Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
2511: X-ray crystallography
2511: X-ray crystallography
X-ray crystallography allows researchers to see structures too small to be seen by even the most powerful microscopes. To visualize the arrangement of atoms within molecules, researchers can use the diffraction patterns obtained by passing X-ray beams through crystals of the molecule. This is a common way for solving the structures of proteins. See image 2512 for a labeled version of this illustration. Featured in The Structures of Life.
Crabtree + Company
View Media
3396: Myelinated axons 1
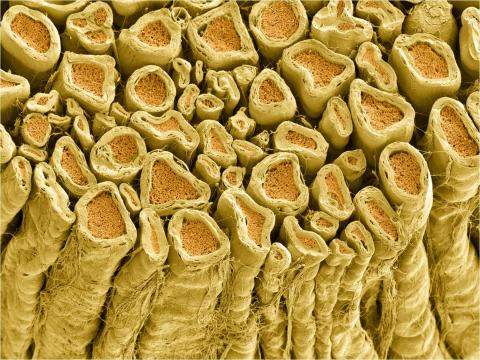
3396: Myelinated axons 1
Myelinated axons in a rat spinal root. Myelin is a type of fat that forms a sheath around and thus insulates the axon to protect it from losing the electrical current needed to transmit signals along the axon. The axoplasm inside the axon is shown in pink. Related to 3397.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media

7012: Adult Hawaiian bobtail squid burying in the sand
7012: Adult Hawaiian bobtail squid burying in the sand
Each morning, the nocturnal Hawaiian bobtail squid, Euprymna scolopes, hides from predators by digging into the sand. At dusk, it leaves the sand again to hunt.
Related to image 7010 and 7011.
Related to image 7010 and 7011.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media

2398: RNase A (1)
2398: RNase A (1)
A crystal of RNase A protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
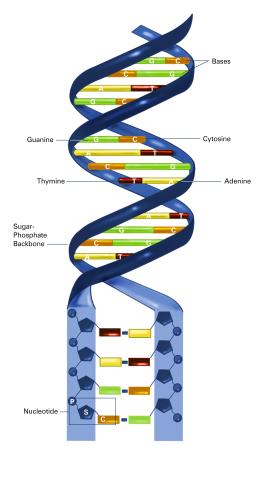
2542: Nucleotides make up DNA (with labels)
2542: Nucleotides make up DNA (with labels)
DNA consists of two long, twisted chains made up of nucleotides. Each nucleotide contains one base, one phosphate molecule, and the sugar molecule deoxyribose. The bases in DNA nucleotides are adenine, thymine, cytosine, and guanine. See image 2541 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

6764: Crystals of CCD-1 in complex with cefotaxime
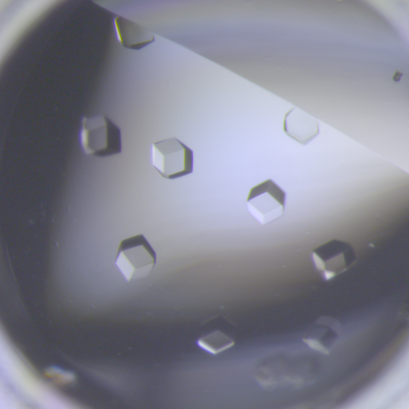
6764: Crystals of CCD-1 in complex with cefotaxime
CCD-1 is an enzyme produced by the bacterium Clostridioides difficile that helps it resist antibiotics. Here, researchers crystallized bound pairs of CCD-1 molecules and molecules of the antibiotic cefotaxime. This enabled their structure to be studied using X-ray crystallography.
Related to images 6765, 6766, and 6767.
Related to images 6765, 6766, and 6767.
Keith Hodgson, Stanford University.
View Media
2434: Fruit fly retina 02
2434: Fruit fly retina 02
Section of a fruit fly retina showing the light-sensing molecules rhodopsin-5 (blue) and rhodopsin-6 (red).
Hermann Steller, Rockefeller University
View Media

7010: Adult and juvenile Hawaiian bobtail squids
7010: Adult and juvenile Hawaiian bobtail squids
An adult Hawaiian bobtail squid, Euprymna scolopes, (~4 cm) surrounded by newly hatched juveniles (~2 mm) in a bowl of seawater.
Related to image 7011 and video 7012.
Related to image 7011 and video 7012.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
3295: Cluster analysis of mysterious protein
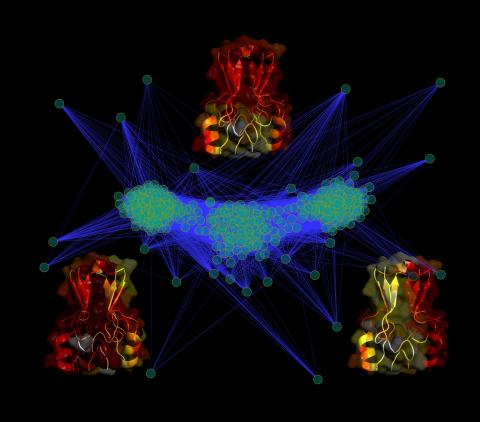
3295: Cluster analysis of mysterious protein
Researchers use cluster analysis to study protein shape and function. Each green circle represents one potential shape of the protein mitoNEET. The longer the blue line between two circles, the greater the differences between the shapes. Most shapes are similar; they fall into three clusters that are represented by the three images of the protein. From a Rice University news release. Graduate student Elizabeth Baxter and Patricia Jennings, professor of chemistry and biochemistry at UCSD, collaborated with José Onuchic, a physicist at Rice University, on this work.
