Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
2554: RNA strand
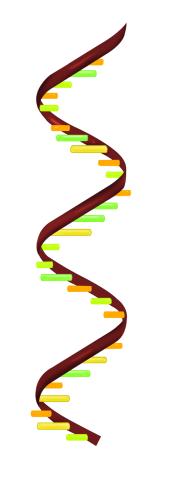
2554: RNA strand
Ribonucleic acid (RNA) has a sugar-phosphate backbone and the bases adenine (A), cytosine (C), guanine (G), and uracil (U). See image 2555 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
6585: Cell-like compartments from frog eggs 2
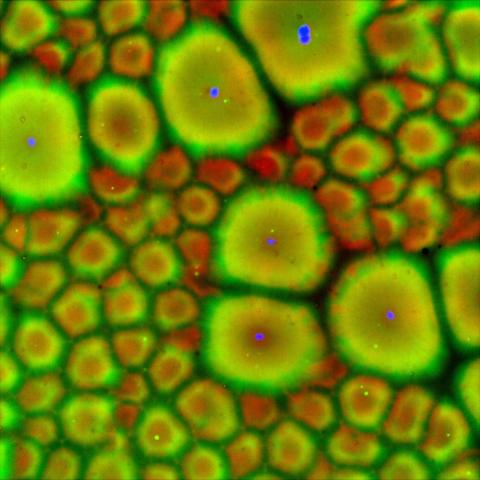
6585: Cell-like compartments from frog eggs 2
Cell-like compartments that spontaneously emerged from scrambled frog eggs, with nuclei (blue) from frog sperm. Endoplasmic reticulum (red) and microtubules (green) are also visible. Regions without nuclei formed smaller compartments. Image created using epifluorescence microscopy.
For more photos of cell-like compartments from frog eggs view: 6584, 6586, 6591, 6592, and 6593.
For videos of cell-like compartments from frog eggs view: 6587, 6588, 6589, and 6590.
Xianrui Cheng, Stanford University School of Medicine.
View Media
6522: Fruit fly ovary
6522: Fruit fly ovary
In this image of a stained fruit fly ovary, the ovary is packed with immature eggs (with DNA stained blue). The cytoskeleton (in pink) is a collection of fibers that gives a cell shape and support. The signal-transmitting molecules like STAT (in yellow) are common to reproductive processes in humans. Researchers used this image to show molecular staining and high-resolution imaging techniques to students.
Crystal D. Rogers, Ph.D., University of California, Davis, School of Veterinary Medicine; and Mariano A. Loza-Coll, Ph.D., California State University, Northridge.
View Media
3628: Skin cancer cells (squamous cell carcinoma)
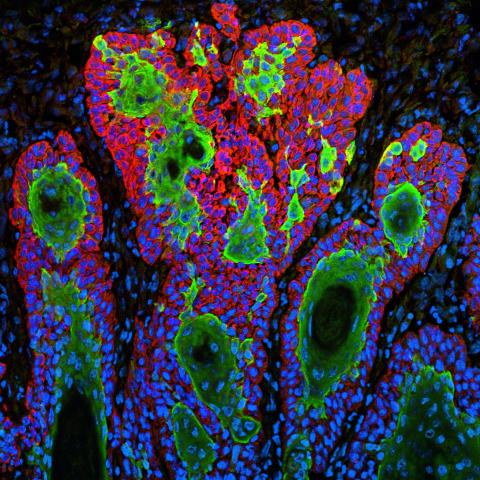
3628: Skin cancer cells (squamous cell carcinoma)
This image shows the uncontrolled growth of cells in squamous cell carcinoma, the second most common form of skin cancer. If caught early, squamous cell carcinoma is usually not life-threatening.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Markus Schober and Elaine Fuchs, The Rockefeller University
View Media
3789: Nucleolus subcompartments spontaneously self-assemble 1
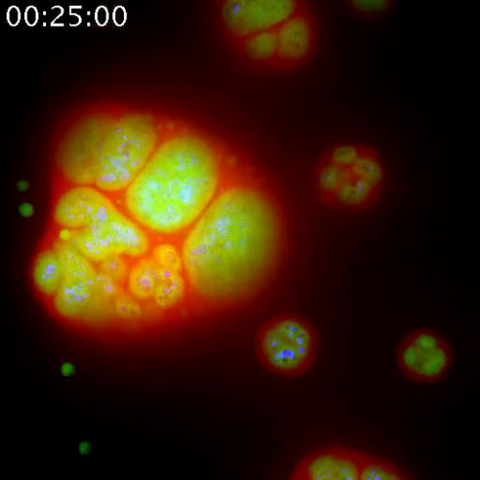
3789: Nucleolus subcompartments spontaneously self-assemble 1
The nucleolus is a small but very important protein complex located in the cell's nucleus. It forms on the chromosomes at the location where the genes for the RNAs are that make up the structure of the ribosome, the indispensable cellular machine that makes proteins from messenger RNAs.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from difference in how the proteins in each compartment mix with water and with each other. These differences let them readily separate from each other into the three nucleolus compartments.
This video of nucleoli in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows how each of the compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue) spontaneously fuse with each other on encounter without mixing with the other compartments. For more details on this research, see this press release from Princeton. Related to video 3791, image 3792 and image 3793.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from difference in how the proteins in each compartment mix with water and with each other. These differences let them readily separate from each other into the three nucleolus compartments.
This video of nucleoli in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows how each of the compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue) spontaneously fuse with each other on encounter without mixing with the other compartments. For more details on this research, see this press release from Princeton. Related to video 3791, image 3792 and image 3793.
Nilesh Vaidya, Princeton University
View Media

2779: Mature, flowering Arabidopsis
2779: Mature, flowering Arabidopsis
This is an adult flowering Arabidopsis thaliana plant with the inbred designation L-er. Arabidopsis is the most widely used model organism for researchers who study plant genetics.
