Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.

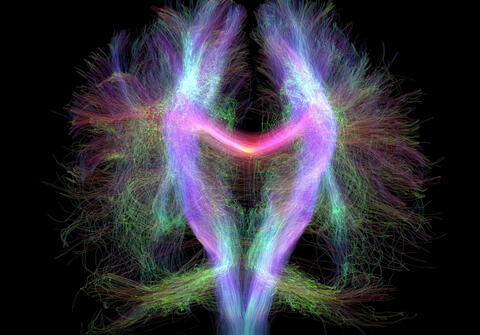
2709: Retroviruses as fossils
2709: Retroviruses as fossils
DNA doesn't leave a fossil record in stone, the way bones do. Instead, the DNA code itself holds the best evidence for organisms' genetic history. Some of the most telling evidence about genetic history comes from retroviruses, the remnants of ancient viral infections.
Emily Harrington, science illustrator
View Media
3638: HIV, the AIDS virus, infecting a human cell
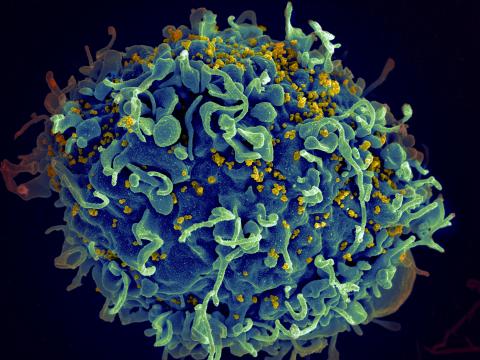
3638: HIV, the AIDS virus, infecting a human cell
This human T cell (blue) is under attack by HIV (yellow), the virus that causes AIDS. The virus specifically targets T cells, which play a critical role in the body's immune response against invaders like bacteria and viruses.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Seth Pincus, Elizabeth Fischer, and Austin Athman, National Institute of Allergy and Infectious Diseases, National Institutes of Health
View Media
3542: Structure of amyloid-forming prion protein
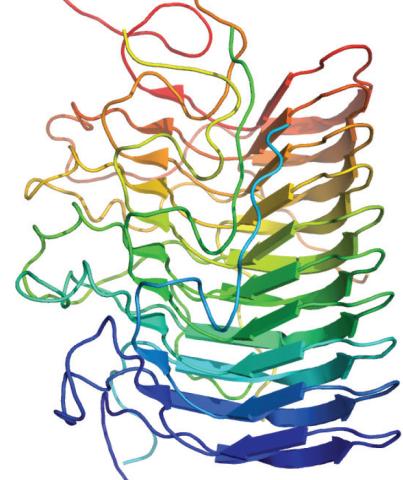
3542: Structure of amyloid-forming prion protein
This structure from an amyloid-forming prion protein shows one way beta sheets can stack. Image originally appeared in a December 2012 PLOS Biology paper.
Douglas Fowler, University of Washington
View Media
3488: Shiga toxin being sorted inside a cell
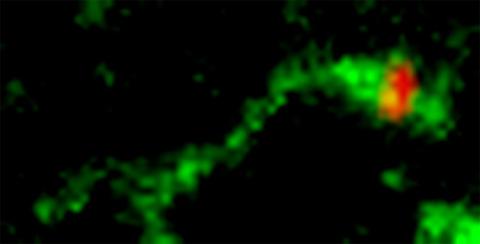
3488: Shiga toxin being sorted inside a cell
Shiga toxin (green) is sorted from the endosome into membrane tubules (red), which then pinch off and move to the Golgi apparatus.
Somshuvra Mukhopadhyay, The University of Texas at Austin, and Adam D. Linstedt, Carnegie Mellon University
View Media
2441: Hydra 05
2441: Hydra 05
Hydra magnipapillata is an invertebrate animal used as a model organism to study developmental questions, for example the formation of the body axis.
Hiroshi Shimizu, National Institute of Genetics in Mishima, Japan
View Media
6800: Magnetic Janus particle activating a T cell
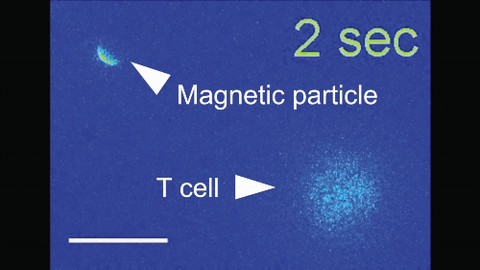
6800: Magnetic Janus particle activating a T cell
A Janus particle being used to activate a T cell, a type of immune cell. A Janus particle is a specialized microparticle with different physical properties on its surface, and this one is coated with nickel on one hemisphere and anti-CD3 antibodies (light blue) on the other. The nickel enables the Janus particle to be moved using a magnet, and the antibodies bind to the T cell and activate it. The T cell in this video was loaded with calcium-sensitive dye to visualize calcium influx, which indicates activation. The intensity of calcium influx was color coded so that warmer color indicates higher intensity. Being able to control Janus particles with simple magnets is a step toward controlling individual cells’ activities without complex magnetic devices.