Patricia Jennings and Elizabeth Baxter, University of California, San Diego
View Media
2323: Motion in the brain
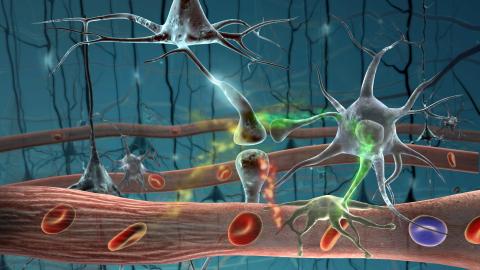
2323: Motion in the brain
Amid a network of blood vessels and star-shaped support cells, neurons in the brain signal each other. The mists of color show the flow of important molecules like glucose and oxygen. This image is a snapshot from a 52-second simulation created by an animation artist. Such visualizations make biological processes more accessible and easier to understand.
Kim Hager and Neal Prakash, University of California, Los Angeles
View Media
2489: Immune cell attacks cell infected with a retrovirus
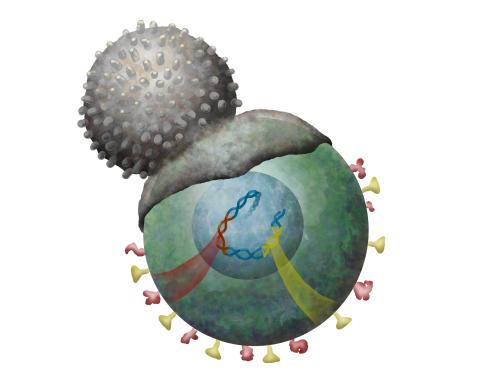
2489: Immune cell attacks cell infected with a retrovirus
T cells engulf and digest cells displaying markers (or antigens) for retroviruses, such as HIV.
Kristy Whitehouse, science illustrator
View Media
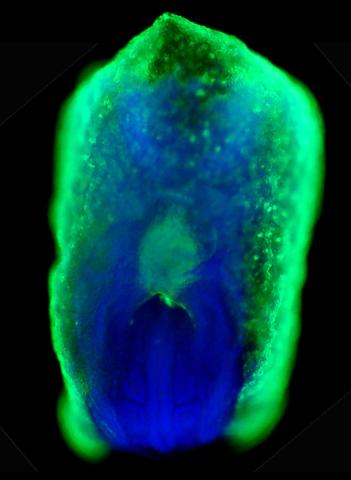
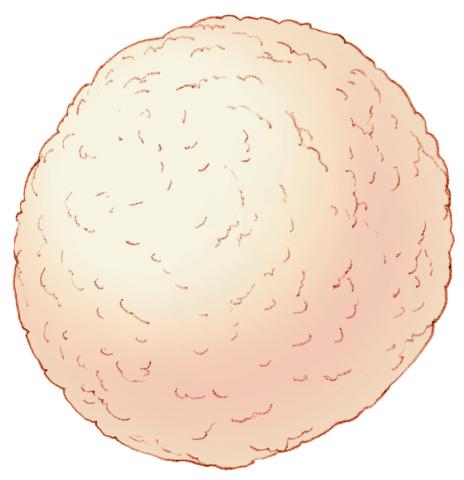
2607: Mouse embryo showing Smad4 protein
2607: Mouse embryo showing Smad4 protein
This eerily glowing blob isn't an alien or a creature from the deep sea--it's a mouse embryo just eight and a half days old. The green shell and core show a protein called Smad4. In the center, Smad4 is telling certain cells to begin forming the mouse's liver and pancreas. Researchers identified a trio of signaling pathways that help switch on Smad4-making genes, starting immature cells on the path to becoming organs. The research could help biologists learn how to grow human liver and pancreas tissue for research, drug testing and regenerative medicine. In addition to NIGMS, NIH's National Institute of Diabetes and Digestive and Kidney Diseases also supported this work.
Kenneth Zaret, Fox Chase Cancer Center
View Media
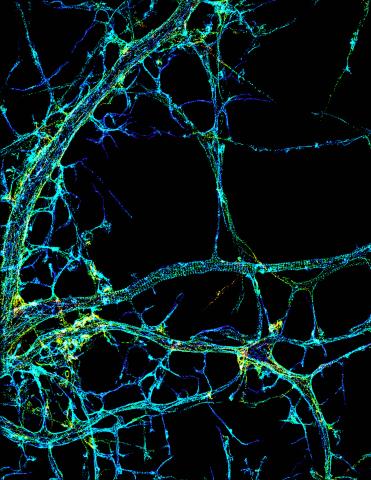
3678: STORM image of axonal cytoskeleton
3678: STORM image of axonal cytoskeleton
This image shows the long, branched structures (axons) of nerve cells. Running horizontally across the middle of the photo is an axon wrapped in rings made of actin protein (green), which plays important roles in nerve cells. The image was captured with a powerful microscopy technique that allows scientists to see single molecules in living cells in real time. The technique is called stochastic optical reconstruction microscopy (STORM). It is based on technology so revolutionary that its developers earned the 2014 Nobel Prize in Chemistry. More information about this image can be found in: K. Xu, G. Zhong, X. Zhuang. Actin, spectrin and associated proteins form a periodic cytoskeleton structure in axons. Science 339, 452-456 (2013).
