Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.

6903: Young squids
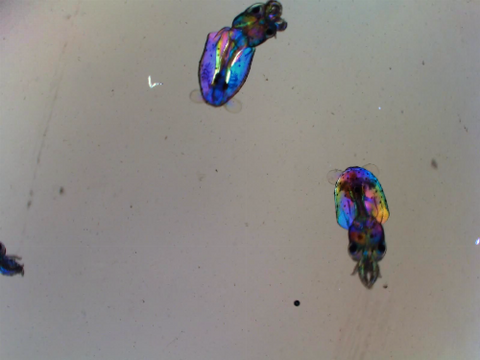
6903: Young squids
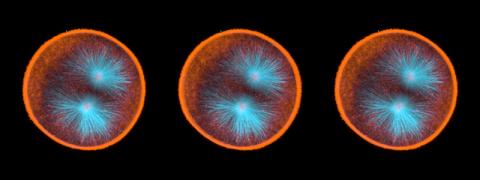
Real-time movie of young squids. Squids are often used as research organisms due to having the largest nervous system of any invertebrate, complex behaviors like instantaneous camouflage, and other unique traits.
This video was taken with polychromatic polarization microscope, as described in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak. The color is generated by interaction of white polarized light with the squid’s transparent soft tissue. The tissue works as a living tunable spectral filter, and the transmission band depends on the molecular orientation. When the young squid is moving, the tissue orientation changes, and its color shifts accordingly.
This video was taken with polychromatic polarization microscope, as described in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak. The color is generated by interaction of white polarized light with the squid’s transparent soft tissue. The tissue works as a living tunable spectral filter, and the transmission band depends on the molecular orientation. When the young squid is moving, the tissue orientation changes, and its color shifts accordingly.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media
3403: Disrupted vascular development in frog embryos
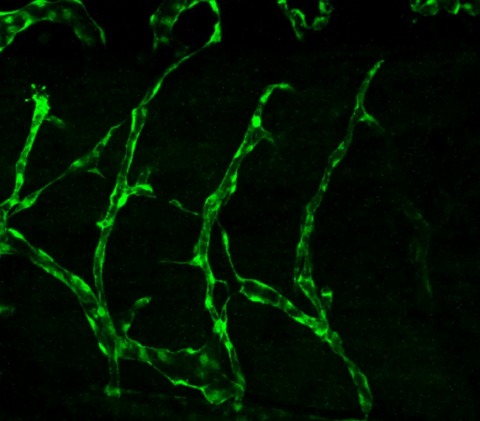
3403: Disrupted vascular development in frog embryos
Disassembly of vasculature in kdr:GFP frogs following addition of 250 µM TBZ. Related to images 3404 and 3505.
Hye Ji Cha, University of Texas at Austin
View Media
2321: Microtubule breakdown
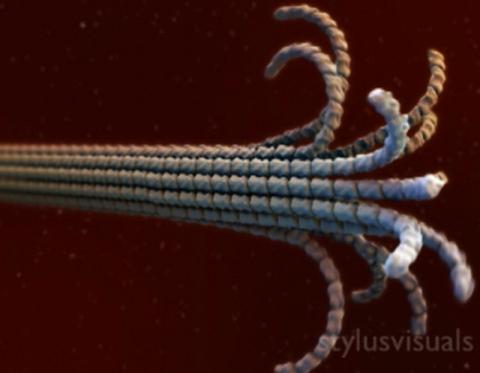
2321: Microtubule breakdown
Like a building supported by a steel frame, a cell contains its own sturdy internal scaffolding made up of proteins, including microtubules. Researchers studying snapshots of microtubules have proposed a model for how these structural elements shorten and lengthen, allowing a cell to move, divide, or change shape. This picture shows an intermediate step in the model: Smaller building blocks called tubulins peel back from the microtubule in thin strips. Knowing the operations of the internal scaffolding will enhance our basic understanding of cellular processes.
Eva Nogales, University of California, Berkeley
View Media
2693: Fruit fly in the pink
2693: Fruit fly in the pink
Fruit flies are a common model organism for basic medical research.
Crabtree + Company
View Media
3334: Four timepoints in gastrulation
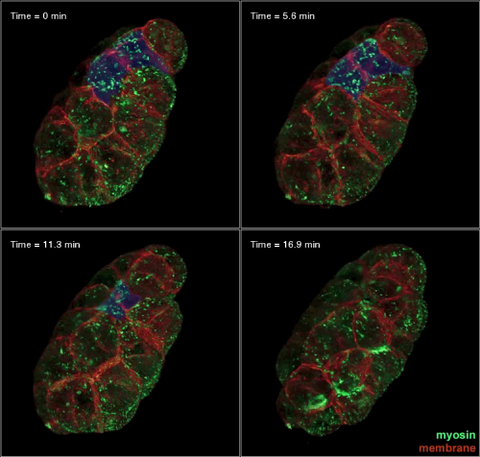
3334: Four timepoints in gastrulation
It has been said that gastrulation is the most important event in a person's life. This part of early embryonic development transforms a simple ball of cells and begins to define cell fate and the body axis. In a study published in Science magazine, NIGMS grantee Bob Goldstein and his research group studied how contractions of actomyosin filaments in C. elegans and Drosophila embryos lead to dramatic rearrangements of cell and embryonic structure. In these images, myosin (green) and plasma membrane (red) are highlighted at four timepoints in gastrulation in the roundworm C. elegans. The blue highlights in the top three frames show how cells are internalized, and the site of closure around the involuting cells is marked with an arrow in the last frame. See related image 3297.