More details can be found in the Angewandte Chemie paper “Remote control of T cell activation using magnetic Janus particles” by Lee et al. This video was captured using epi-fluorescence microscopy.
Related to video 6801.
More details can be found in the Angewandte Chemie paper “Remote control of T cell activation using magnetic Janus particles” by Lee et al. This video was captured using epi-fluorescence microscopy.
Related to video 6801.
Yan Yu, Indiana University, Bloomington.
View Media

6751: Petri dish containing C. elegans
6751: Petri dish containing C. elegans
This Petri dish contains microscopic roundworms called Caenorhabditis elegans. Researchers used these particular worms to study how C. elegans senses the color of light in its environment.
H. Robert Horvitz and Dipon Ghosh, Massachusetts Institute of Technology.
View Media
2473: Glowing glycans
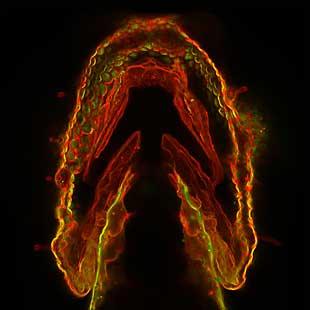
2473: Glowing glycans
Sugars light up the cells in this jaw of a 3-day-old zebrafish embryo and highlight a scientific first: labeling and tracking the movements of sugar chains called glycans in a living organism. Here, recently produced glycans (red) are on the cell surface while those made earlier in development (green) have migrated into the cells. In some areas, old and new glycans mingle (yellow). A better understanding of such traffic patterns could shed light on how organisms develop and may uncover markers for disease, such as cancer. Featured in the May 21, 2008 of Biomedical Beat.
Carolyn Bertozzi, University of California, Berkeley
View Media
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
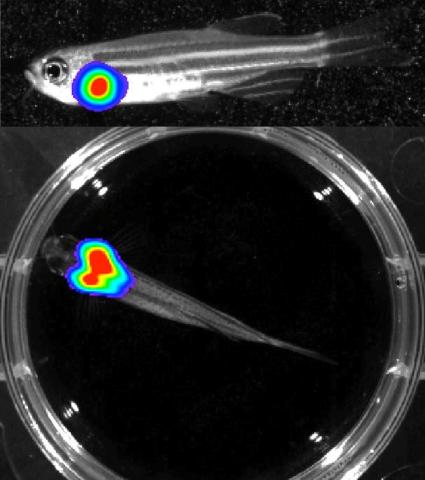
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression. This is the lateral and overhead (Bottom) view.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
6557: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 24 hours
6557: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 24 hours
Floral pattern emerging as two bacterial species, motile Acinetobacter baylyi and non-motile Escherichia coli (green), are grown together for 24 hours on 0.75% agar surface from a small inoculum in the center of a Petri dish.
See 6553 for a photo of this process at 48 hours on 1% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6550 for a video of this process.
See 6553 for a photo of this process at 48 hours on 1% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6550 for a video of this process.
L. Xiong et al, eLife 2020;9: e48885
View Media
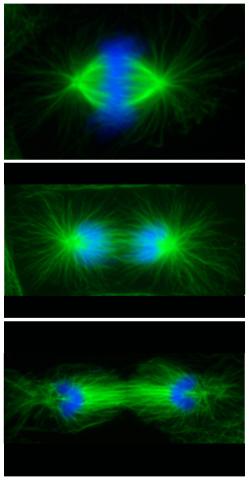
3442: Cell division phases in Xenopus frog cells
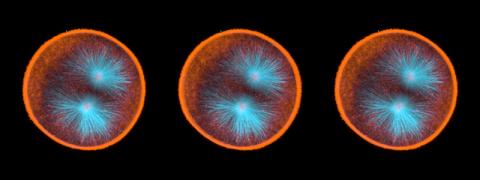
3442: Cell division phases in Xenopus frog cells
These images show three stages of cell division in Xenopus XL177 cells, which are derived from tadpole epithelial cells. They are (from top): metaphase, anaphase and telophase. The microtubules are green and the chromosomes are blue. Related to 3443.
Claire Walczak, who took them while working as a postdoc in the laboratory of Timothy Mitchison
View Media
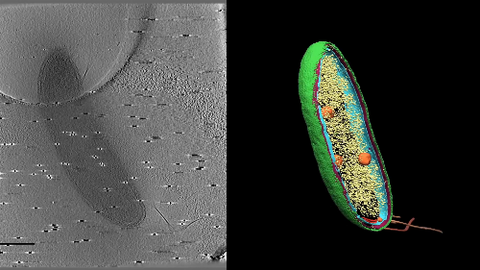
6569: Cryo-electron tomography of a Caulobacter bacterium
6569: Cryo-electron tomography of a Caulobacter bacterium
3D image of Caulobacter bacterium with various components highlighted: cell membranes (red and blue), protein shell (green), protein factories known as ribosomes (yellow), and storage granules (orange).