Xiaowei Zhuang Laboratory, Howard Hughes Medical Institute, Harvard University
View Media
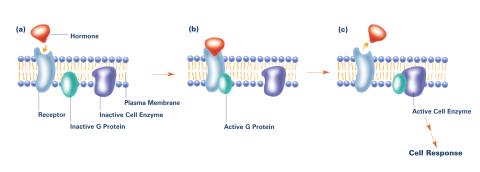
2538: G switch (with labels and stages)
2538: G switch (with labels and stages)
The G switch allows our bodies to respond rapidly to hormones. G proteins act like relay batons to pass messages from circulating hormones into cells. A hormone (red) encounters a receptor (blue) in the membrane of a cell. Next, a G protein (green) becomes activated and makes contact with the receptor to which the hormone is attached. Finally, the G protein passes the hormone's message to the cell by switching on a cell enzyme (purple) that triggers a response. See image 2536 and 2537 for other versions of this image. Featured in Medicines By Design.
Crabtree + Company
View Media
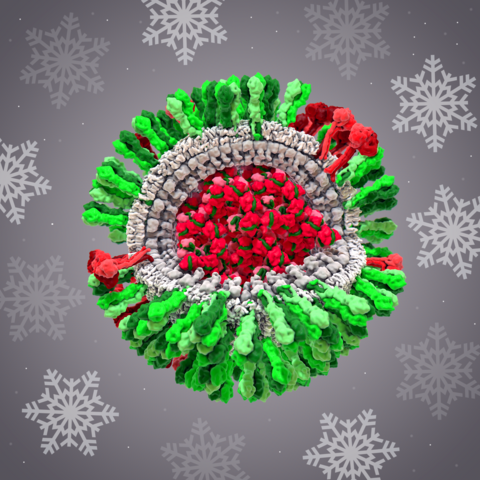

6355: H1N1 Influenza Virus
6355: H1N1 Influenza Virus
CellPack image of the H1N1 influenza virus, with hemagglutinin and neuraminidase glycoproteins in green and red, respectively, on the outer envelope (white); matrix protein in gray, and ribonucleoprotein particles inside the virus in red and green. Related to image 6356.
Dr. Rommie Amaro, University of California, San Diego
View Media
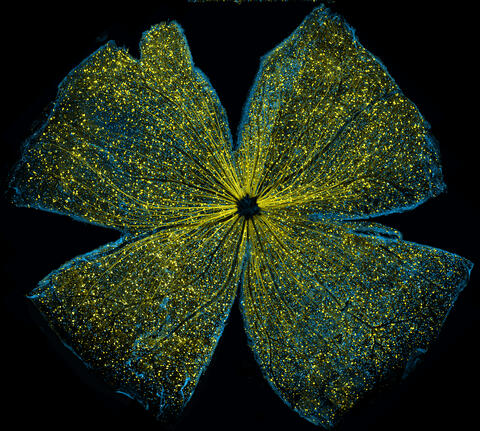
5793: Mouse retina
5793: Mouse retina
What looks like the gossamer wings of a butterfly is actually the retina of a mouse, delicately snipped to lay flat and sparkling with fluorescent molecules. The image is from a research project investigating the promise of gene therapy for glaucoma. It was created at an NIGMS-funded advanced microscopy facility that develops technology for imaging across many scales, from whole organisms to cells to individual molecules.
The ability to obtain high-resolution imaging of tissue as large as whole mouse retinas was made possible by a technique called large-scale mosaic confocal microscopy, which was pioneered by the NIGMS-funded National Center for Microscopy and Imaging Research. The technique is similar to Google Earth in that it computationally stitches together many small, high-resolution images.
The ability to obtain high-resolution imaging of tissue as large as whole mouse retinas was made possible by a technique called large-scale mosaic confocal microscopy, which was pioneered by the NIGMS-funded National Center for Microscopy and Imaging Research. The technique is similar to Google Earth in that it computationally stitches together many small, high-resolution images.
Tom Deerinck and Keunyoung (“Christine”) Kim, NCMIR
View Media
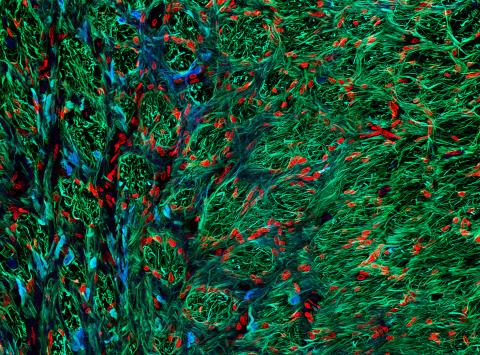
5852: Optic nerve astrocytes
5852: Optic nerve astrocytes
Astrocytes in the cross section of a human optic nerve head
Tom Deerinck and Keunyoung (“Christine”) Kim, NCMIR
View Media
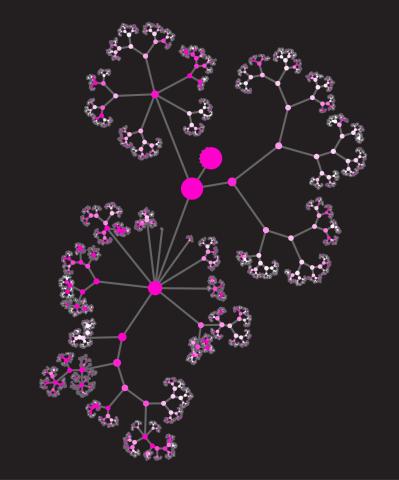
3436: Network diagram of genes, cellular components and processes (unlabeled)
3436: Network diagram of genes, cellular components and processes (unlabeled)
This image shows the hierarchical ontology of genes, cellular components and processes derived from large genomic datasets. From Dutkowski et al. A gene ontology inferred from molecular networks Nat Biotechnol. 2013 Jan;31(1):38-45. Related to 3437.