Bob Goldstein, University of North Carolina, Chapel Hill
View Media
5795: Mouse cerebellum
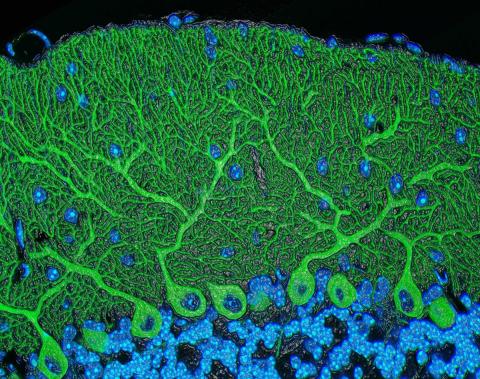
5795: Mouse cerebellum
The cerebellum is the brain's locomotion control center. Found at the base of your brain, the cerebellum is a single layer of tissue with deep folds like an accordion. People with damage to this region of the brain often have difficulty with balance, coordination and fine motor skills.
This image of a mouse cerebellum is part of a collection of such images in different colors and at different levels of magnification from the National Center for Microscopy and Imaging Research (NCMIR). Related to image 5800.
This image of a mouse cerebellum is part of a collection of such images in different colors and at different levels of magnification from the National Center for Microscopy and Imaging Research (NCMIR). Related to image 5800.
National Center for Microscopy and Imaging Research (NCMIR)
View Media

2335: Virtual snow world
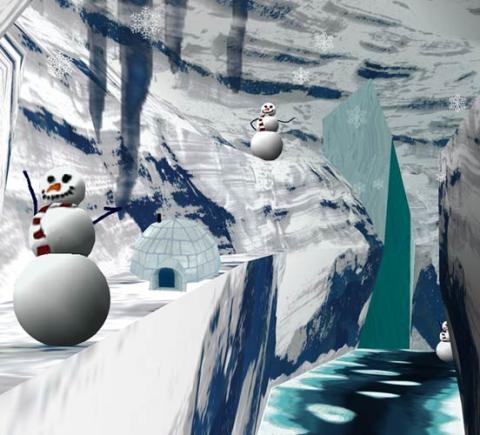
2335: Virtual snow world
Glide across an icy canyon, where you see smiling snowmen and waddling penguins. Toss a snowball, hear it smash against an igloo, and then watch it explode in bright colors. Psychologists David Patterson and Hunter Hoffman of the University of Washington in Seattle developed this virtual "Snow World" to test whether immersing someone in a pretend reality could ease pain during burn treatment and other medical procedures. They found that people fully engaged in the virtual reality experience reported 60 percent less pain. The technology offers a promising way to manage pain.
David Patterson and Hunter Hoffmann, University of Washington
View Media
3411: O2 reacting with a flavin-dependent enzyme
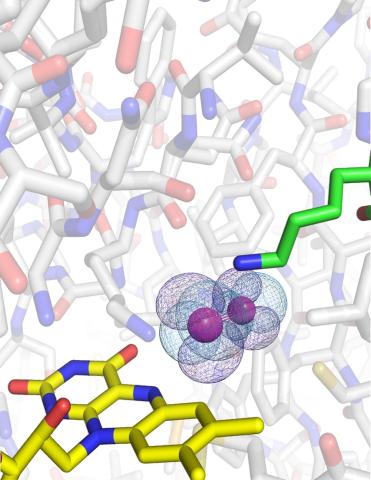
3411: O2 reacting with a flavin-dependent enzyme
Department of Biological Chemistry, University of Michigan
View Media

6549: The Structure of Cilia’s Doublet Microtubules
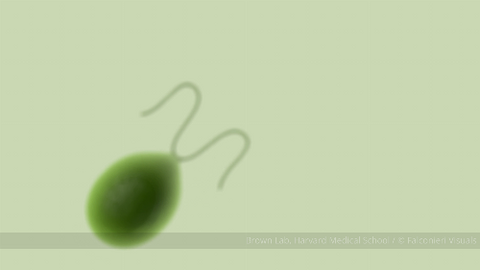
6549: The Structure of Cilia’s Doublet Microtubules
Cilia (cilium in singular) are complex molecular machines found on many of our cells. One component of cilia is the doublet microtubule, a major part of cilia’s skeletons that give them support and shape. This animated video illustrates the structure of doublet microtubules, which contain 451 protein chains that were mapped using cryo-electron microscopy. Image can be found here 6548.