Peter Dahlberg, Stanford University.
View Media

2305: Beaded bacteriophage
2305: Beaded bacteriophage
This sculpture made of purple and clear glass beads depicts bacteriophage Phi174, a virus that infects bacteria. It rests on a surface that portrays an adaptive landscape, a conceptual visualization. The ridges represent the gene combinations associated with the greatest fitness levels of the virus, as measured by how quickly the virus can reproduce itself. Phi174 is an important model system for studies of viral evolution because its genome can readily be sequenced as it evolves under defined laboratory conditions.
Holly Wichman, University of Idaho. (Surface by A. Johnston; photo by J. Palmersheim)
View Media
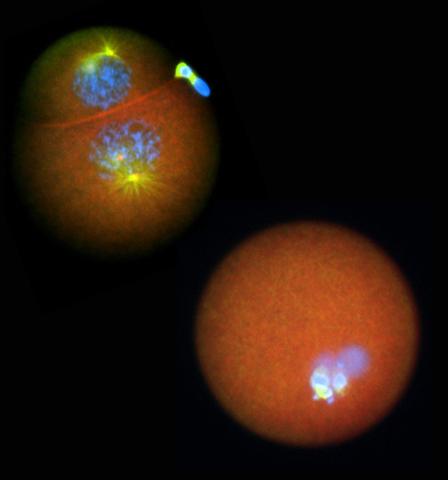
2762: Nucleolinus
2762: Nucleolinus
The nucleolinus is a cellular compartment that has been a lonely bystander in scientific endeavors. Although it's found in a range of species, its function has been mysterious—mainly because the structure is hard to visualize. An August 2010 study showed that the nucleolinus is crucial for cell division. When researchers zapped the structure with a laser, an egg cell didn't complete division. When the oocyte was fertilized after laser microsurgery (bottom right), the resulting zygote didn't form vital cell division structures (blue and yellow).
Mary Anne Alliegro, Marine Biological Laboratory
View Media
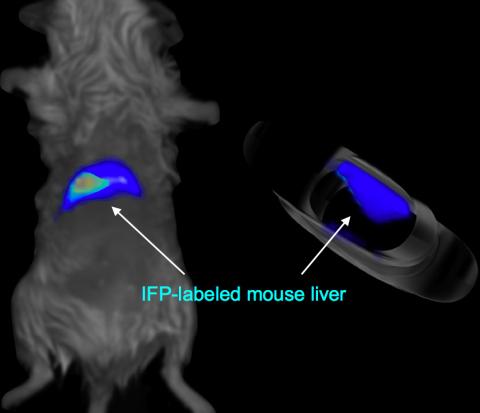
2601: Mouse liver labeled with fluorescent probe
2601: Mouse liver labeled with fluorescent probe
A mouse liver glows after being tagged with specially designed infrared-fluorescent protein (IFP). Since its discovery in 1962, green fluorescent protein (GFP) has become an invaluable resource in biomedical imaging. But because of its short wavelength, the light that makes GFP glow doesn't penetrate far in whole animals. So University of California, San Diego cell biologist Roger Tsien--who shared the 2008 Nobel Prize in chemistry for groundbreaking work with GFP--made infrared-fluorescent proteins (IFPs) that shine under longer-wavelength light, allowing whole-body imaging in small animals.
Xiaokun Shu, University of California, San Diego
View Media

2362: Automated crystal screening system
2362: Automated crystal screening system
Automated crystal screening systems such as the one shown here are becoming a common feature at synchrotron and other facilities where high-throughput crystal structure determination is being carried out. These systems rapidly screen samples to identify the best candidates for further study.
Southeast Collaboratory for Structural Genomics
View Media

2579: Bottles of warfarin
2579: Bottles of warfarin
In 2007, the FDA modified warfarin's label to indicate that genetic makeup may affect patient response to the drug. The widely used blood thinner is sold under the brand name Coumadin®. Scientists involved in the NIH Pharmacogenetics Research Network are investigating whether genetic information can be used to improve optimal dosage prediction for patients.
Alisa Machalek, NIGMS/NIH
View Media
2763: Fused, dicentric chromosomes
2763: Fused, dicentric chromosomes
This fused chromosome has two functional centromeres, shown as two sets of red and green dots. Centromeres are DNA/protein complexes that are key to splitting the chromosomes evenly during cell division. When dicentric chromosomes like this one are formed in a person, fertility problems or other difficulties may arise. Normal chromosomes carrying a single centromere (one set of red and green dots) are also visible in this image.