Janusz Dutkowski and Trey Ideker
View Media
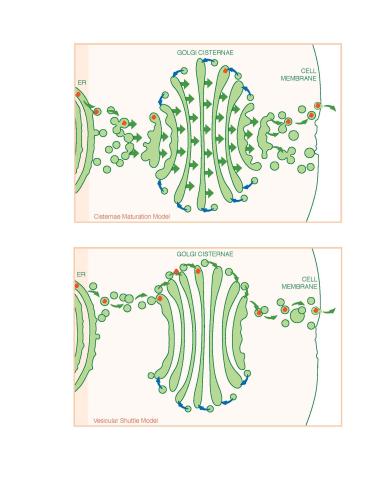
1278: Golgi theories
1278: Golgi theories
Two models for how material passes through the Golgi apparatus: the vesicular shuttle model and the cisternae maturation model.
Judith Stoffer
View Media
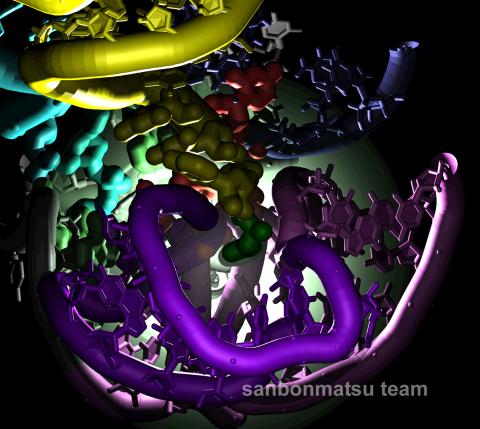
2336: Natural nanomachine in action
2336: Natural nanomachine in action
Using a supercomputer to simulate the movement of atoms in a ribosome, researchers looked into the core of this protein-making nanomachine and took snapshots. The picture shows an amino acid (green) being delivered by transfer RNA (yellow) into a corridor (purple) in the ribosome. In the corridor, a series of chemical reactions will string together amino acids to make a protein. The research project, which tracked the movement of more than 2.6 million atoms, was the largest computer simulation of a biological structure to date. The results shed light on the manufacturing of proteins and could aid the search for new antibiotics, which typically work by disabling the ribosomes of bacteria.
Kevin Sanbonmatsu, Los Alamos National Laboratory
View Media

6344: Drosophila
6344: Drosophila
Two adult fruit flies (Drosophila)
Dr. Vicki Losick, MDI Biological Laboratory, www.mdibl.org
View Media
3573: Myotonic dystrophy type 2 genetic defect
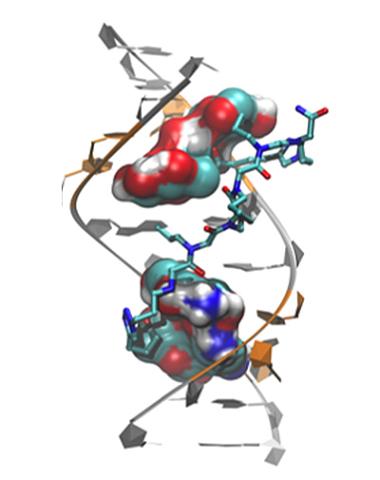
3573: Myotonic dystrophy type 2 genetic defect
Scientists revealed a detailed image of the genetic defect that causes myotonic dystrophy type 2 and used that information to design drug candidates to counteract the disease.
Matthew Disney, Scripps Research Institute and Ilyas Yildirim, Northwestern University
View Media
2574: Simulation of uncontrolled avian flu outbreak
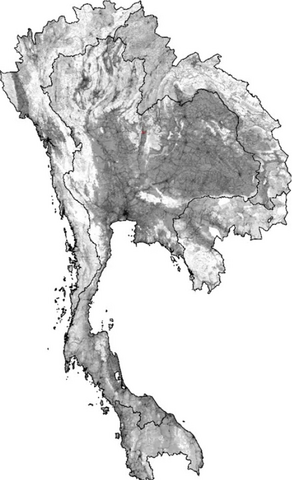
2574: Simulation of uncontrolled avian flu outbreak
This video simulation shows what an uncontrolled outbreak of transmissible avian flu among people living in Thailand might look like. Red indicates new cases while green indicates areas where the epidemic has finished. The video shows the spread of infection and recovery over 300 days in Thailand and neighboring countries.
Neil M. Ferguson, Imperial College London
View Media

6751: Petri dish containing C. elegans
6751: Petri dish containing C. elegans
This Petri dish contains microscopic roundworms called Caenorhabditis elegans. Researchers used these particular worms to study how C. elegans senses the color of light in its environment.
H. Robert Horvitz and Dipon Ghosh, Massachusetts Institute of Technology.
View Media
2793: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 04
2793: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 04
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
3540: Structure of heme, side view
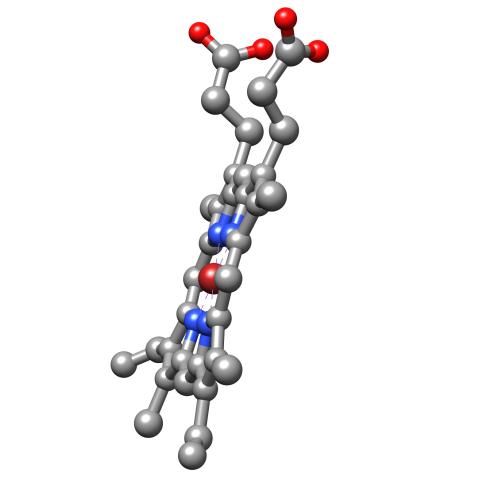
3540: Structure of heme, side view
Molecular model of the struture of heme. Heme is a small, flat molecule with an iron ion (dark red) at its center. Heme is an essential component of hemoglobin, the protein in blood that carries oxygen throughout our bodies. This image first appeared in the September 2013 issue of Findings Magazine. View side view of heme here 3539.