Jeff Dangl, University of North Carolina, Chapel Hill
View Media
7021: Single-cell “radios” image
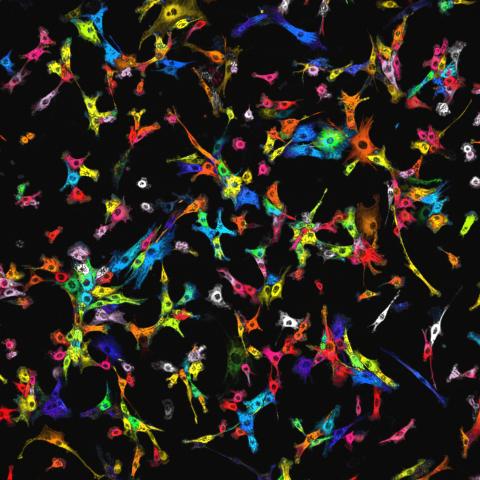
7021: Single-cell “radios” image
Individual cells are color-coded based on their identity and signaling activity using a protein circuit technology developed by the Coyle Lab. Just as a radio allows you to listen to an individual frequency, this technology allows researchers to tune into the specific “radio station” of each cell through genetically encoded proteins from a bacterial system called MinDE. The proteins generate an oscillating fluorescent signal that transmits information about cell shape, state, and identity that can be decoded using digital signal processing tools originally designed for telecommunications. The approach allows researchers to look at the dynamics of a single cell in the presence of many other cells.
Related to video 7022.
Related to video 7022.
Scott Coyle, University of Wisconsin-Madison.
View Media
2419: Mapping brain differences
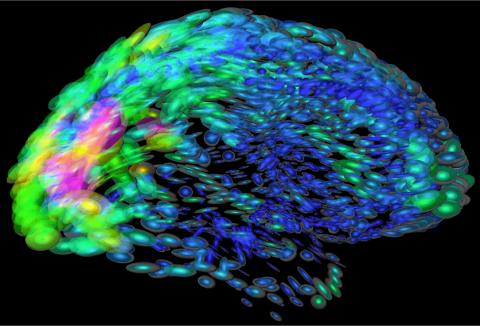
2419: Mapping brain differences
This image of the human brain uses colors and shapes to show neurological differences between two people. The blurred front portion of the brain, associated with complex thought, varies most between the individuals. The blue ovals mark areas of basic function that vary relatively little. Visualizations like this one are part of a project to map complex and dynamic information about the human brain, including genes, enzymes, disease states, and anatomy. The brain maps represent collaborations between neuroscientists and experts in math, statistics, computer science, bioinformatics, imaging, and nanotechnology.
Arthur Toga, University of California, Los Angeles
View Media
1011: Lily mitosis 11
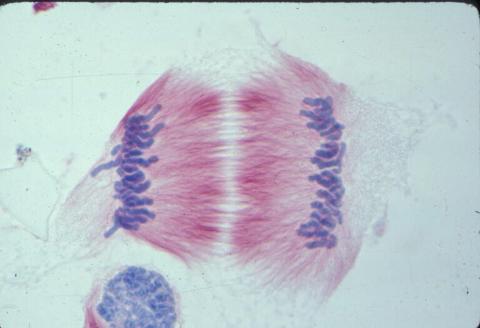
1011: Lily mitosis 11
A light microscope image of cells from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and have separated into the opposite sides of a dividing cell.
Related to images 1010, 1012, 1013, 1014, 1015, 1016, 1017, 1018, 1019, and 1021.
Related to images 1010, 1012, 1013, 1014, 1015, 1016, 1017, 1018, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
2325: Multicolor STORM
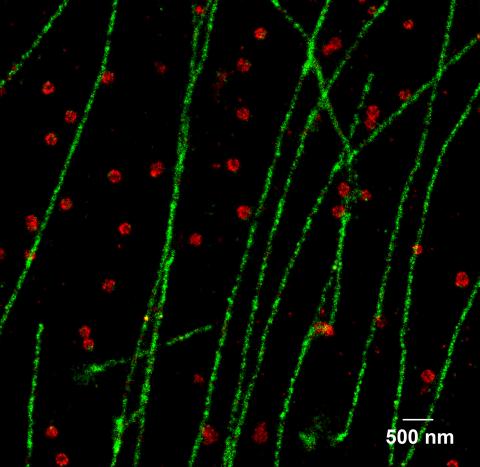
2325: Multicolor STORM
In 2006, scientists developed an optical microscopy technique enabling them to clearly see individual molecules within cells. In 2007, they took the technique, abbreviated STORM, a step further. They identified multicolored probes that let them peer into cells and clearly see multiple cellular components at the same time, such as these microtubules (green) and small hollows called clathrin-coated pits (red). Unlike conventional methods, the multicolor STORM technique produces a crisp and high resolution picture. A sharper view of how cellular components interact will likely help scientists answer some longstanding questions about cell biology.
Xiaowei Zhuang, Harvard University
View Media
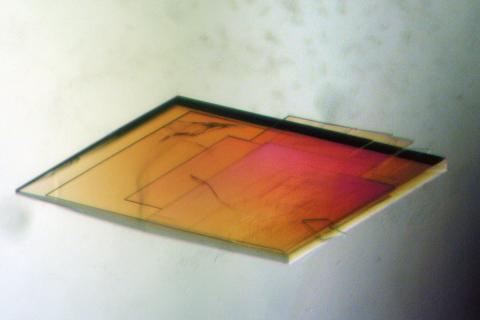

2403: Pig trypsin crystal
2403: Pig trypsin crystal
A crystal of pig trypsin protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
2601: Mouse liver labeled with fluorescent probe
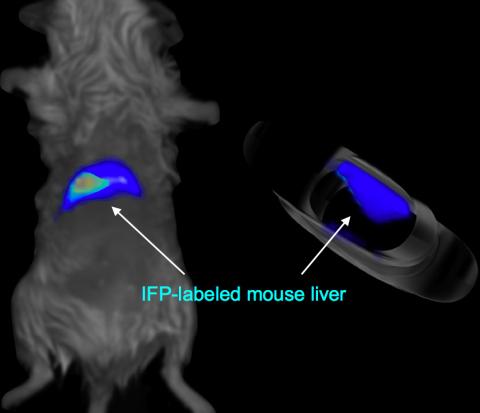
2601: Mouse liver labeled with fluorescent probe
A mouse liver glows after being tagged with specially designed infrared-fluorescent protein (IFP). Since its discovery in 1962, green fluorescent protein (GFP) has become an invaluable resource in biomedical imaging. But because of its short wavelength, the light that makes GFP glow doesn't penetrate far in whole animals. So University of California, San Diego cell biologist Roger Tsien--who shared the 2008 Nobel Prize in chemistry for groundbreaking work with GFP--made infrared-fluorescent proteins (IFPs) that shine under longer-wavelength light, allowing whole-body imaging in small animals.