Beth A. Sullivan, Duke University
View Media
2754: Myosin V binding to actin
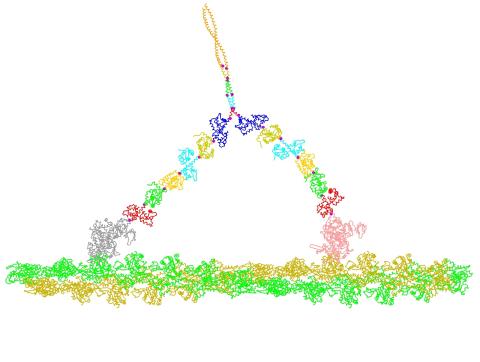
2754: Myosin V binding to actin
This simulation of myosin V binding to actin was created using the software tool Protein Mechanica. With Protein Mechanica, researchers can construct models using information from a variety of sources: crystallography, cryo-EM, secondary structure descriptions, as well as user-defined solid shapes, such as spheres and cylinders. The goal is to enable experimentalists to quickly and easily simulate how different parts of a molecule interact.
Simbios, NIH Center for Biomedical Computation at Stanford
View Media

1311: Housekeeping cell illustration
6486: CRISPR Illustration Frame 2
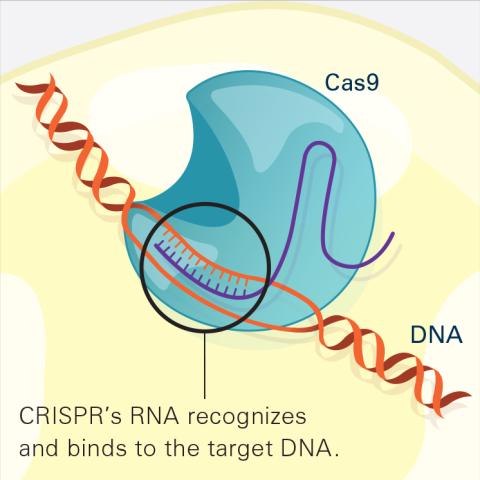
6486: CRISPR Illustration Frame 2
This illustration shows, in simplified terms, how the CRISPR-Cas9 system can be used as a gene-editing tool. The CRISPR system has two components joined together: a finely tuned targeting device (a small strand of RNA programmed to look for a specific DNA sequence) and a strong cutting device (an enzyme called Cas9 that can cut through a double strand of DNA). In this frame (2 of 4), the CRISPR machine locates the target DNA sequence once inserted into a cell.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
National Institute of General Medical Sciences.
View Media
2566: Haplotypes
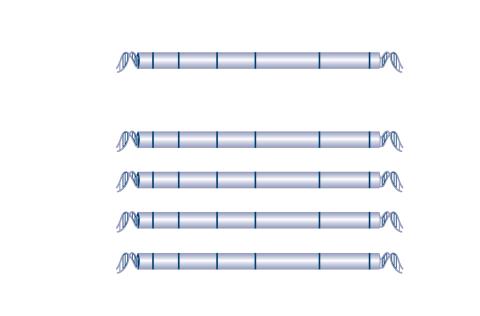
2566: Haplotypes
Haplotypes are combinations of gene variants that are likely to be inherited together within the same chromosomal region. In this example, an original haplotype (top) evolved over time to create three newer haplotypes that each differ by a few nucleotides (red). See image 2567 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
2794: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 01
2794: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 01
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
2795: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 02
2795: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 02
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
2550: Introns
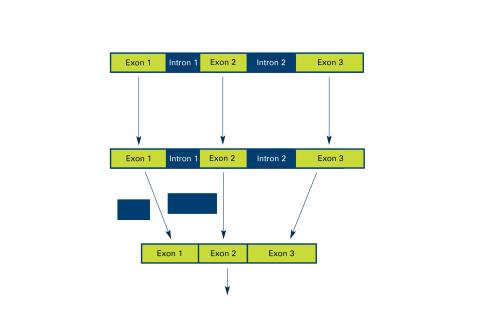
2550: Introns
Genes are often interrupted by stretches of DNA (introns, blue) that do not contain instructions for making a protein. The DNA segments that do contain protein-making instructions are known as exons (green). See image 2551 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
7021: Single-cell “radios” image
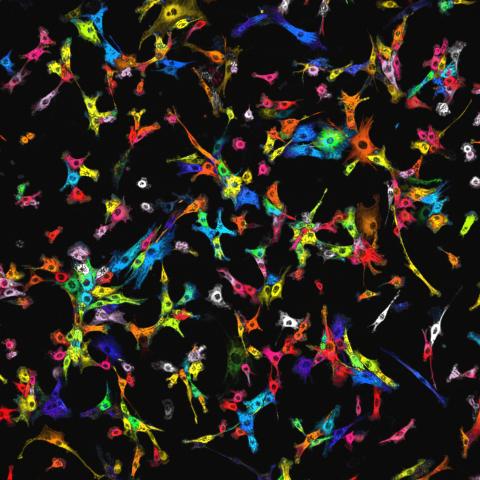
7021: Single-cell “radios” image
Individual cells are color-coded based on their identity and signaling activity using a protein circuit technology developed by the Coyle Lab. Just as a radio allows you to listen to an individual frequency, this technology allows researchers to tune into the specific “radio station” of each cell through genetically encoded proteins from a bacterial system called MinDE. The proteins generate an oscillating fluorescent signal that transmits information about cell shape, state, and identity that can be decoded using digital signal processing tools originally designed for telecommunications. The approach allows researchers to look at the dynamics of a single cell in the presence of many other cells.