Rachel Kramer Green, RCSB Protein Data Bank
View Media
6933: Zebrafish head vasculature video
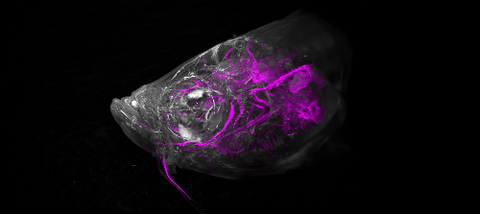
6933: Zebrafish head vasculature video
Various views of a zebrafish head with blood vessels shown in purple. Researchers often study zebrafish because they share many genes with humans, grow and reproduce quickly, and have see-through eggs and embryos, which make it easy to study early stages of development.
This video was captured using a light sheet microscope.
Related to image 6934.
This video was captured using a light sheet microscope.
Related to image 6934.
Prayag Murawala, MDI Biological Laboratory and Hannover Medical School.
View Media
3427: Antitoxin GhoS (Illustration 1)
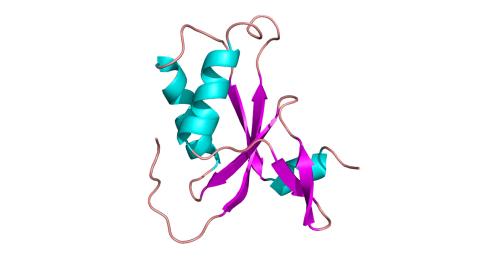
3427: Antitoxin GhoS (Illustration 1)
Structure of the bacterial antitoxin protein GhoS. GhoS inhibits the production of a bacterial toxin, GhoT, which can contribute to antibiotic resistance. GhoS is the first known bacterial antitoxin that works by cleaving the messenger RNA that carries the instructions for making the toxin. More information can be found in the paper: Wang X, Lord DM, Cheng HY, Osbourne DO, Hong SH, Sanchez-Torres V, Quiroga C, Zheng K, Herrmann T, Peti W, Benedik MJ, Page R, Wood TK. A new type V toxin-antitoxin system where mRNA for toxin GhoT is cleaved by antitoxin GhoS. Nat Chem Biol. 2012 Oct;8(10):855-61. Related to 3428.
Rebecca Page and Wolfgang Peti, Brown University and Thomas K. Wood, Pennsylvania State University
View Media
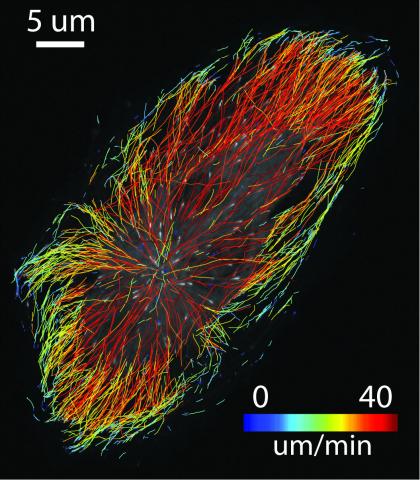
2784: Microtubule dynamics in real time
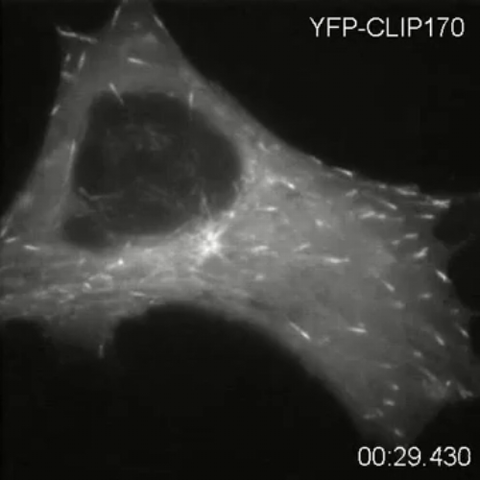
2784: Microtubule dynamics in real time
Cytoplasmic linker protein (CLIP)-170 is a microtubule plus-end-tracking protein that regulates microtubule dynamics and links microtubule ends to different intracellular structures. In this movie, the gene for CLIP-170 has been fused with green fluorescent protein (GFP). When the protein is expressed in cells, the activities can be monitored in real time. Here, you can see CLIP-170 streaming towards the edges of the cell.
Gary Borisy, Marine Biology Laboratory
View Media
6797: Yeast cells with accumulated cell wall material
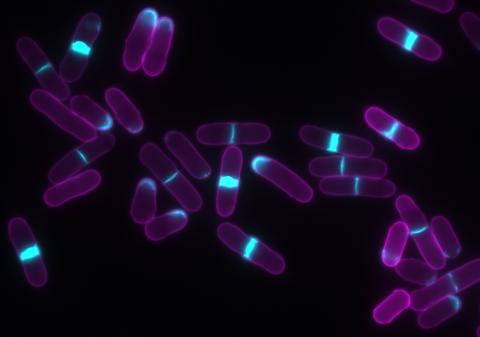
6797: Yeast cells with accumulated cell wall material
Yeast cells that abnormally accumulate cell wall material (blue) at their ends and, when preparing to divide, in their middles. This image was captured using wide-field microscopy with deconvolution.
