenergic receptor protein
Crystal structure of the beta2-adrenergic receptor protein. This is the first known structure of a human G protein-coupled receptor, a large family of proteins that control critical bodily functions and the action of about half of today's pharmaceuticals. Featured as one of the November 2007 Protein Structure Initiative Structures of the Month.
The Stevens Laboratory, The Scripps Research Institute
View Media
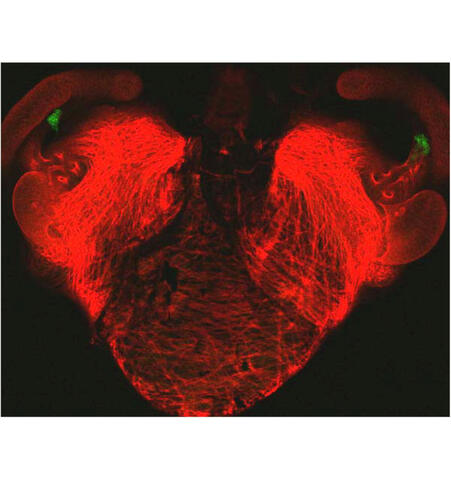
7018: Bacterial cells aggregating above the light organ of the Hawaiian bobtail squid
7018: Bacterial cells aggregating above the light organ of the Hawaiian bobtail squid
A light organ (~0.5 mm across) of a juvenile Hawaiian bobtail squid, Euprymna scolopes. Movement of cilia on the surface of the organ aggregates bacterial symbionts (green) into two areas above sets of pores that lead to interior crypts. This image was taken using a confocal fluorescence microscope.
Related to images 7016, 7017, 7019, and 7020.
Related to images 7016, 7017, 7019, and 7020.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
3288: Smooth muscle from human ES cells
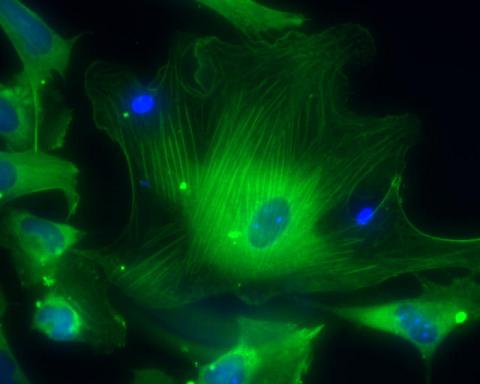
3288: Smooth muscle from human ES cells
These smooth muscle cells were derived from human embryonic stem cells. The nuclei are stained blue, and the proteins of the cytoskeleton are stained green. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Alexey Terskikh lab, Burnham Institute for Medical Research, via CIRM
View Media

2723: iPS cell facility at the Coriell Institute for Medical Research
2723: iPS cell facility at the Coriell Institute for Medical Research
This lab space was designed for work on the induced pluripotent stem (iPS) cell collection, part of the NIGMS Human Genetic Cell Repository at the Coriell Institute for Medical Research.
Courtney Sill, Coriell Institute for Medical Research
View Media
5816: Cas9 protein involved in the CRISPR gene-editing technology
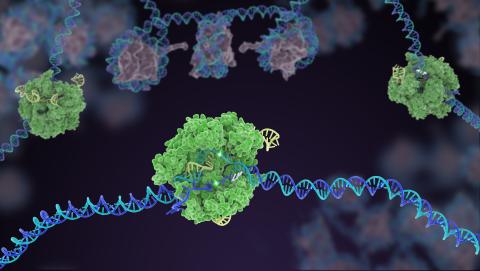
5816: Cas9 protein involved in the CRISPR gene-editing technology
In the gene-editing tool CRISPR, a small strand of RNA identifies a specific chunk of DNA. Then the enzyme Cas9 (green) swoops in and cuts the double-stranded DNA (blue/purple) in two places, removing the specific chunk.