Related to video 7022.
Related to video 7022.
Scott Coyle, University of Wisconsin-Madison.
View Media
3756: Protective membrane and membrane proteins of the dengue virus visualized with cryo-EM
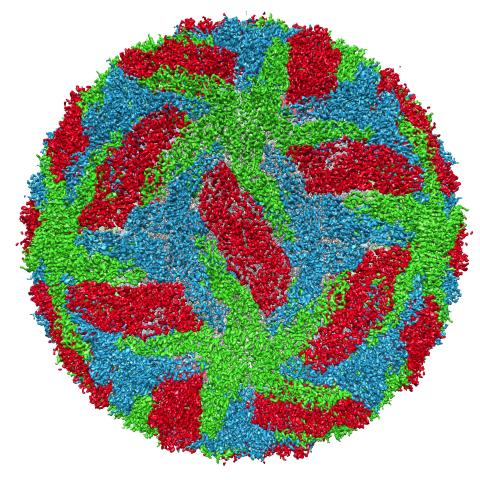
3756: Protective membrane and membrane proteins of the dengue virus visualized with cryo-EM
Dengue virus is a mosquito-borne illness that infects millions of people in the tropics and subtropics each year. Like many viruses, dengue is enclosed by a protective membrane. The proteins that span this membrane play an important role in the life cycle of the virus. Scientists used cryo-EM to determine the structure of a dengue virus at a 3.5-angstrom resolution to reveal how the membrane proteins undergo major structural changes as the virus matures and infects a host. For more on cryo-EM see the blog post Cryo-Electron Microscopy Reveals Molecules in Ever Greater Detail. You can watch a rotating view of the dengue virus surface structure in video 3748.
Hong Zhou, UCLA
View Media

3266: Biopixels
3266: Biopixels
Bioengineers were able to coax bacteria to blink in unison on microfluidic chips. This image shows a small chip with about 500 blinking bacterial colonies or biopixels. Related to images 3265 and 3268. From a UC San Diego news release, "Researchers create living 'neon signs' composed of millions of glowing bacteria."
Jeff Hasty Lab, UC San Diego
View Media
6580: Bacterial nanowire model
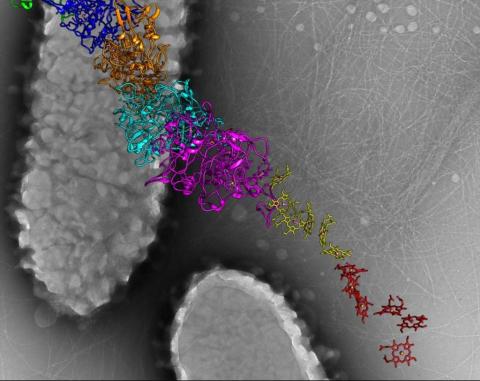
6580: Bacterial nanowire model
A model of a Geobacter sulfurreducens nanowire created from cryo-electron microscopy images. The bacterium conducts electricity through these nanowires, which are made up of protein and iron-containing molecules.
Edward Egelman, University of Virginia.
View Media

2531: Drugs enter skin
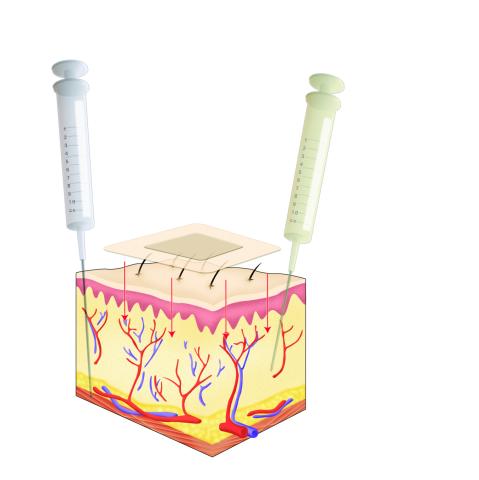
2531: Drugs enter skin
Drugs enter different layers of skin via intramuscular, subcutaneous, or transdermal delivery methods. See image 2532 for a labeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
2337: Beta2-adrenergic receptor protein
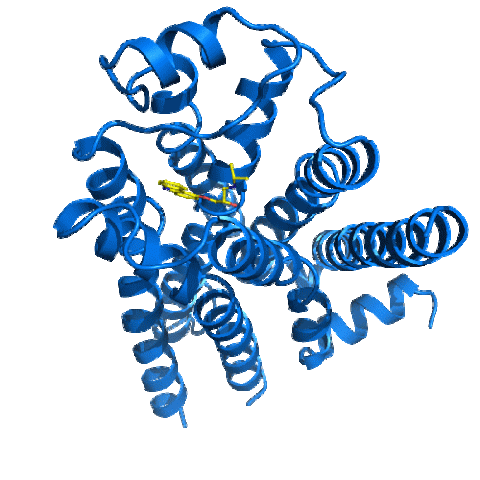
2337: Beta2-adrenergic receptor protein
Crystal structure of the beta2-adrenergic receptor protein. This is the first known structure of a human G protein-coupled receptor, a large family of proteins that control critical bodily functions and the action of about half of today's pharmaceuticals. Featured as one of the November 2007 Protein Structure Initiative Structures of the Month.