Related to images 6791, 6792, 6793, 6794, 6798, and videos 6795 and 6796.
Related to images 6791, 6792, 6793, 6794, 6798, and videos 6795 and 6796.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media
1013: Lily mitosis 03
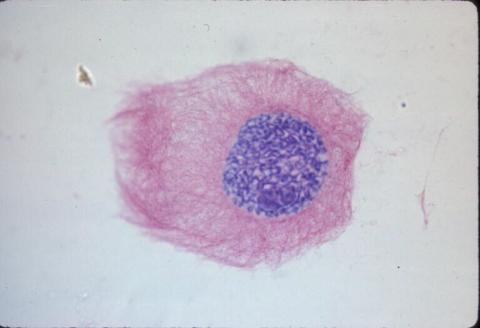
1013: Lily mitosis 03
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue.
Related to images 1010, 1011, 1012, 1014, 1015, 1016, 1017, 1018, 1019, and 1021.
Related to images 1010, 1011, 1012, 1014, 1015, 1016, 1017, 1018, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
6930: Mouse brain 2
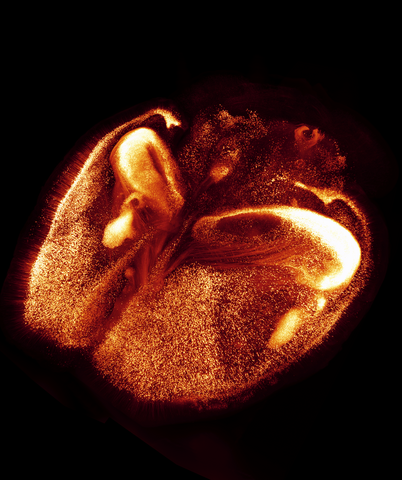
6930: Mouse brain 2
A mouse brain that was genetically modified so that subpopulations of its neurons glow. Researchers often study mice because they share many genes with people and can shed light on biological processes, development, and diseases in humans.
This image was captured using a light sheet microscope.
Related to image 6929 and video 6931.
This image was captured using a light sheet microscope.
Related to image 6929 and video 6931.
Prayag Murawala, MDI Biological Laboratory and Hannover Medical School.
View Media
3364: Nociceptin/orphanin FQ peptide opioid receptor
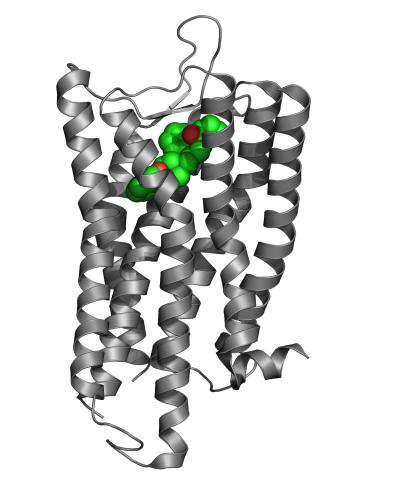
3364: Nociceptin/orphanin FQ peptide opioid receptor
The receptor is shown bound to an antagonist, compound-24
Raymond Stevens, The Scripps Research Institute
View Media
2314: Finding one bug
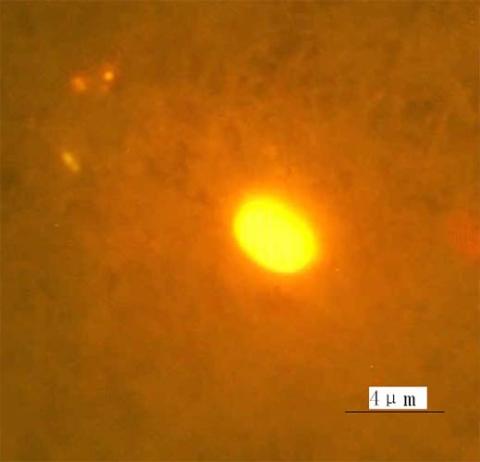
2314: Finding one bug
A nanometer-sized biosensor can detect a single deadly bacterium in tainted ground beef. How? Researchers attached nanoparticles, each packed with thousands of dye molecules, to an antibody that recognizes the microbe E. coli O157:H7. When the nanoball-antibody combo comes into contact with the E. coli bacterium, it glows. Here is the transition, a single bacterial cell glows brightly when it encounters nanoparticle-antibody biosensors, each packed with thousands of dye molecules.
Weihong Tan, University of Florida in Gainesville
View Media
2557: Dicer generates microRNAs (with labels)
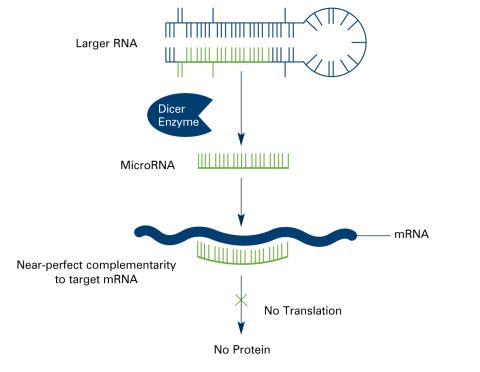
2557: Dicer generates microRNAs (with labels)
The enzyme Dicer generates microRNAs by chopping larger RNA molecules into tiny Velcro®-like pieces. MicroRNAs stick to mRNA molecules and prevent the mRNAs from being made into proteins. See image 2556 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
3478: DDR2 Receptors Attach to Collagen in Breast Tumor
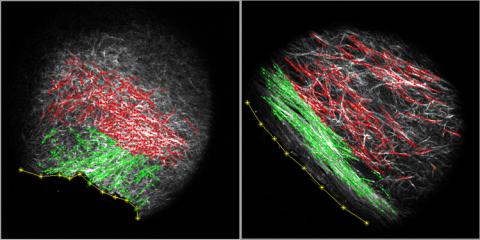
3478: DDR2 Receptors Attach to Collagen in Breast Tumor
On the left, the boundary of a breast tumor (yellow) attaches to collagen fibers that are closest to it (green) using DDR2. On the right, a tumor without DDR2 remains disconnected from the collagen.