Brown Lab, Harvard Medical School and Veronica Falconieri Hays
View Media
3742: Confocal microscopy of perineuronal nets in the brain 2
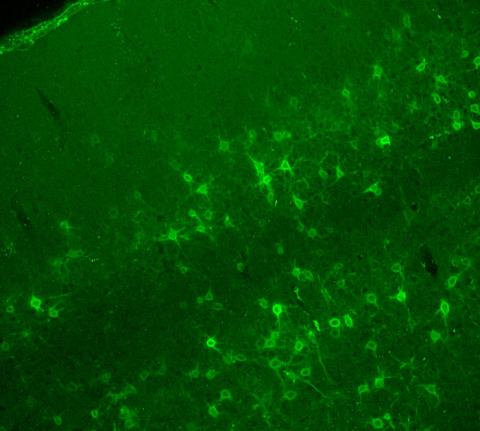
3742: Confocal microscopy of perineuronal nets in the brain 2
The photo shows a confocal microscopy image of perineuronal nets (PNNs), which are specialized extracellular matrix (ECM) structures in the brain. The PNN surrounds some nerve cells in brain regions including the cortex, hippocampus and thalamus. Researchers study the PNN to investigate their involvement stabilizing the extracellular environment and forming nets around nerve cells and synapses in the brain. Abnormalities in the PNNs have been linked to a variety of disorders, including epilepsy and schizophrenia, and they limit a process called neural plasticity in which new nerve connections are formed. To visualize the PNNs, researchers labeled them with Wisteria floribunda agglutinin (WFA)-fluorescein. Related to image 3741.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media

2305: Beaded bacteriophage
2305: Beaded bacteriophage
This sculpture made of purple and clear glass beads depicts bacteriophage Phi174, a virus that infects bacteria. It rests on a surface that portrays an adaptive landscape, a conceptual visualization. The ridges represent the gene combinations associated with the greatest fitness levels of the virus, as measured by how quickly the virus can reproduce itself. Phi174 is an important model system for studies of viral evolution because its genome can readily be sequenced as it evolves under defined laboratory conditions.
Holly Wichman, University of Idaho. (Surface by A. Johnston; photo by J. Palmersheim)
View Media

3272: Ear hair cells derived from embryonic stem cells
3272: Ear hair cells derived from embryonic stem cells
Mouse embryonic stem cells matured into this bundle of hair cells similar to the ones that transmit sound in the ear. These cells could one day be transplanted as a therapy for some forms of deafness, or they could be used to screen drugs to treat deafness. The hairs are shown at 23,000 times magnification via scanning electron microscopy. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Stefen Heller, Stanford University, via CIRM
View Media
2758: Cross section of a Drosophila melanogaster pupa
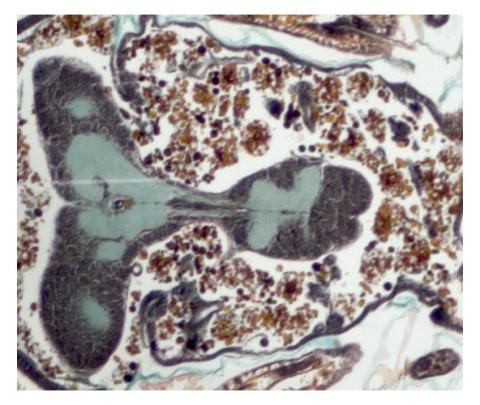
2758: Cross section of a Drosophila melanogaster pupa
This photograph shows a magnified view of a Drosophila melanogaster pupa in cross section. Compare this normal pupa to one that lacks an important receptor, shown in image 2759.
Christina McPhee and Eric Baehrecke, University of Massachusetts Medical School
View Media
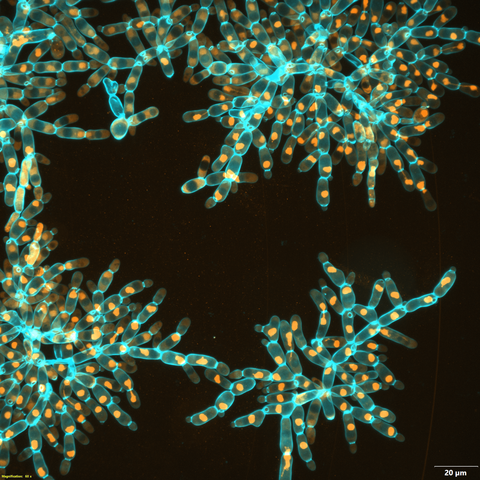
6971: Snowflake yeast 3
6971: Snowflake yeast 3
Multicellular yeast called snowflake yeast that researchers created through many generations of directed evolution from unicellular yeast. Here, the researchers visualized nuclei in orange to help them study changes in how the yeast cells divided. Cell walls are shown in blue. This image was captured using spinning disk confocal microscopy.
Related to images 6969 and 6970.
Related to images 6969 and 6970.
William Ratcliff, Georgia Institute of Technology.
View Media
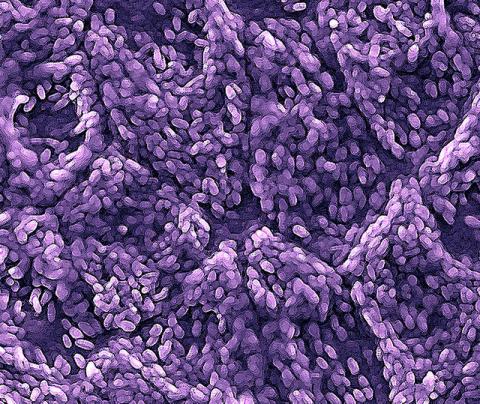
3286: Retinal pigment epithelium derived from human ES cells
3286: Retinal pigment epithelium derived from human ES cells
This color-enhanced image is a scanning electron microscope image of retinal pigment epithelial (RPE) cells derived from human embryonic stem cells. The cells are remarkably similar to normal RPE cells, growing in a hexagonal shape in a single, well-defined layer. This kind of retinal cell is responsible for macular degeneration, the most common cause of blindness. Image and caption information courtesy of the California Institute for Regenerative Medicine. Related to image 3287.