The Stevens Laboratory, The Scripps Research Institute
View Media
5886: Mouse Brain Cross Section
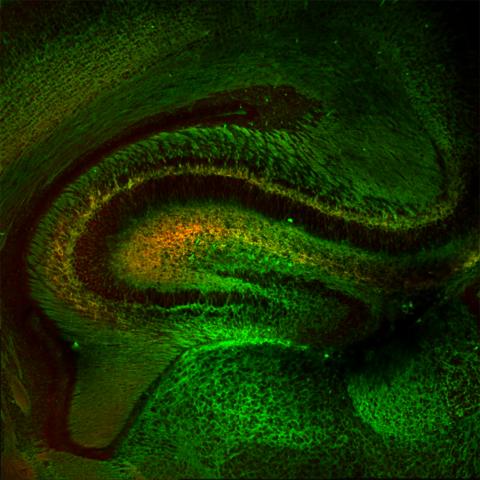
5886: Mouse Brain Cross Section
The brain sections are treated with fluorescent antibodies specific to a particular protein and visualized using serial electron microscopy (SEM).
Anton Maximov, The Scripps Research Institute, La Jolla, CA
View Media

5872: Mouse retina close-up
5872: Mouse retina close-up
Keunyoung ("Christine") Kim National Center for Microscopy and Imaging Research (NCMIR)
View Media
3408: Kluyveromyces polysporus Argonaute bound to guide RNA
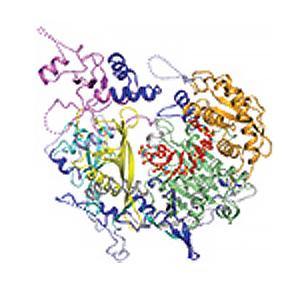
3408: Kluyveromyces polysporus Argonaute bound to guide RNA
A segment of siRNA, shown in red, guides a "slicer" protein called Argonaute (multi-colored twists and corkscrews) to the target RNA molecules.
Kotaro Nakanishi and David Weinberg, Massachusetts Institute of Technology
View Media
2363: PSI: from genes to structures
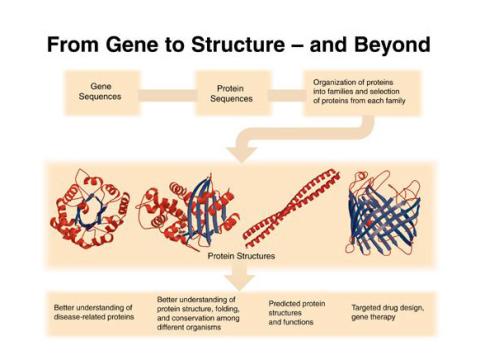
2363: PSI: from genes to structures
The goal of the Protein Structure Initiative (PSI) is to determine the three-dimensional shapes of a wide range of proteins by solving the structures of representative members of each protein family found in nature. The collection of structures should serve as a valuable resource for biomedical research scientists.
National Institute of General Medical Sciences
View Media
6964: Crawling cell
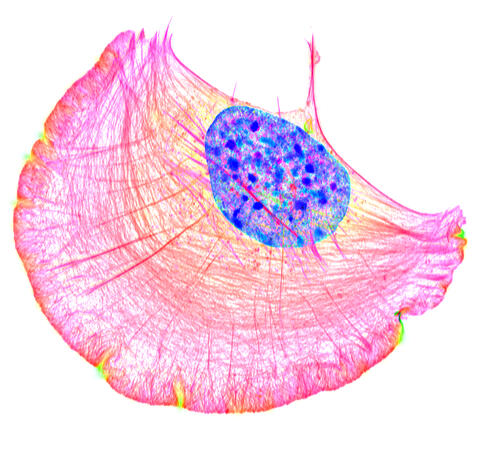
6964: Crawling cell
A crawling cell with DNA shown in blue and actin filaments, which are a major component of the cytoskeleton, visible in pink. Actin filaments help enable cells to crawl. This image was captured using structured illumination microscopy.
Dylan T. Burnette, Vanderbilt University School of Medicine.