Callie Corsa and Suzanne Ponik, Washington University School of Medicine in St. Louis
View Media
2762: Nucleolinus
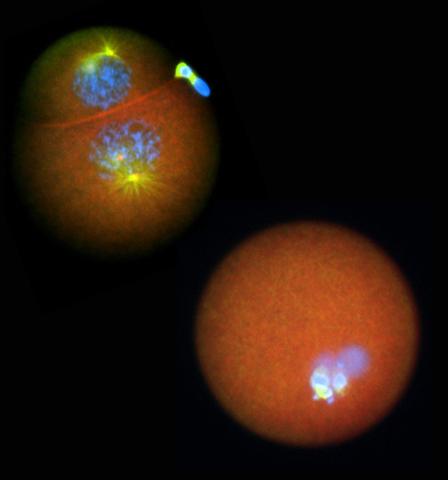
2762: Nucleolinus
The nucleolinus is a cellular compartment that has been a lonely bystander in scientific endeavors. Although it's found in a range of species, its function has been mysterious—mainly because the structure is hard to visualize. An August 2010 study showed that the nucleolinus is crucial for cell division. When researchers zapped the structure with a laser, an egg cell didn't complete division. When the oocyte was fertilized after laser microsurgery (bottom right), the resulting zygote didn't form vital cell division structures (blue and yellow).
Mary Anne Alliegro, Marine Biological Laboratory
View Media
6788: Mitosis and meiosis compared-labeled
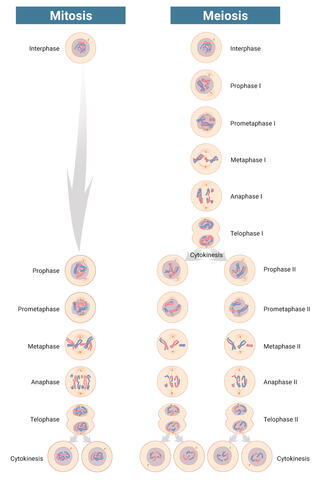
6788: Mitosis and meiosis compared-labeled
Meiosis is used to make sperm and egg cells. During meiosis, a cell's chromosomes are copied once, but the cell divides twice. During mitosis, the chromosomes are copied once, and the cell divides once. For simplicity, cells are illustrated with only three pairs of chromosomes.
See image 1333 for an unlabeled version of this illustration.
See image 1333 for an unlabeled version of this illustration.
Judith Stoffer
View Media
7013: An adult Hawaiian bobtail squid
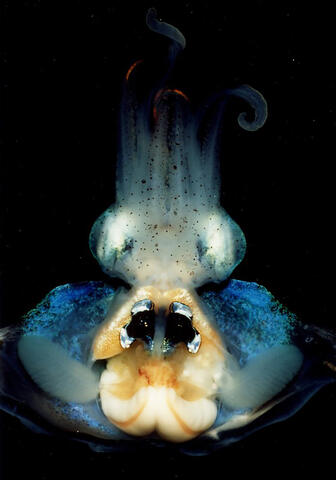
7013: An adult Hawaiian bobtail squid
An adult female Hawaiian bobtail squid, Euprymna scolopes, with its mantle cavity exposed from the underside. Some internal organs are visible, including the two lobes of the light organ that contains bioluminescent bacteria, Vibrio fischeri. The light organ includes accessory tissues like an ink sac (black) that serves as a shutter, and a silvery reflector that directs the light out of the underside of the animal.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
6520: HeLa cell undergoing division into two daughter cells
6520: HeLa cell undergoing division into two daughter cells
Here, a human HeLa cell (a type of immortal cell line used in laboratory experiments) is undergoing cell division. They come from cervical cancer cells that were obtained in 1951 from Henrietta Lacks, a patient at the Johns Hopkins Hospital. The final stage of division, called cytokinesis, occurs after the genomes—shown in yellow—have split into two new daughter cells. The myosin II is a motor protein shown in blue, and the actin filaments, which are types of protein that support cell structure, are shown in red.
Dylan T. Burnette, Ph.D., Vanderbilt University School of Medicine.