Xiaokun Shu, University of California, San Diego
View Media
2565: Recombinant DNA (with labels)
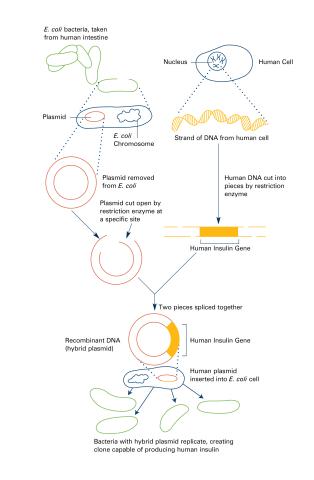
2565: Recombinant DNA (with labels)
To splice a human gene (in this case, the one for insulin) into a plasmid, scientists take the plasmid out of an E. coli bacterium, cut the plasmid with a restriction enzyme, and splice in insulin-making human DNA. The resulting hybrid plasmid can be inserted into another E. coli bacterium, where it multiplies along with the bacterium. There, it can produce large quantities of insulin. See image 2564 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
2364: High-throughput protein structure determination pipeline
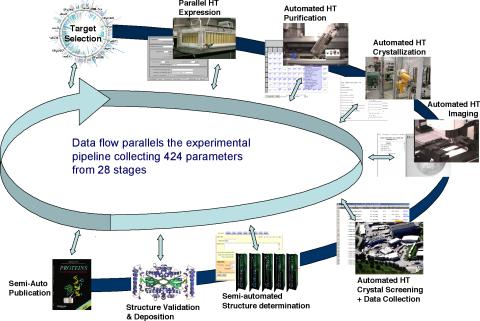
2364: High-throughput protein structure determination pipeline
This slide shows the technologies that the Joint Center for Structural Genomics developed for going from gene to structure and how the technologies have been integrated into a high-throughput pipeline, including all of the steps from target selection, parallel expression, protein purification, automated crystallization trials, automated crystal screening, structure determination, validation, and publication.
Joint Center for Structural Genomics
View Media
3314: Human opioid receptor structure superimposed on poppy
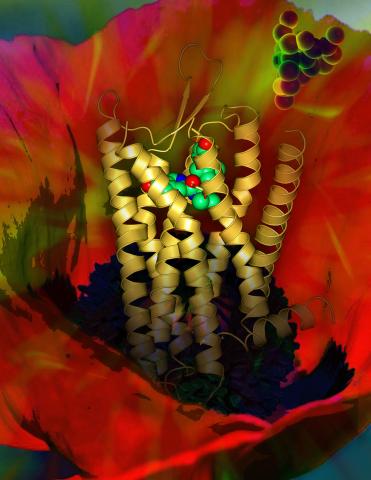
3314: Human opioid receptor structure superimposed on poppy
Opioid receptors on the surfaces of brain cells are involved in pleasure, pain, addiction, depression, psychosis, and other conditions. The receptors bind to both innate opioids and drugs ranging from hospital anesthetics to opium. Researchers at The Scripps Research Institute, supported by the NIGMS Protein Structure Initiative, determined the first three-dimensional structure of a human opioid receptor, a kappa-opioid receptor. In this illustration, the submicroscopic receptor structure is shown while bound to an agonist (or activator). The structure is superimposed on a poppy flower, the source of opium.
Raymond Stevens, The Scripps Research Institute
View Media
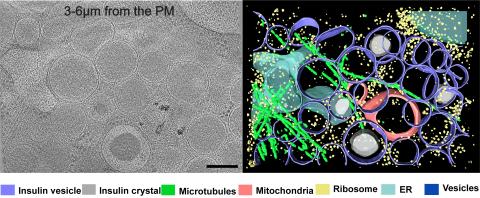

6607: Cryo-ET cell cross-section visualizing insulin vesicles
6607: Cryo-ET cell cross-section visualizing insulin vesicles
On the left, a cross-section slice of a rat pancreas cell captured using cryo-electron tomography (cryo-ET). On the right, a color-coded, 3D version of the image highlighting cell structures. Visible features include insulin vesicles (purple rings), insulin crystals (gray circles), microtubules (green rods), ribosomes (small yellow circles). The black line at the bottom right of the left image represents 200 nm. Related to image 6608.
Xianjun Zhang, University of Southern California.
View Media
2796: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 03
2796: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 03
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media

3735: Scanning electron microscopy of collagen fibers
3735: Scanning electron microscopy of collagen fibers
This image shows collagen, a fibrous protein that's the main component of the extracellular matrix (ECM). Collagen is a strong, ropelike molecule that forms stretch-resistant fibers. The most abundant protein in our bodies, collagen accounts for about a quarter of our total protein mass. Among its many functions is giving strength to our tendons, ligaments and bones and providing scaffolding for skin wounds to heal. There are about 20 different types of collagen in our bodies, each adapted to the needs of specific tissues.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
3742: Confocal microscopy of perineuronal nets in the brain 2
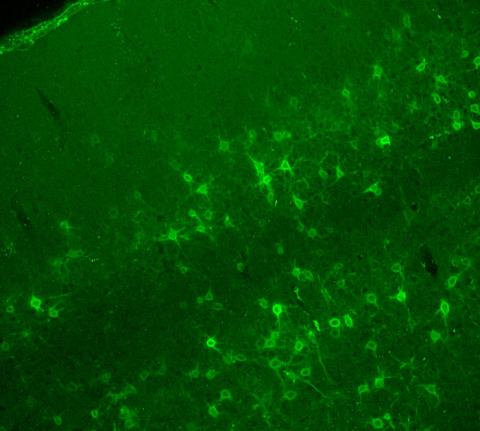
3742: Confocal microscopy of perineuronal nets in the brain 2
The photo shows a confocal microscopy image of perineuronal nets (PNNs), which are specialized extracellular matrix (ECM) structures in the brain. The PNN surrounds some nerve cells in brain regions including the cortex, hippocampus and thalamus. Researchers study the PNN to investigate their involvement stabilizing the extracellular environment and forming nets around nerve cells and synapses in the brain. Abnormalities in the PNNs have been linked to a variety of disorders, including epilepsy and schizophrenia, and they limit a process called neural plasticity in which new nerve connections are formed. To visualize the PNNs, researchers labeled them with Wisteria floribunda agglutinin (WFA)-fluorescein. Related to image 3741.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
1306: Vesicular shuttle model
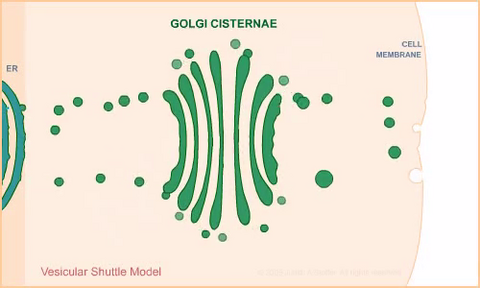
1306: Vesicular shuttle model
Animation for the vesicular shuttle model of Golgi transport.