David Hinton lab, University of Southern California, via CIRM
View Media
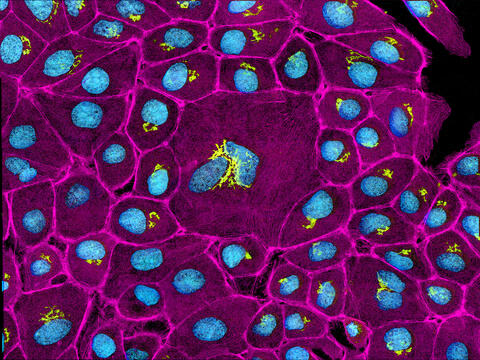
3647: Epithelial cells
3647: Epithelial cells
This image mostly shows normal cultured epithelial cells expressing green fluorescent protein targeted to the Golgi apparatus (yellow-green) and stained for actin (magenta) and DNA (cyan). The middle cell is an abnormal large multinucleated cell. All the cells in this image have a Golgi but not all are expressing the targeted recombinant fluorescent protein.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
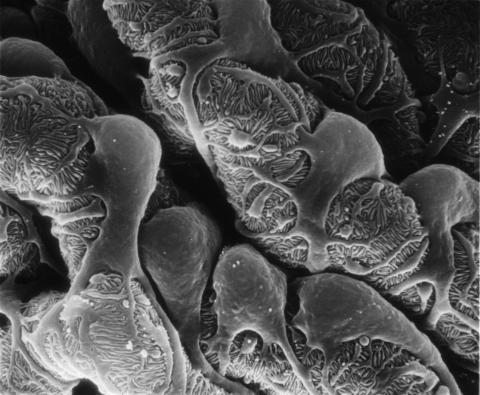
3565: Podocytes from a chronically diseased kidney
3565: Podocytes from a chronically diseased kidney
This scanning electron microscope (SEM) image shows podocytes--cells in the kidney that play a vital role in filtering waste from the bloodstream--from a patient with chronic kidney disease. This image first appeared in Princeton Journal Watch on October 4, 2013.
Olga Troyanskaya, Princeton University and Matthias Kretzler, University of Michigan
View Media

3327: Diversity oriented synthesis: generating skeletal diversity using folding processes
3327: Diversity oriented synthesis: generating skeletal diversity using folding processes
This 1 1/2-minute video animation was produced for chemical biologist Stuart Schreiber's lab page. The animation shows how diverse chemical structures can be produced in the lab.
Eric Keller
View Media

2531: Drugs enter skin
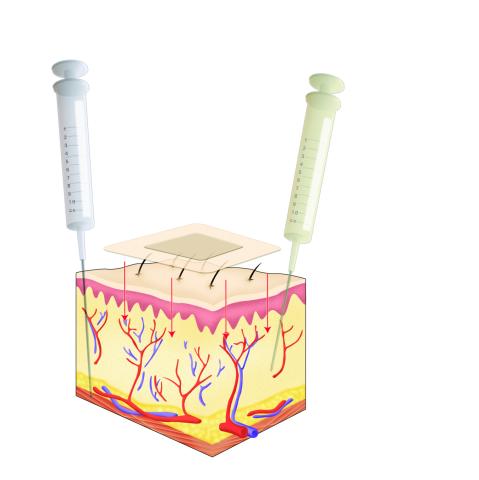
2531: Drugs enter skin
Drugs enter different layers of skin via intramuscular, subcutaneous, or transdermal delivery methods. See image 2532 for a labeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
6583: Closeup of fluorescent C. elegans showing muscle and ribosomal protein
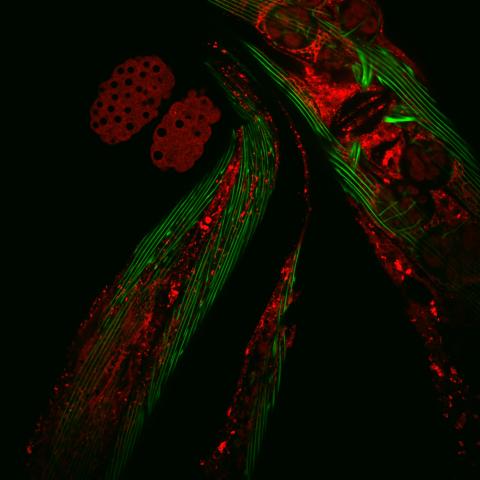
6583: Closeup of fluorescent C. elegans showing muscle and ribosomal protein
Closeup of C. elegans, tiny roundworms, with a ribosomal protein glowing red and muscle fibers glowing green. Researchers used these worms to study a molecular pathway that affects aging. The ribosomal protein is involved in protein translation and may play a role in dietary restriction-induced longevity. Image created using confocal microscopy.
View single roundworm here 6581.
View group of roundworms here 6582.
View single roundworm here 6581.
View group of roundworms here 6582.
Jarod Rollins, Mount Desert Island Biological Laboratory.