Janet Iwasa
View Media
3765: Trypanosoma brucei, the cause of sleeping sickness
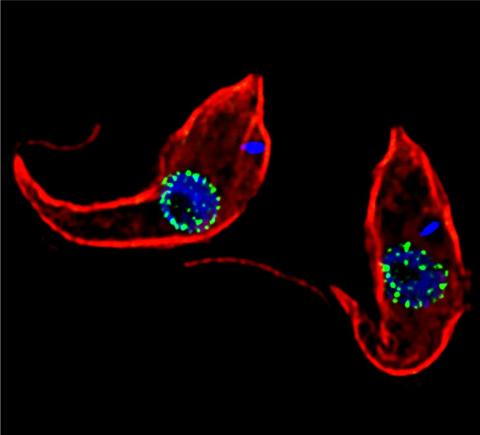
3765: Trypanosoma brucei, the cause of sleeping sickness
Trypanosoma brucei is a single-cell parasite that causes sleeping sickness in humans. Scientists have been studying trypanosomes for some time because of their negative effects on human and also animal health, especially in sub-Saharan Africa. Moreover, because these organisms evolved on a separate path from those of animals and plants more than a billion years ago, researchers study trypanosomes to find out what traits they may harbor that are common to or different from those of other eukaryotes (i.e., those organisms having a nucleus and mitochondria). This image shows the T. brucei cell membrane in red, the DNA in the nucleus and kinetoplast (a structure unique to protozoans, including trypanosomes, which contains mitochondrial DNA) in blue and nuclear pore complexes (which allow molecules to pass into or out of the nucleus) in green. Scientists have found that the trypanosome nuclear pore complex has a unique mechanism by which it attaches to the nuclear envelope. In addition, the trypanosome nuclear pore complex differs from those of other eukaryotes because its components have a near-complete symmetry, and it lacks almost all of the proteins that in other eukaryotes studied so far are required to assemble the pore.
Michael Rout, Rockefeller University
View Media
2473: Glowing glycans
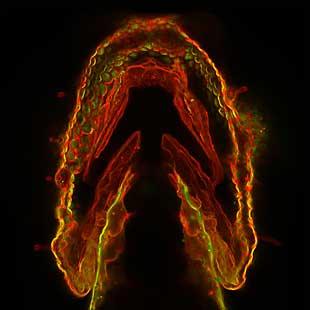
2473: Glowing glycans
Sugars light up the cells in this jaw of a 3-day-old zebrafish embryo and highlight a scientific first: labeling and tracking the movements of sugar chains called glycans in a living organism. Here, recently produced glycans (red) are on the cell surface while those made earlier in development (green) have migrated into the cells. In some areas, old and new glycans mingle (yellow). A better understanding of such traffic patterns could shed light on how organisms develop and may uncover markers for disease, such as cancer. Featured in the May 21, 2008 of Biomedical Beat.
Carolyn Bertozzi, University of California, Berkeley
View Media
2756: Xenopus laevis embryos
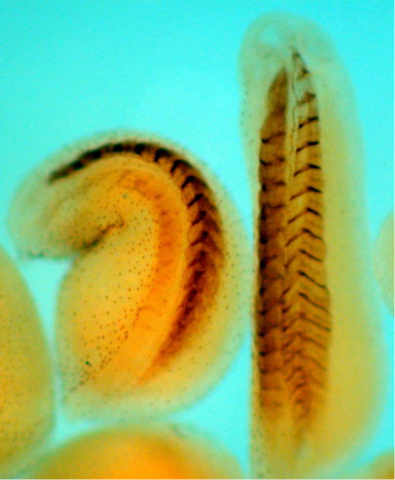
2756: Xenopus laevis embryos
Xenopus laevis, the African clawed frog, has long been used as a model organism for studying embryonic development. The frog embryo on the left lacks the developmental factor Sizzled. A normal embryo is shown on the right.
Michael Klymkowsky, University of Colorado, Boulder
View Media
2324: Movements of myosin
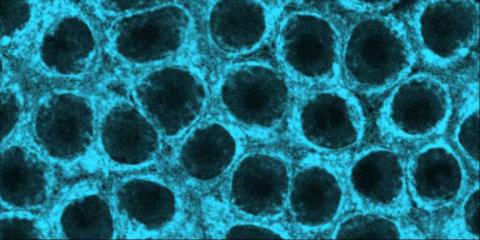
2324: Movements of myosin
Inside the fertilized egg cell of a fruit fly, we see a type of myosin (related to the protein that helps muscles contract) made to glow by attaching a fluorescent protein. After fertilization, the myosin proteins are distributed relatively evenly near the surface of the embryo. The proteins temporarily vanish each time the cells' nuclei--initially buried deep in the cytoplasm--divide. When the multiplying nuclei move to the surface, they shift the myosin, producing darkened holes. The glowing myosin proteins then gather, contract, and start separating the nuclei into their own compartments.