Judith Stoffer
View Media
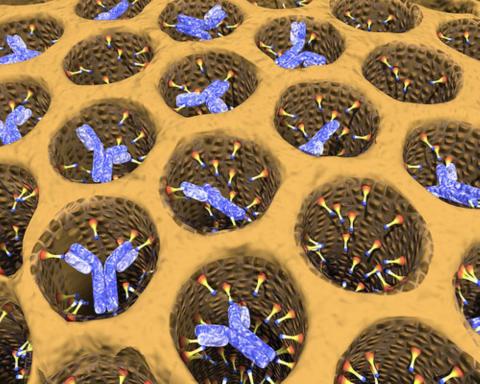

2750: Antibodies in silica honeycomb
2750: Antibodies in silica honeycomb
Antibodies are among the most promising therapies for certain forms of cancer, but patients must take them intravenously, exposing healthy tissues to the drug and increasing the risk of side effects. A team of biochemists packed the anticancer antibodies into porous silica particles to deliver a heavy dose directly to tumors in mice.
Chenghong Lei, Pacific Northwest National Laboratory & Karl Erik Hellstrom, University of Washington
View Media
1160: Vibrio bacteria
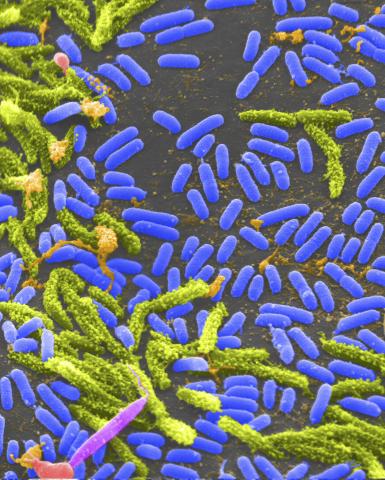
1160: Vibrio bacteria
Vibrio, a type (genus) of rod-shaped bacteria. Some Vibrio species cause cholera in humans.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
6608: Cryo-ET cross-section of a rat pancreas cell
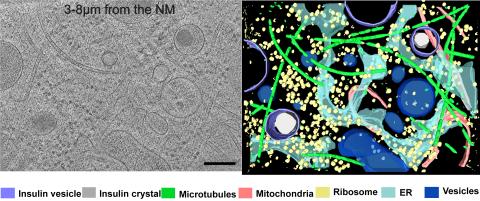
6608: Cryo-ET cross-section of a rat pancreas cell
On the left, a cross-section slice of a rat pancreas cell captured using cryo-electron tomography (cryo-ET). On the right, a 3D, color-coded version of the image highlighting cell structures. Visible features include microtubules (neon-green rods), ribosomes (small yellow circles), and vesicles (dark-blue circles). These features are surrounded by the partially visible endoplasmic reticulum (light blue). The black line at the bottom right of the left image represents 200 nm. Related to image 6607.
Xianjun Zhang, University of Southern California.
View Media
2383: PanC from M. tuberculosis
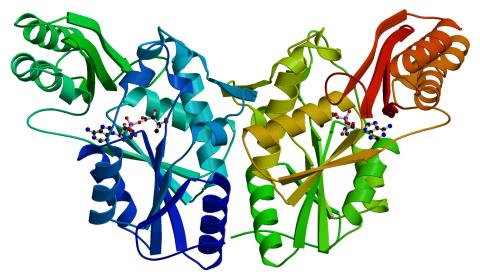
2383: PanC from M. tuberculosis
Model of an enzyme, PanC, that is involved in the last step of vitamin B5 biosynthesis in Mycobacterium tuberculosis. PanC is essential for the growth of M. tuberculosis, which causes most cases of tuberculosis, and is therefore a potential drug target.
Mycobacterium Tuberculosis Center, PSI
View Media
2347: Cysteine dioxygenase from mouse
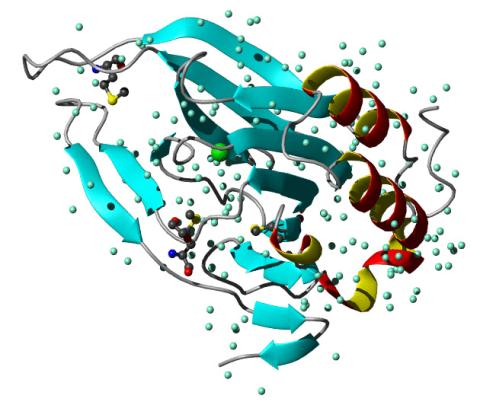
2347: Cysteine dioxygenase from mouse
Model of the mammalian iron enzyme cysteine dioxygenase from a mouse.
Center for Eukaryotic Structural Genomics, PSI
View Media
6993: RNA polymerase
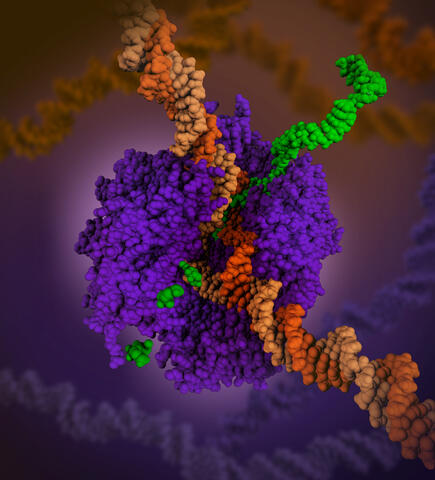
6993: RNA polymerase
RNA polymerase (purple) is a complex enzyme at the heart of transcription. During this process, the enzyme unwinds the DNA double helix and uses one strand (darker orange) as a template to create the single-stranded messenger RNA (green), later used by ribosomes for protein synthesis.