View Media
3754: Circadian rhythm neurons in the fruit fly brain
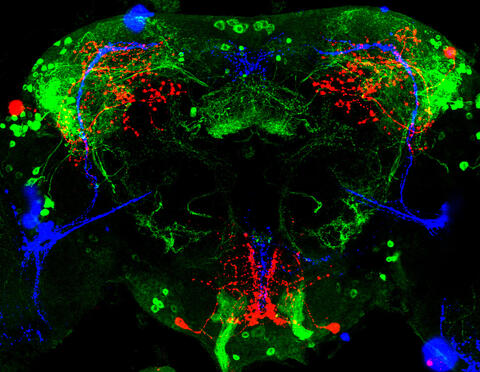
3754: Circadian rhythm neurons in the fruit fly brain
Some nerve cells (neurons) in the brain keep track of the daily cycle. This time-keeping mechanism, called the circadian clock, is found in all animals including us. The circadian clock controls our daily activities such as sleep and wakefulness. Researchers are interested in finding the neuron circuits involved in this time keeping and how the information about daily time in the brain is relayed to the rest of the body. In this image of a brain of the fruit fly Drosophila the time-of-day information flowing through the brain has been visualized by staining the neurons involved: clock neurons (shown in blue) function as "pacemakers" by communicating with neurons that produce a short protein called leucokinin (LK) (red), which, in turn, relays the time signal to other neurons, called LK-R neurons (green). This signaling cascade set in motion by the pacemaker neurons helps synchronize the fly's daily activity with the 24-hour cycle. To learn more about what scientists have found out about circadian pacemaker neurons in the fruit fly see this news release by New York University. This work was featured in the Biomedical Beat blog post Cool Image: A Circadian Circuit.
Justin Blau, New York University
View Media
6851: Himastatin, 360-degree view
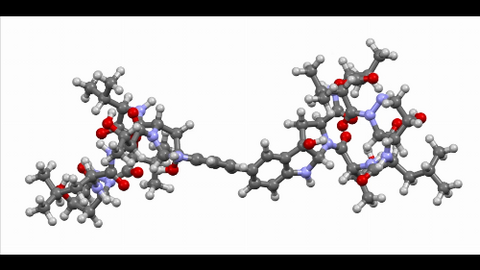
6851: Himastatin, 360-degree view
A 360-degree view of the molecule himastatin, which was first isolated from the bacterium Streptomyces himastatinicus. Himastatin shows antibiotic activity. The researchers who created this video developed a new, more concise way to synthesize himastatin so it can be studied more easily.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
2484: RNA Polymerase II
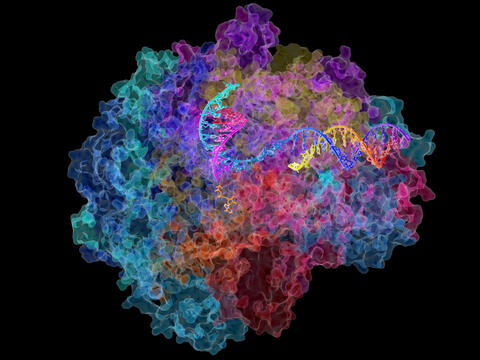
2484: RNA Polymerase II
NIGMS-funded researchers led by Roger Kornberg solved the structure of RNA polymerase II. This is the enzyme in mammalian cells that catalyzes the transcription of DNA into messenger RNA, the molecule that in turn dictates the order of amino acids in proteins. For his work on the mechanisms of mammalian transcription, Kornberg received the Nobel Prize in Chemistry in 2006.
David Bushnell, Ken Westover and Roger Kornberg, Stanford University
View Media

7011: Hawaiian bobtail squid
7011: Hawaiian bobtail squid
An adult Hawaiian bobtail squid, Euprymna scolopes, swimming next to a submerged hand.
Related to image 7010 and video 7012.
Related to image 7010 and video 7012.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
2426: Zinc finger
2426: Zinc finger
The structure of a gene-regulating zinc finger protein bound to DNA.
Jeremy M. Berg, National Institute of General Medical Sciences
View Media
1082: Natcher Building 02
1082: Natcher Building 02
NIGMS staff are located in the Natcher Building on the NIH campus.
Alisa Machalek, National Institute of General Medical Sciences
View Media
3268: Fluorescent E. coli bacteria
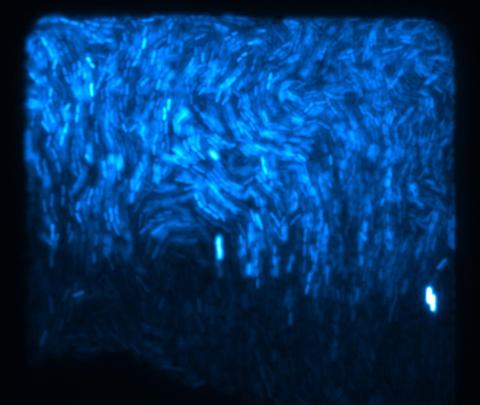
3268: Fluorescent E. coli bacteria
Bioengineers were able to coax bacteria to blink in unison on microfluidic chips. They called each blinking bacterial colony a biopixel. Thousands of fluorescent E. coli bacteria, shown here, make up a biopixel. Related to images 3265 and 3266. From a UC San Diego news release, "Researchers create living 'neon signs' composed of millions of glowing bacteria."