View Media
2608: Human embryonic stem cells
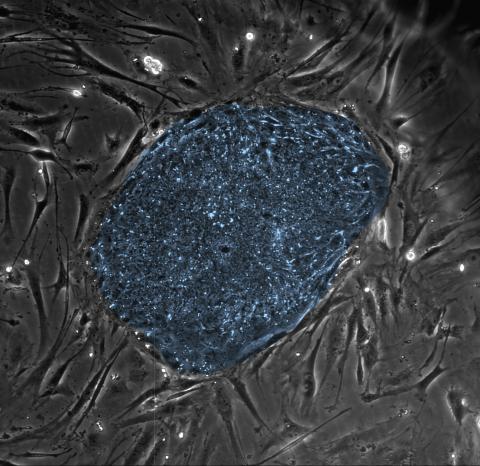
2608: Human embryonic stem cells
The center cluster of cells, colored blue, shows a colony of human embryonic stem cells. These cells, which arise at the earliest stages of development, are capable of differentiating into any of the 220 types of cells in the human body and can provide access to cells for basic research and potential therapies. This image is from the lab of the University of Wisconsin-Madison's James Thomson.
James Thomson, University of Wisconsin-Madison
View Media

3423: White Poppy (cropped)
3423: White Poppy (cropped)
A cropped image of a white poppy. View poppy uncropped here 3424.
Judy Coyle, Donald Danforth Plant Science Center
View Media
2474: Dinosaur evolutionary tree
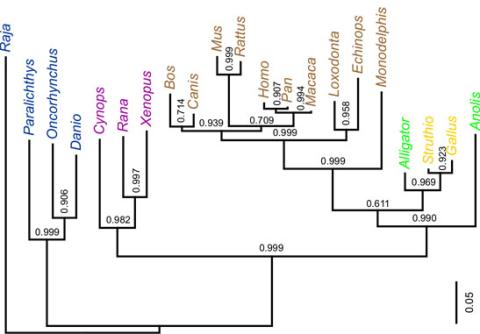
2474: Dinosaur evolutionary tree
Analysis of 68 million-year-old collagen molecule fragments preserved in a T. rex femur confirmed what paleontologists have said for decades: Dinosaurs are close relatives of chickens, ostriches, and to a lesser extent, alligators. A Harvard University research team, including NIGMS-supported postdoctoral research fellow Chris Organ, used sophisticated statistical and computational tools to compare the ancient protein to ones from 21 living species. Because evolutionary processes produce similarities across species, the methods and results may help illuminate other areas of the evolutionary tree. Featured in the May 21, 2008 Biomedical Beat.
Chris Organ, Harvard University
View Media
2490: Cascade reaction promoted by water
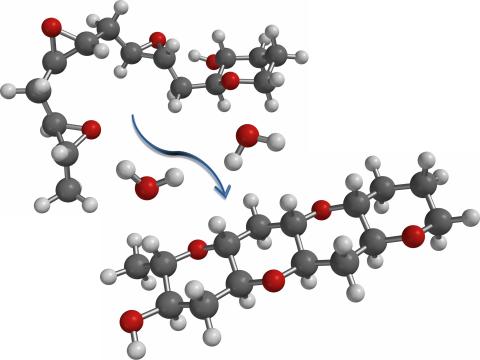
2490: Cascade reaction promoted by water
This illustration of an epoxide-opening cascade promoted by water emulates the proposed biosynthesis of some of the Red Tide toxins.
Tim Jamison, Massachusetts Institute of Technology
View Media
6809: Fruit fly egg ooplasmic streaming
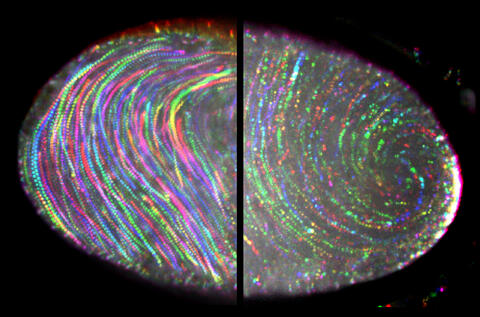
6809: Fruit fly egg ooplasmic streaming
Two fruit fly (Drosophila melanogaster) egg cells, one on each side of the central black line. The colorful swirls show the circular movement of cytoplasm—called ooplasmic streaming—that occurs in late egg cell development in wild-type (right) and mutant (left) oocytes. This image was captured using confocal microscopy.
More information on the research that produced this image can be found in the Journal of Cell Biology paper “Ooplasmic flow cooperates with transport and anchorage in Drosophila oocyte posterior determination” by Lu et al.
More information on the research that produced this image can be found in the Journal of Cell Biology paper “Ooplasmic flow cooperates with transport and anchorage in Drosophila oocyte posterior determination” by Lu et al.
Vladimir I. Gelfand, Feinberg School of Medicine, Northwestern University.