Victoria Foe, University of Washington
View Media
3402: Hsp33 Heat Shock Protein Inactive to Active
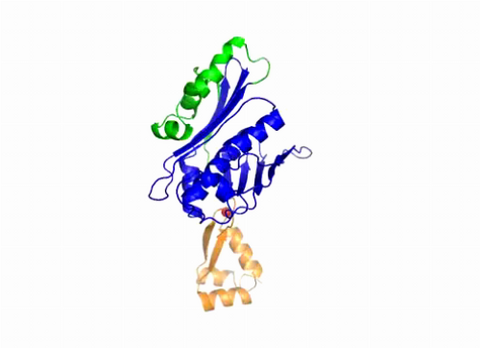
3402: Hsp33 Heat Shock Protein Inactive to Active
When the heat shock protein hsp33 is folded, it is inactive and contains a zinc ion, stabilizing the redox sensitive domain (orange). In the presence of an environmental stressor, the protein releases the zinc ion, which leads to the unfolding of the redox domain. This unfolding causes the chaperone to activate by reaching out its "arm" (green) to protect other proteins.
Dana Reichmann, University of Michigan
View Media
2331: Statistical cartography
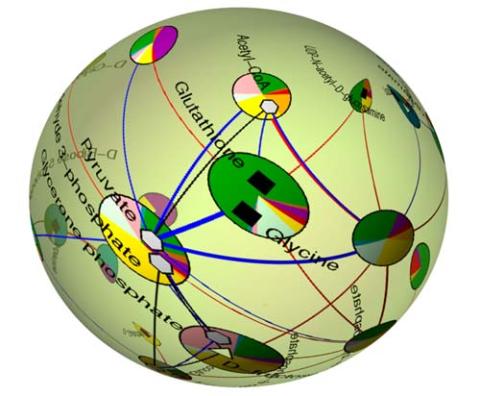
2331: Statistical cartography
Like a world of its own, this sphere represents all the known chemical reactions in the E. coli bacterium. The colorful circles on the surface symbolize sets of densely interconnected reactions. The lines between the circles show additional connecting reactions. The shapes inside the circles are landmark molecules, like capital cities on a map, that either act as hubs for many groups of reactions, are highly conserved among species, or both. Molecules that connect far-flung reactions on the sphere are much more conserved during evolution than molecules that connect reactions within a single circle. This statistical cartography could reveal insights about other complex systems, such as protein interactions and gene regulation networks.
Luis A. Nunes Amaral, Northwestern University
View Media
6993: RNA polymerase
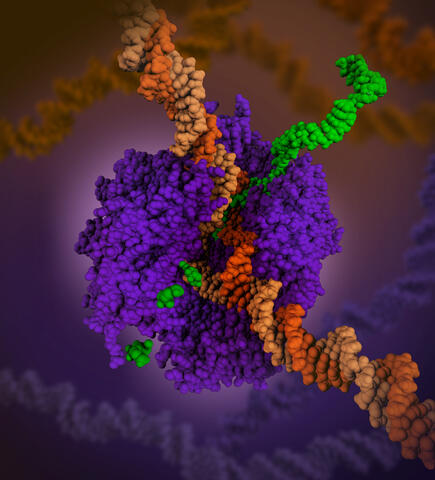
6993: RNA polymerase
RNA polymerase (purple) is a complex enzyme at the heart of transcription. During this process, the enzyme unwinds the DNA double helix and uses one strand (darker orange) as a template to create the single-stranded messenger RNA (green), later used by ribosomes for protein synthesis.
From the RNA polymerase II elongation complex of Saccharomyces cerevisiae (PDB entry 1I6H) as seen in PDB-101's What is a Protein? video.
From the RNA polymerase II elongation complex of Saccharomyces cerevisiae (PDB entry 1I6H) as seen in PDB-101's What is a Protein? video.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
2567: Haplotypes (with labels)
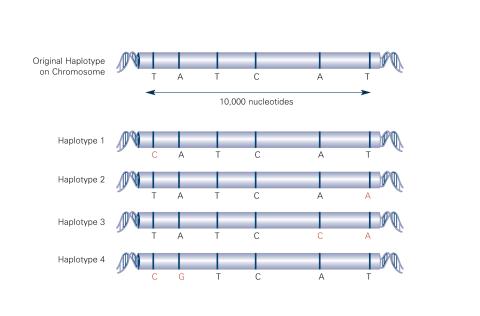
2567: Haplotypes (with labels)
Haplotypes are combinations of gene variants that are likely to be inherited together within the same chromosomal region. In this example, an original haplotype (top) evolved over time to create three newer haplotypes that each differ by a few nucleotides (red). See image 2566 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
6562: Drosophila (fruit fly) myosin 1D motility assay
6562: Drosophila (fruit fly) myosin 1D motility assay
Actin gliding powered by myosin 1D. Note the counterclockwise motion of the gliding actin filaments.