Jeff Hasty Lab, UC San Diego
View Media
2304: Bacteria working to eat
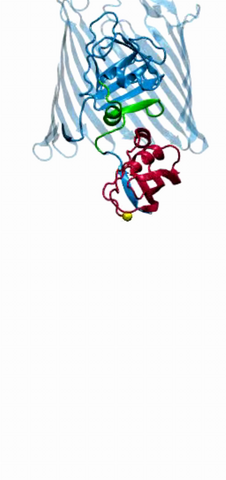
2304: Bacteria working to eat
Gram-negative bacteria perform molecular acrobatics just to eat. Because they're encased by two membranes, they must haul nutrients across both. To test one theory of how the bacteria manage this feat, researchers used computer simulations of two proteins involved in importing vitamin B12. Here, the protein (red) anchored in the inner membrane of bacteria tugs on a much larger protein (green and blue) in the outer membrane. Part of the larger protein unwinds, creating a pore through which the vitamin can pass.
Emad Tajkhorshid, University of Illinois at Urbana-Champaign
View Media
2432: ARTS triggers apoptosis
2432: ARTS triggers apoptosis
Cell showing overproduction of the ARTS protein (red). ARTS triggers apoptosis, as shown by the activation of caspase-3 (green) a key tool in the cell's destruction. The nucleus is shown in blue. Image is featured in October 2015 Biomedical Beat blog post Cool Images: A Halloween-Inspired Cell Collection.
Hermann Steller, Rockefeller University
View Media

3718: A Bacillus subtilis biofilm grown in a Petri dish
3718: A Bacillus subtilis biofilm grown in a Petri dish
Bacterial biofilms are tightly knit communities of bacterial cells growing on, for example, solid surfaces, such as in water pipes or on teeth. Here, cells of the bacterium Bacillus subtilis have formed a biofilm in a laboratory culture. Researchers have discovered that the bacterial cells in a biofilm communicate with each other through electrical signals via specialized potassium ion channels to share resources, such as nutrients, with each other. This insight may help scientists to improve sanitation systems to prevent biofilms, which often resist common treatments, from forming and to develop better medicines to combat bacterial infections. See the Biomedical Beat blog post Bacterial Biofilms: A Charged Environment for more information.
Gürol Süel, UCSD
View Media

5843: Color coding of the Drosophila brain - video
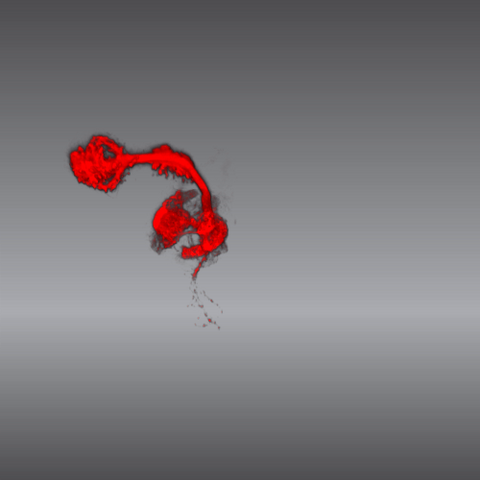
5843: Color coding of the Drosophila brain - video
This video results from a research project to visualize which regions of the adult fruit fly (Drosophila) brain derive from each neural stem cell. First, researchers collected several thousand fruit fly larvae and fluorescently stained a random stem cell in the brain of each. The idea was to create a population of larvae in which each of the 100 or so neural stem cells was labeled at least once. When the larvae grew to adults, the researchers examined the flies’ brains using confocal microscopy. With this technique, the part of a fly’s brain that derived from a single, labeled stem cell “lights up.” The scientists photographed each brain and digitally colorized its lit-up area. By combining thousands of such photos, they created a three-dimensional, color-coded map that shows which part of the Drosophila brain comes from each of its ~100 neural stem cells. In other words, each colored region shows which neurons are the progeny or “clones” of a single stem cell. This work established a hierarchical structure as well as nomenclature for the neurons in the Drosophila brain. Further research will relate functions to structures of the brain.
Related to images 5838 and 5868.
Related to images 5838 and 5868.
Yong Wan from Charles Hansen’s lab, University of Utah. Data preparation and visualization by Masayoshi Ito in the lab of Kei Ito, University of Tokyo.
View Media
3491: Kinesin moves cellular cargo
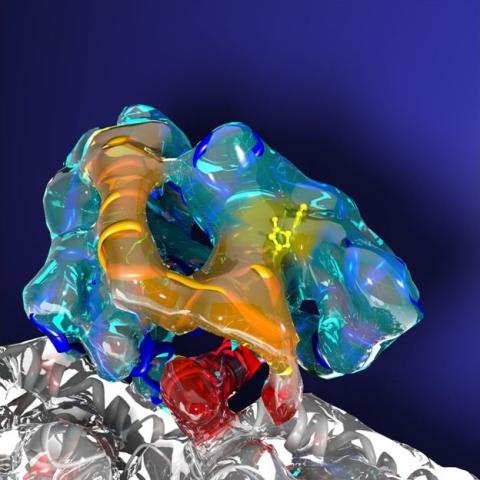
3491: Kinesin moves cellular cargo
A protein called kinesin (blue) is in charge of moving cargo around inside cells and helping them divide. It's powered by biological fuel called ATP (bright yellow) as it scoots along tube-like cellular tracks called microtubules (gray).