View Media
2792: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 03
2792: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 03
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
1316: Mitosis - interphase
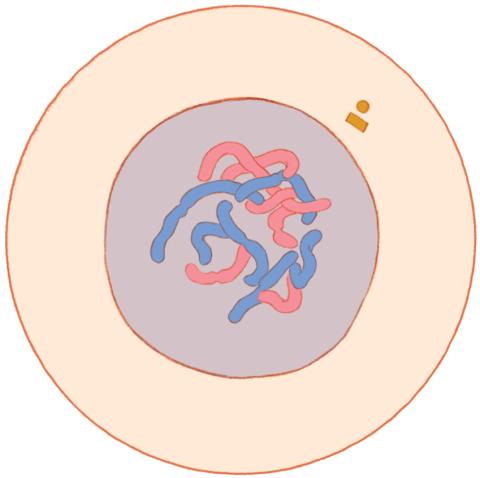
1316: Mitosis - interphase
A cell in interphase, at the start of mitosis: Chromosomes duplicate, and the copies remain attached to each other. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
3558: Bioluminescent imaging in adult zebrafish - lateral view
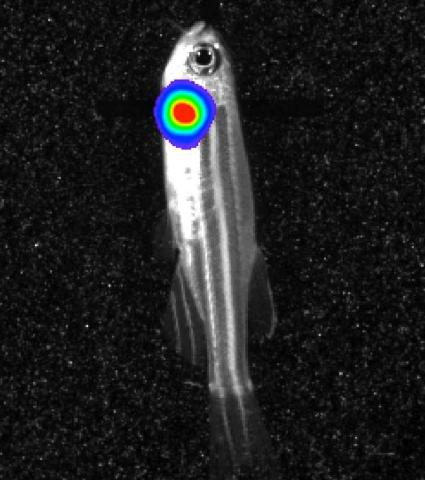
3558: Bioluminescent imaging in adult zebrafish - lateral view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression (lateral view).
For imagery of both the lateral and overhead view go to 3556.
For imagery of the overhead view go to 3557.
For more information about the illumated area go to 3559.
For imagery of both the lateral and overhead view go to 3556.
For imagery of the overhead view go to 3557.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
6995: Measles virus
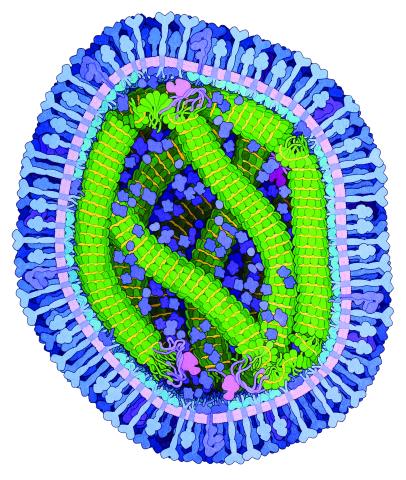
6995: Measles virus
A cross section of the measles virus in which six proteins work together to infect cells. The measles virus is extremely infectious; 9 out of 10 people exposed will contract the disease. Fortunately, an effective vaccine protects against infection.
For a zoomed-in look at the six important proteins, see Measles Virus Proteins.
For a zoomed-in look at the six important proteins, see Measles Virus Proteins.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
3566: Mouse colon with gut bacteria
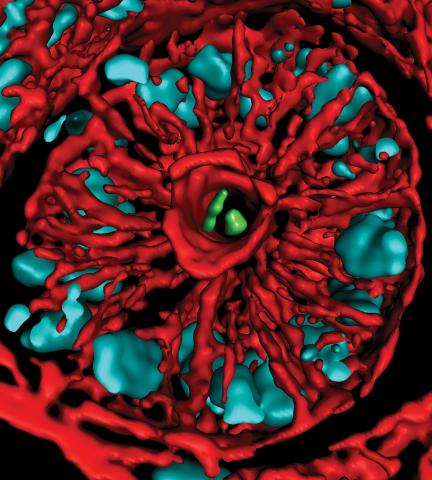
3566: Mouse colon with gut bacteria
A section of mouse colon with gut bacteria (center, in green) residing within a protective pocket. Understanding how microorganisms colonize the gut could help devise ways to correct for abnormal changes in bacterial communities that are associated with disorders like inflammatory bowel disease.
Sarkis K. Mazmanian, California Institute of Technology
View Media
6548: Partial Model of a Cilium’s Doublet Microtubule
6548: Partial Model of a Cilium’s Doublet Microtubule
Cilia (cilium in singular) are complex molecular machines found on many of our cells. One component of cilia is the doublet microtubule, a major part of cilia’s skeletons that give them support and shape. This animated image is a partial model of a doublet microtubule’s structure based on cryo-electron microscopy images. Video can be found here 6549.
Brown Lab, Harvard Medical School and Veronica Falconieri Hays.
View Media
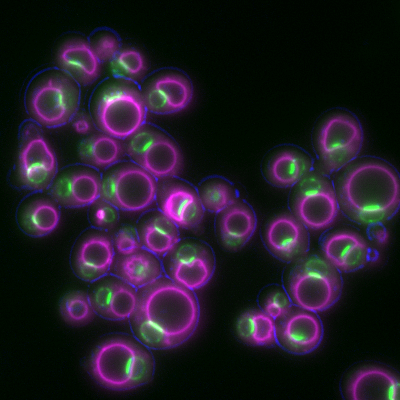
6772: Yeast cells responding to a glucose shortage
6772: Yeast cells responding to a glucose shortage
These yeast cells were exposed to a glucose (sugar) shortage. This caused the cells to compartmentalize HMGCR (green)—an enzyme involved in making cholesterol—to a patch on the nuclear envelope next to the vacuole/lysosome (purple). This process enhanced HMGCR activity and helped the yeast adapt to the glucose shortage. Researchers hope that understanding how yeast regulate cholesterol could ultimately lead to new ways to treat high cholesterol in people. This image was captured using a fluorescence microscope.
Mike Henne, University of Texas Southwestern Medical Center.
View Media
6850: Himastatin and bacteria
6850: Himastatin and bacteria
A model of the molecule himastatin overlaid on an image of Bacillus subtilis bacteria. Scientists first isolated himastatin from the bacterium Streptomyces himastatinicus, and the molecule shows antibiotic activity. The researchers who created this image developed a new, more concise way to synthesize himastatin so it can be studied more easily. They also tested the effects of himastatin and derivatives of the molecule on B. subtilis.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6848 and video 6851.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6848 and video 6851.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media