Serapion Pyrpassopoulos and E. Michael Ostap, University of Pennsylvania
View Media
3766: TFIID complex binds DNA to start gene transcription
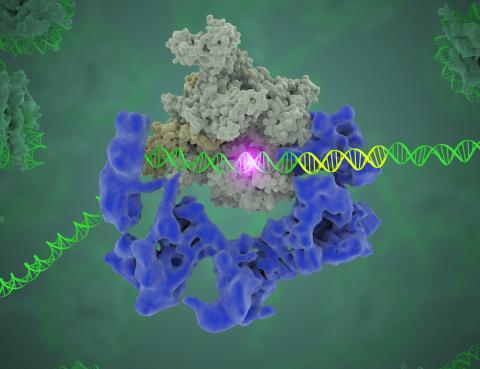
3766: TFIID complex binds DNA to start gene transcription
Gene transcription is a process by which the genetic information encoded in DNA is transcribed into RNA. It's essential for all life and requires the activity of proteins, called transcription factors, that detect where in a DNA strand transcription should start. In eukaryotes (i.e., those that have a nucleus and mitochondria), a protein complex comprising 14 different proteins is responsible for sniffing out transcription start sites and starting the process. This complex, called TFIID, represents the core machinery to which an enzyme, named RNA polymerase, can bind to and read the DNA and transcribe it to RNA. Scientists have used cryo-electron microscopy (cryo-EM) to visualize the TFIID-RNA polymerase-DNA complex in unprecedented detail. In this illustration, TFIID (blue) contacts the DNA and recruits the RNA polymerase (gray) for gene transcription. The start of the transcribed gene is shown with a flash of light. To learn more about the research that has shed new light on gene transcription, see this news release from Berkeley Lab. Related to video 5730.
Eva Nogales, Berkeley Lab
View Media
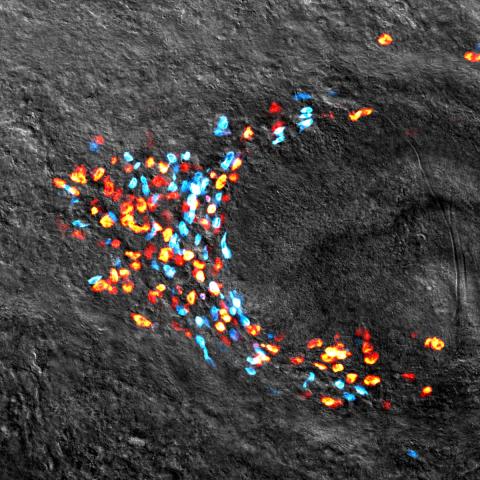
3306: Planarian stem cell colony
3306: Planarian stem cell colony
Planarians are freshwater flatworms that have powerful abilities to regenerate their bodies, which would seem to make them natural model organisms in which to study stem cells. But until recently, scientists had not been able to efficiently find the genes that regulate the planarian stem cell system. In this image, a single stem cell has given rise to a colony of stem cells in a planarian. Proliferating cells are red, and differentiating cells are blue. Quantitatively measuring the size and ratios of these two cell types provides a powerful framework for studying the roles of stem cell regulatory genes in planarians.
Peter Reddien, Whitehead Institute
View Media

1120: Superconducting magnet
1120: Superconducting magnet
Superconducting magnet for NMR research, from the February 2003 profile of Dorothee Kern in Findings.
Mike Lovett
View Media
5762: Panorama view of golden mitochondria
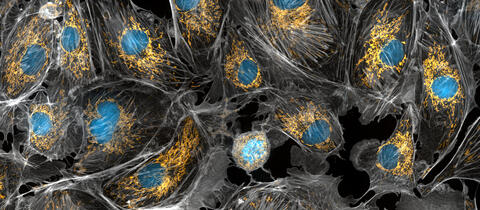
5762: Panorama view of golden mitochondria
Mitochondria are the powerhouses of the cells, generating the energy the cells need to do their tasks and to stay alive. Researchers have studied mitochondria for some time because when these cell organelles don't work as well as they should, several diseases develop. In this photograph of cow cells taken with a microscope, the mitochondria were stained in bright yellow to visualize them in the cell. The large blue dots are the cell nuclei and the gray web is the cytoskeleton of the cells.