View Media
7017: The nascent juvenile light organ of the Hawaiian bobtail squid
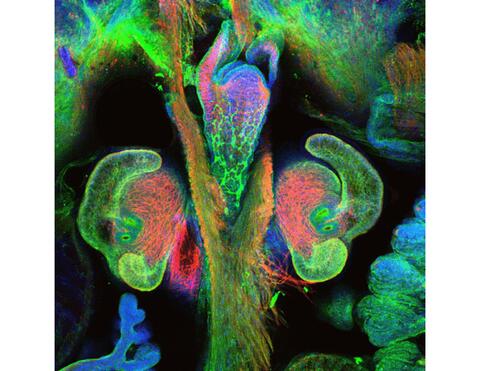
7017: The nascent juvenile light organ of the Hawaiian bobtail squid
A light organ (~0.5 mm across) of a Hawaiian bobtail squid, Euprymna scolopes, with different tissues are stained various colors. The two pairs of ciliated appendages, or “arms,” on the sides of the organ move Vibrio fischeri bacterial cells closer to the two sets of three pores (two seen in this image) at the base of the arms that each lead to an interior crypt. This image was taken using a confocal fluorescence microscope.
Related to images 7016, 7018, 7019, and 7020.
Related to images 7016, 7018, 7019, and 7020.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
3437: Network diagram of genes, cellular components and processes (labeled)
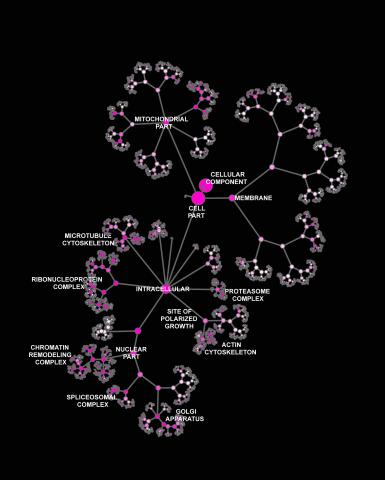
3437: Network diagram of genes, cellular components and processes (labeled)
This image shows the hierarchical ontology of genes, cellular components and processes derived from large genomic datasets. From Dutkowski et al. A gene ontology inferred from molecular networks Nat Biotechnol. 2013 Jan;31(1):38-45. Related to 3436.
Janusz Dutkowski and Trey Ideker, University of California, San Diego
View Media

2737: Cytoscape network diagram 1
2737: Cytoscape network diagram 1
Molecular biologists are increasingly relying on bioinformatics software to visualize molecular interaction networks and to integrate these networks with data such as gene expression profiles. Related to 2749.
Keiichiro Ono, Trey Ideker lab, University of California, San Diego
View Media
1160: Vibrio bacteria
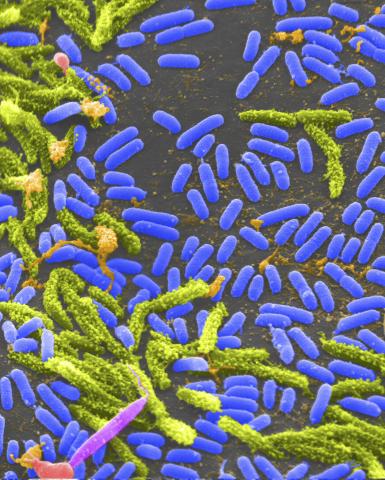
1160: Vibrio bacteria
Vibrio, a type (genus) of rod-shaped bacteria. Some Vibrio species cause cholera in humans.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media

3526: 800 MHz NMR magnet
3526: 800 MHz NMR magnet
Scientists use nuclear magnetic spectroscopy (NMR) to determine the detailed, 3D structures of molecules.
Asokan Anbanandam, University of Kansas
View Media
3436: Network diagram of genes, cellular components and processes (unlabeled)
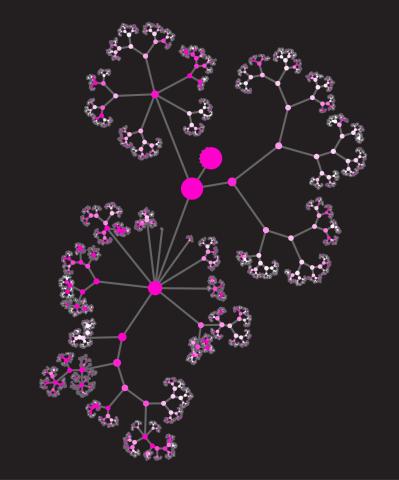
3436: Network diagram of genes, cellular components and processes (unlabeled)
This image shows the hierarchical ontology of genes, cellular components and processes derived from large genomic datasets. From Dutkowski et al. A gene ontology inferred from molecular networks Nat Biotechnol. 2013 Jan;31(1):38-45. Related to 3437.
Janusz Dutkowski and Trey Ideker
View Media
2767: Research mentor and student
2767: Research mentor and student
A research mentor (Lori Eidson) and student (Nina Waldron, on the microscope) were 2009 members of the BRAIN (Behavioral Research Advancements In Neuroscience) program at Georgia State University in Atlanta. This program is an undergraduate summer research experience funded in part by NIGMS.
Elizabeth Weaver, Georgia State University
View Media