From the RNA polymerase II elongation complex of Saccharomyces cerevisiae (PDB entry 1I6H) as seen in PDB-101's What is a Protein? video.
From the RNA polymerase II elongation complex of Saccharomyces cerevisiae (PDB entry 1I6H) as seen in PDB-101's What is a Protein? video.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
6888: Chromatin in human fibroblast
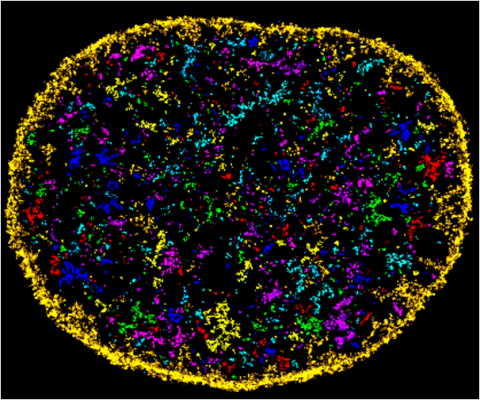
6888: Chromatin in human fibroblast
The nucleus of a human fibroblast cell with chromatin—a substance made up of DNA and proteins—shown in various colors. Fibroblasts are one of the most common types of cells in mammalian connective tissue, and they play a key role in wound healing and tissue repair. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6887 and 6893.
Related to images 6887 and 6893.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media
2318: Gene silencing
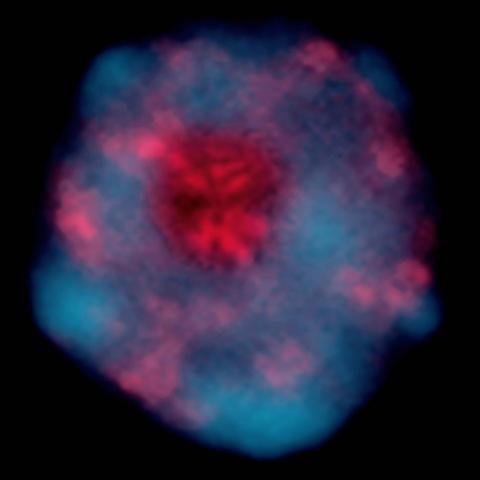
2318: Gene silencing
Pretty in pink, the enzyme histone deacetylase (HDA6) stands out against a background of blue-tinted DNA in the nucleus of an Arabidopsis plant cell. Here, HDA6 concentrates in the nucleolus (top center), where ribosomal RNA genes reside. The enzyme silences the ribosomal RNA genes from one parent while those from the other parent remain active. This chromosome-specific silencing of ribosomal RNA genes is an unusual phenomenon observed in hybrid plants.
Olga Pontes and Craig Pikaard, Washington University
View Media
6793: Yeast cells with endocytic actin patches
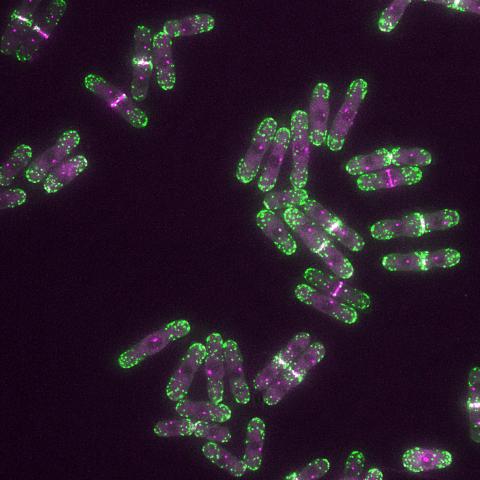
6793: Yeast cells with endocytic actin patches
Yeast cells with endocytic actin patches (green). These patches help cells take in outside material. When a cell is in interphase, patches concentrate at its ends. During later stages of cell division, patches move to where the cell splits. This image was captured using wide-field microscopy with deconvolution.
Related to images 6791, 6792, 6794, 6797, 6798, and videos 6795 and 6796.
Related to images 6791, 6792, 6794, 6797, 6798, and videos 6795 and 6796.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media
6780: Calling Cards in a mouse brain
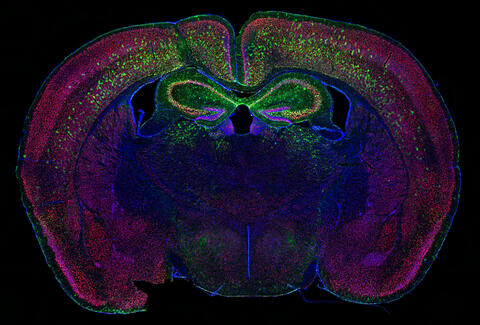
6780: Calling Cards in a mouse brain
The green spots in this mouse brain are cells labeled with Calling Cards, a technology that records molecular events in brain cells as they mature. Understanding these processes during healthy development can guide further research into what goes wrong in cases of neuropsychiatric disorders. Also fluorescently labeled in this image are neurons (red) and nuclei (blue). Calling Cards and its application are described in the Cell paper “Self-Reporting Transposons Enable Simultaneous Readout of Gene Expression and Transcription Factor Binding in Single Cells” by Moudgil et al.; and the Proceedings of the National Academy of Sciences paper “A viral toolkit for recording transcription factor–DNA interactions in live mouse tissues” by Cammack et al. The technology was also featured in the NIH Director’s Blog post The Amazing Brain: Tracking Molecular Events with Calling Cards.
Related to video
Related to video
Allen Yen, Lab of Joseph Dougherty, Washington University School of Medicine in St. Louis.
View Media
5885: 3-D Architecture of a Synapse
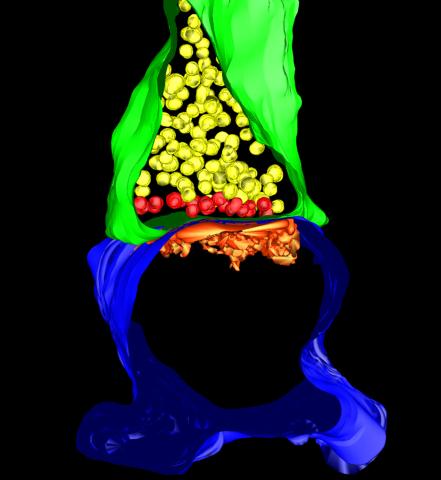
5885: 3-D Architecture of a Synapse
This image shows the structure of a synapse, or junction between two nerve cells in three dimensions. From the brain of a mouse.