Charles Sindelar, Yale University
View Media
3725: Fluorescent microscopy of kidney tissue--close-up
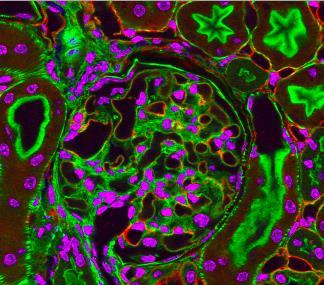
3725: Fluorescent microscopy of kidney tissue--close-up
This photograph of kidney tissue, taken using fluorescent light microscopy, shows a close-up view of part of image 3723. Kidneys filter the blood, removing waste and excessive fluid, which is excreted in urine. The filtration system is made up of components that include glomeruli (for example, the round structure taking up much of the image's center is a glomerulus) and tubules (seen in cross-section here with their inner lining stained green). Related to image 3675 .
Tom Deerinck , National Center for Microscopy and Imaging Research
View Media
5770: EM of yeast cell division
5770: EM of yeast cell division
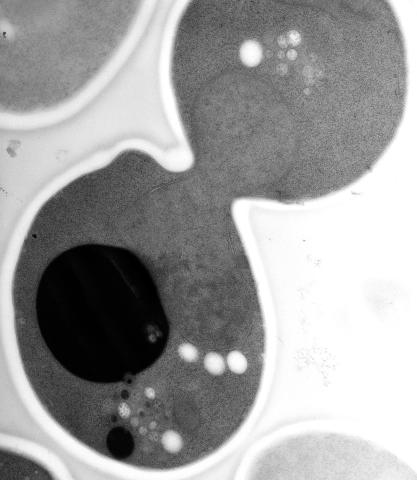
Cell division is an incredibly coordinated process. It not only ensures that the new cells formed during this event have a full set of chromosomes, but also that they are endowed with all the cellular materials, including proteins, lipids and small functional compartments called organelles, that are required for normal cell activity. This proper apportioning of essential cell ingredients helps each cell get off to a running start.
This image shows an electron microscopy (EM) thin section taken at 10,000x magnification of a dividing yeast cell over-expressing the protein ubiquitin, which is involved in protein degradation and recycling. The picture features mother and daughter endosome accumulations (small organelles with internal vesicles), a darkly stained vacuole and a dividing nucleus in close contact with a cadre of lipid droplets (unstained spherical bodies). Other dynamic events are also visible, such as spindle microtubules in the nucleus and endocytic pits at the plasma membrane.
These extensive details were revealed thanks to a preservation method involving high-pressure freezing, freeze-substitution and Lowicryl HM20 embedding.
This image shows an electron microscopy (EM) thin section taken at 10,000x magnification of a dividing yeast cell over-expressing the protein ubiquitin, which is involved in protein degradation and recycling. The picture features mother and daughter endosome accumulations (small organelles with internal vesicles), a darkly stained vacuole and a dividing nucleus in close contact with a cadre of lipid droplets (unstained spherical bodies). Other dynamic events are also visible, such as spindle microtubules in the nucleus and endocytic pits at the plasma membrane.
These extensive details were revealed thanks to a preservation method involving high-pressure freezing, freeze-substitution and Lowicryl HM20 embedding.
Matthew West and Greg Odorizzi, University of Colorado
View Media
6794: Yeast cells with Fimbrin Fim1
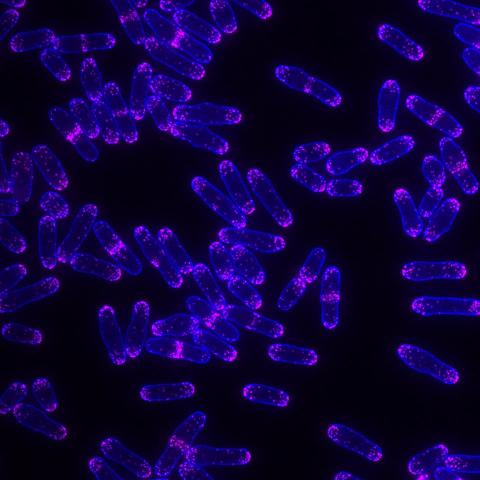
6794: Yeast cells with Fimbrin Fim1
Yeast cells with the protein Fimbrin Fim1 shown in magenta. This protein plays a role in cell division. This image was captured using wide-field microscopy with deconvolution.
Related to images 6791, 6792, 6793, 6797, 6798, and videos 6795 and 6796.
Related to images 6791, 6792, 6793, 6797, 6798, and videos 6795 and 6796.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media
3417: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 5
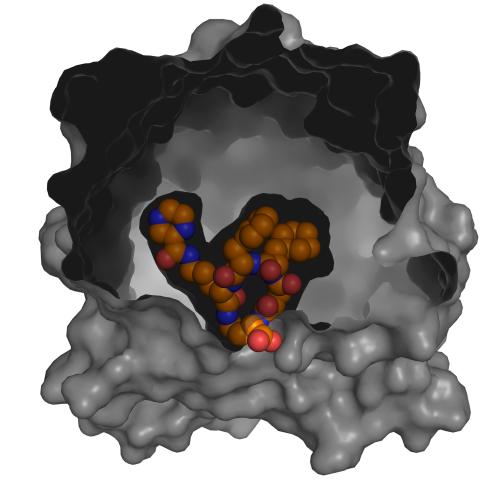
3417: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 5
X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor. Related to images 3413, 3414, 3415, 3416, 3418, and 3419.