Torsten Wittmann, University of California, San Francisco
View Media
5838: Color coding of the Drosophila brain - image
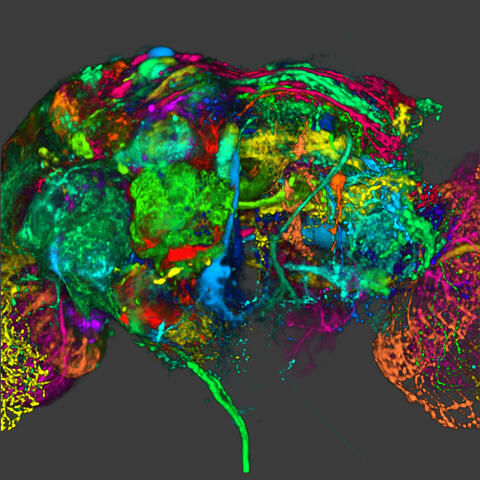
5838: Color coding of the Drosophila brain - image
This image results from a research project to visualize which regions of the adult fruit fly (Drosophila) brain derive from each neural stem cell. First, researchers collected several thousand fruit fly larvae and fluorescently stained a random stem cell in the brain of each. The idea was to create a population of larvae in which each of the 100 or so neural stem cells was labeled at least once. When the larvae grew to adults, the researchers examined the flies’ brains using confocal microscopy. With this technique, the part of a fly’s brain that derived from a single, labeled stem cell “lights up. The scientists photographed each brain and digitally colorized its lit-up area. By combining thousands of such photos, they created a three-dimensional, color-coded map that shows which part of the Drosophila brain comes from each of its ~100 neural stem cells. In other words, each colored region shows which neurons are the progeny or “clones” of a single stem cell. This work established a hierarchical structure as well as nomenclature for the neurons in the Drosophila brain. Further research will relate functions to structures of the brain.
Related to image 5868 and video 5843
Related to image 5868 and video 5843
Yong Wan from Charles Hansen’s lab, University of Utah. Data preparation and visualization by Masayoshi Ito in the lab of Kei Ito, University of Tokyo.
View Media
1011: Lily mitosis 11
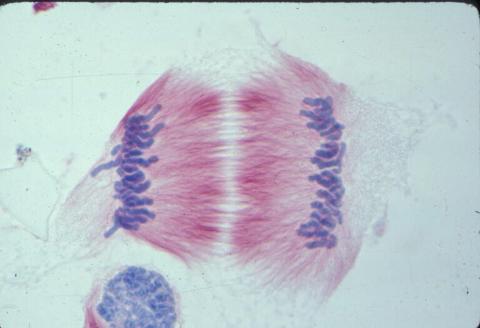
1011: Lily mitosis 11
A light microscope image of cells from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and have separated into the opposite sides of a dividing cell.
Related to images 1010, 1012, 1013, 1014, 1015, 1016, 1017, 1018, 1019, and 1021.
Related to images 1010, 1012, 1013, 1014, 1015, 1016, 1017, 1018, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
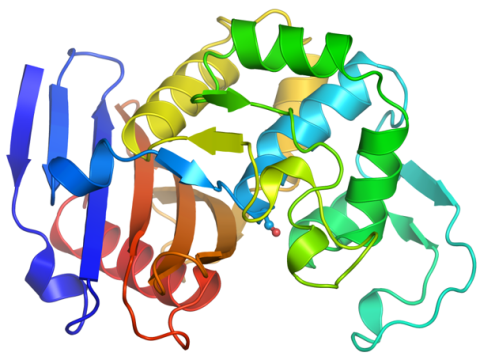
6766: Ribbon diagram of a cefotaxime-CCD-1 complex
6766: Ribbon diagram of a cefotaxime-CCD-1 complex
CCD-1 is an enzyme produced by the bacterium Clostridioides difficile that helps it resist antibiotics. Using X-ray crystallography, researchers determined the structure of a CCD-1 molecule and a molecule of the antibiotic cefotaxime bound together. The structure revealed that CCD-1 provides extensive hydrogen bonding and stabilization of the antibiotic in the active site, leading to efficient degradation of the antibiotic.
Related to images 6764, 6765, and 6767.
Related to images 6764, 6765, and 6767.
Keith Hodgson, Stanford University.
View Media
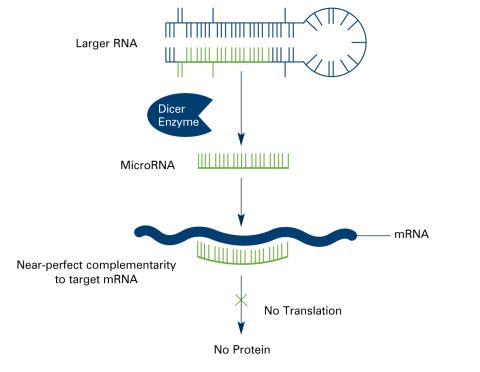
2557: Dicer generates microRNAs (with labels)
2557: Dicer generates microRNAs (with labels)
The enzyme Dicer generates microRNAs by chopping larger RNA molecules into tiny Velcro®-like pieces. MicroRNAs stick to mRNA molecules and prevent the mRNAs from being made into proteins. See image 2556 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