Anton Maximov, The Scripps Research Institute, La Jolla, CA
View Media
3488: Shiga toxin being sorted inside a cell
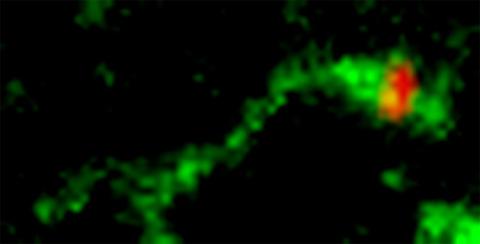
3488: Shiga toxin being sorted inside a cell
Shiga toxin (green) is sorted from the endosome into membrane tubules (red), which then pinch off and move to the Golgi apparatus.
Somshuvra Mukhopadhyay, The University of Texas at Austin, and Adam D. Linstedt, Carnegie Mellon University
View Media
3402: Hsp33 Heat Shock Protein Inactive to Active
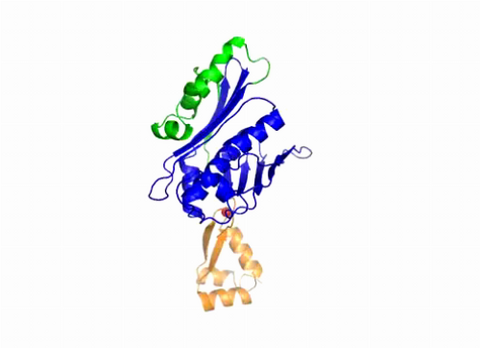
3402: Hsp33 Heat Shock Protein Inactive to Active
When the heat shock protein hsp33 is folded, it is inactive and contains a zinc ion, stabilizing the redox sensitive domain (orange). In the presence of an environmental stressor, the protein releases the zinc ion, which leads to the unfolding of the redox domain. This unfolding causes the chaperone to activate by reaching out its "arm" (green) to protect other proteins.
Dana Reichmann, University of Michigan
View Media

2723: iPS cell facility at the Coriell Institute for Medical Research
2723: iPS cell facility at the Coriell Institute for Medical Research
This lab space was designed for work on the induced pluripotent stem (iPS) cell collection, part of the NIGMS Human Genetic Cell Repository at the Coriell Institute for Medical Research.
Courtney Sill, Coriell Institute for Medical Research
View Media
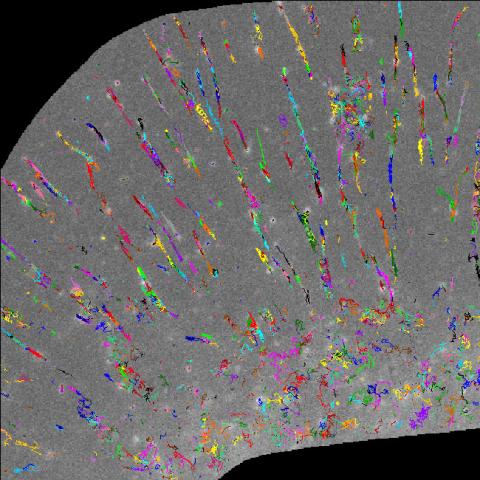
2801: Trajectories of labeled cell receptors
3270: Dopaminergic neurons from ES cells
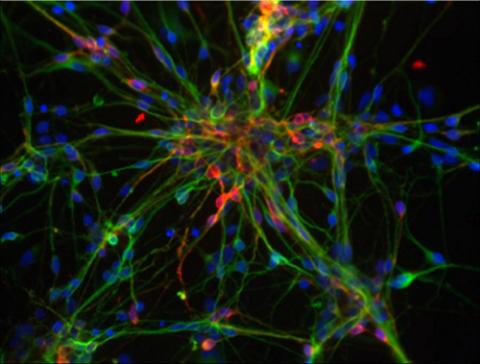
3270: Dopaminergic neurons from ES cells
Human embryonic stem cells differentiated into dopaminergic neurons, the type that degenerate in Parkinson's disease. Image courtesy of the California Institute for Regenerative Medicine. Related to images 3271 and 3285.
Jeannie Liu, Lab of Jan Nolta, University of California, Davis, via CIRM
View Media
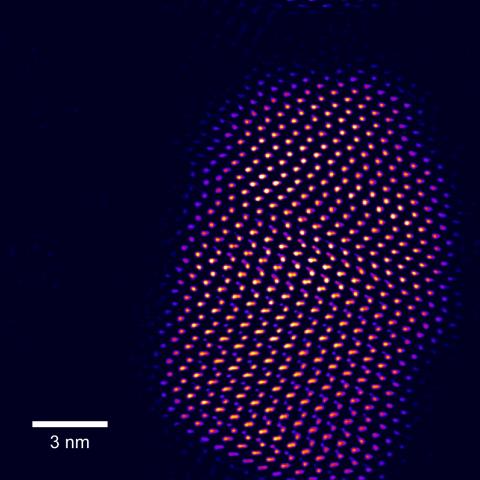

2332: Tiny points of light in a quantum dot
2332: Tiny points of light in a quantum dot
This fingertip-shaped group of lights is a microscopic crystal called a quantum dot. About 10,000 times thinner than a sheet of paper, the dot radiates brilliant colors under ultraviolet light. Dots such as this one allow researchers to label and track individual molecules in living cells and may be used for speedy disease diagnosis, DNA testing, and screening for illegal drugs.
Sandra Rosenthal and James McBride, Vanderbilt University, and Stephen Pennycook, Oak Ridge National Laboratory
View Media
2367: Map of protein structures 02
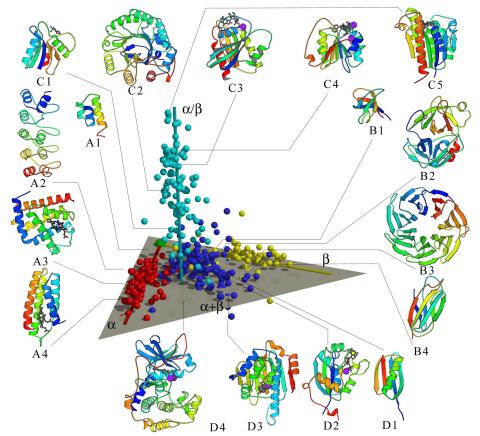
2367: Map of protein structures 02
A global "map of the protein structure universe" indicating the positions of specific proteins. The preponderance of small, less-structured proteins near the origin, with the more highly structured, large proteins towards the ends of the axes, may suggest the evolution of protein structures.
Berkeley Structural Genomics Center, PSI
View Media
2309: Cellular polarity
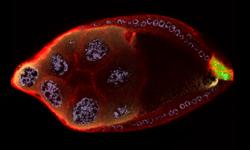
2309: Cellular polarity
As an egg cell develops, a process called polarization controls what parts ultimately become the embryo's head and tail. This picture shows an egg of the fruit fly Drosophila. Red and green mark two types of signaling proteins involved in polarization. Disrupting these signals can scramble the body plan of the embryo, leading to severe developmental disorders.
Wu-Min Deng, Florida State University
View Media
6568: Correlative imaging by annotation with single molecules (CIASM) process
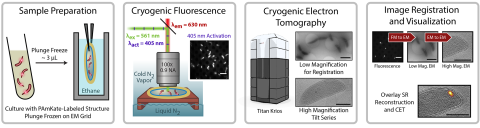
6568: Correlative imaging by annotation with single molecules (CIASM) process
These images illustrate a technique combining cryo-electron tomography and super-resolution fluorescence microscopy called correlative imaging by annotation with single molecules (CIASM). CIASM enables researchers to identify small structures and individual molecules in cells that they couldn’t using older techniques.
Peter Dahlberg, Stanford University.
View Media
1316: Mitosis - interphase
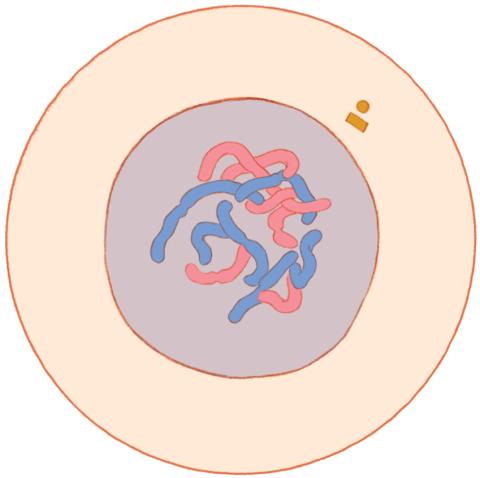
1316: Mitosis - interphase
A cell in interphase, at the start of mitosis: Chromosomes duplicate, and the copies remain attached to each other. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
6352: CRISPR surveillance complex
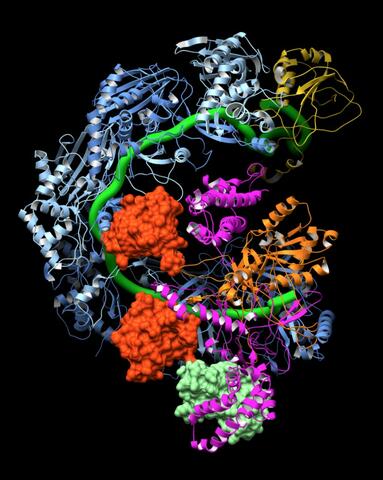
6352: CRISPR surveillance complex
This image shows how the CRISPR surveillance complex is disabled by two copies of anti-CRISPR protein AcrF1 (red) and one AcrF2 (light green). These anti-CRISPRs block access to the CRISPR RNA (green tube) preventing the surveillance complex from scanning and targeting invading viral DNA for destruction.
NRAMM National Resource for Automated Molecular Microscopy http://nramm.nysbc.org/nramm-images/ Source: Bridget Carragher
View Media
2382: PanB from M. tuberculosis (2)
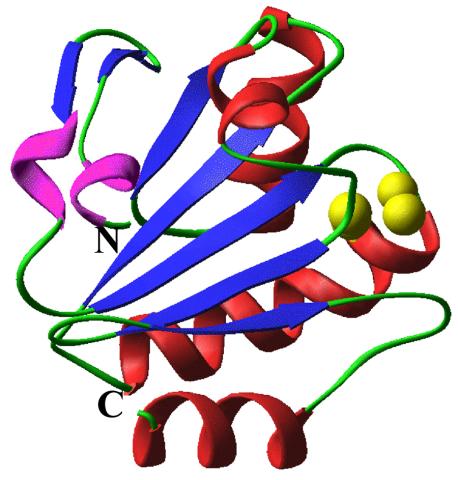
2382: PanB from M. tuberculosis (2)
Model of an enzyme, PanB, from Mycobacterium tuberculosis, the bacterium that causes most cases of tuberculosis. This enzyme is an attractive drug target.
Mycobacterium Tuberculosis Center, PSI-1
View Media
6750: C. elegans with blue and yellow lights in the background
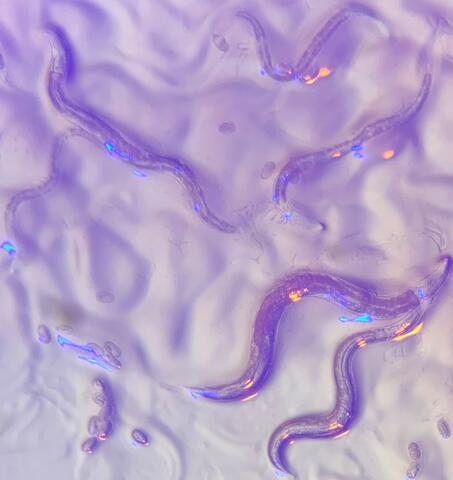
6750: C. elegans with blue and yellow lights in the background
These microscopic roundworms, called Caenorhabditis elegans, lack eyes and the opsin proteins used by visual systems to detect colors. However, researchers found that the worms can still sense the color of light in a way that enables them to avoid pigmented toxins made by bacteria. This image was captured using a stereo microscope.
H. Robert Horvitz and Dipon Ghosh, Massachusetts Institute of Technology.
View Media
3590: Fruit fly spermatids
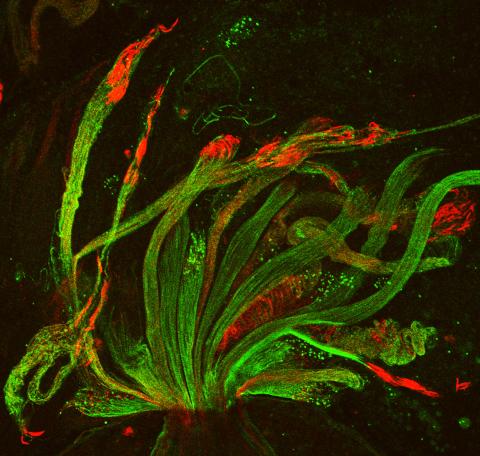
3590: Fruit fly spermatids
Developing spermatids (precursors of mature sperm cells) begin as small, round cells and mature into long-tailed, tadpole-shaped ones. In the sperm cell's head is the cell nucleus; in its tail is the power to outswim thousands of competitors to fertilize an egg. As seen in this microscopy image, fruit fly spermatids start out as groups of interconnected cells. A small lipid molecule called PIP2 helps spermatids tell their heads from their tails. Here, PIP2 (red) marks the nuclei and a cell skeleton-building protein called tubulin (green) marks the tails. When PIP2 levels are too low, some spermatids get mixed up and grow with their heads at the wrong end. Because sperm development is similar across species, studies in fruit flies could help researchers understand male infertility in humans.
Lacramioara Fabian, The Hospital for Sick Children, Toronto, Canada
View Media
6982: Insulin production and fat sensing in fruit flies
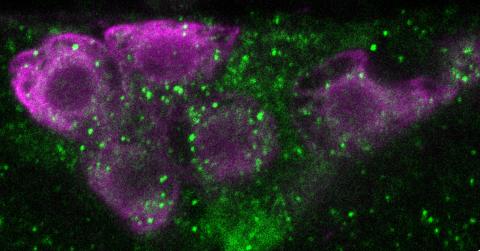
6982: Insulin production and fat sensing in fruit flies
Fourteen neurons (magenta) in the adult Drosophila brain produce insulin, and fat tissue sends packets of lipids to the brain via the lipoprotein carriers (green). This image was captured using a confocal microscope and shows a maximum intensity projection of many slices.
Related to images 6983, 6984, and 6985.
Related to images 6983, 6984, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media
5761: A panorama view of cells
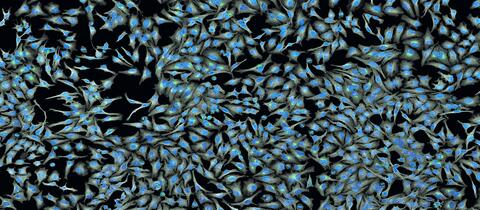
5761: A panorama view of cells
This photograph shows a panoramic view of HeLa cells, a cell line many researchers use to study a large variety of important research questions. The cells' nuclei containing the DNA are stained in blue and the cells' cytoskeletons in gray.
Tom Deerinck, National Center for Microscopy and Imaging Research
View Media
2725: Supernova bacteria
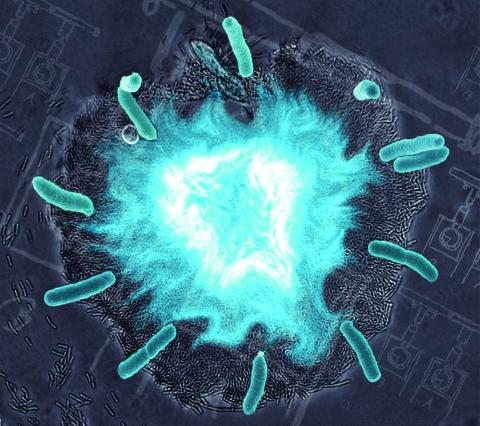
2725: Supernova bacteria
Bacteria engineered to act as genetic clocks flash in synchrony. Here, a "supernova" burst in a colony of coupled genetic clocks just after reaching critical cell density. Superimposed: A diagram from the notebook of Christiaan Huygens, who first characterized synchronized oscillators in the 17th century.
Jeff Hasty, UCSD
View Media
6788: Mitosis and meiosis compared-labeled
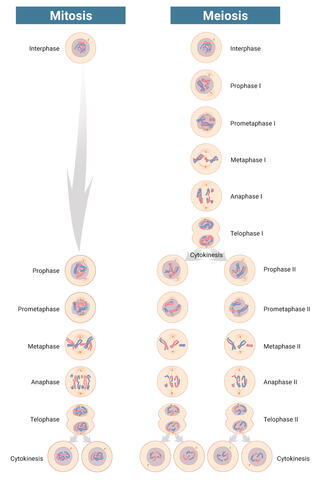
6788: Mitosis and meiosis compared-labeled
Meiosis is used to make sperm and egg cells. During meiosis, a cell's chromosomes are copied once, but the cell divides twice. During mitosis, the chromosomes are copied once, and the cell divides once. For simplicity, cells are illustrated with only three pairs of chromosomes.
See image 1333 for an unlabeled version of this illustration.
See image 1333 for an unlabeled version of this illustration.
Judith Stoffer
View Media
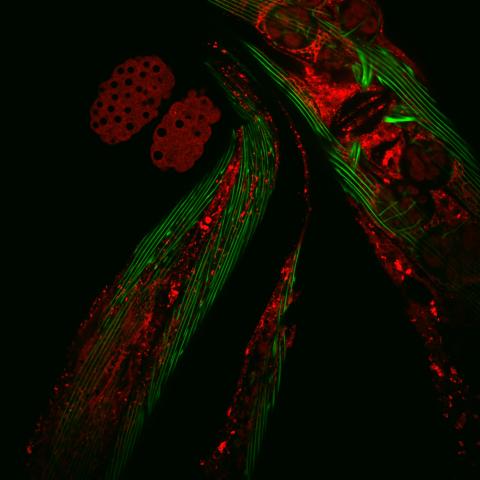
6583: Closeup of fluorescent C. elegans showing muscle and ribosomal protein
6583: Closeup of fluorescent C. elegans showing muscle and ribosomal protein
Closeup of C. elegans, tiny roundworms, with a ribosomal protein glowing red and muscle fibers glowing green. Researchers used these worms to study a molecular pathway that affects aging. The ribosomal protein is involved in protein translation and may play a role in dietary restriction-induced longevity. Image created using confocal microscopy.
View single roundworm here 6581.
View group of roundworms here 6582.
View single roundworm here 6581.
View group of roundworms here 6582.
Jarod Rollins, Mount Desert Island Biological Laboratory.
View Media