Markus A. Seeliger, Stony Brook University Medical School and David R. Liu, Harvard University
View Media
6795: Dividing yeast cells with nuclear envelopes and spindle pole bodies
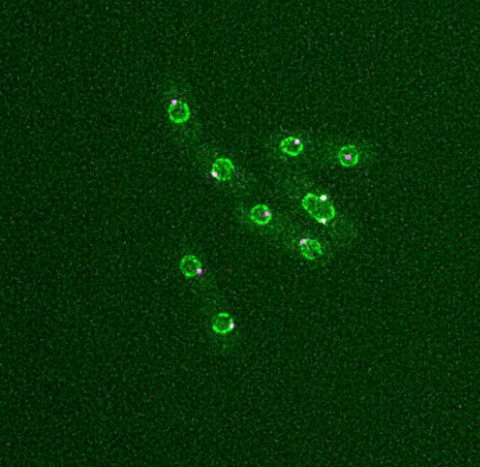
6795: Dividing yeast cells with nuclear envelopes and spindle pole bodies
Time-lapse video of yeast cells undergoing cell division. Nuclear envelopes are shown in green, and spindle pole bodies, which help pull apart copied genetic information, are shown in magenta. This video was captured using wide-field microscopy with deconvolution.
Related to images 6791, 6792, 6793, 6794, 6797, 6798, and video 6796.
Related to images 6791, 6792, 6793, 6794, 6797, 6798, and video 6796.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media

2747: Cell division with late aligning chromosomes
2747: Cell division with late aligning chromosomes
This video shows an instance of abnormal mitosis where chromosomes are late to align. The video demonstrates the spindle checkpoint in action: just one unaligned chromosome can delay anaphase and the completion of mitosis. The cells shown are S3 tissue cultured cells from Xenopus laevis, African clawed frog.
Gary Gorbsky, Oklahoma Medical Research Foundation
View Media
2746: Active site of sulfite oxidase
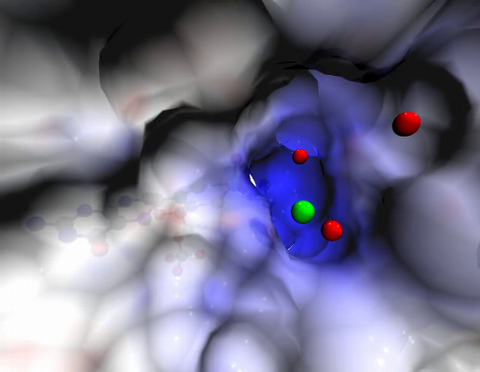
2746: Active site of sulfite oxidase
Sulfite oxidase is an enzyme that is essential for normal neurological development in children. This video shows the active site of the enzyme and its molybdenum cofactor visible as a faint ball-and-stick representation buried within the protein. The positively charged channel (blue) at the active site contains a chloride ion (green) and three water molecules (red). As the protein oscillates, one can see directly down the positively charged channel. At the bottom is the molybdenum atom of the active site (light blue) and its oxo group (red) that is transferred to sulfite to form sulfate in the catalytic reaction.
John Enemark, University of Arizona
View Media
3686: Hippocampal neuron from rodent brain
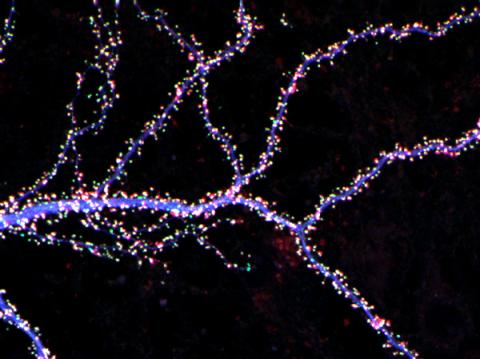
3686: Hippocampal neuron from rodent brain
Hippocampal neuron from rodent brain with dendrites shown in blue. The hundreds of tiny magenta, green and white dots are the dendritic spines of excitatory synapses.
Shelley Halpain, UC San Diego
View Media
3442: Cell division phases in Xenopus frog cells
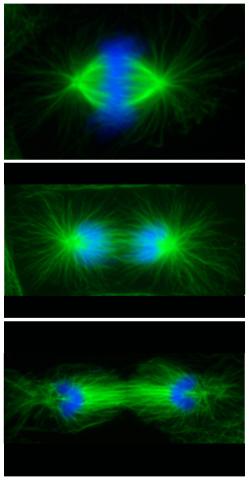
3442: Cell division phases in Xenopus frog cells
These images show three stages of cell division in Xenopus XL177 cells, which are derived from tadpole epithelial cells. They are (from top): metaphase, anaphase and telophase. The microtubules are green and the chromosomes are blue. Related to 3443.
Claire Walczak, who took them while working as a postdoc in the laboratory of Timothy Mitchison
View Media
2456: Z rings in bacterial division
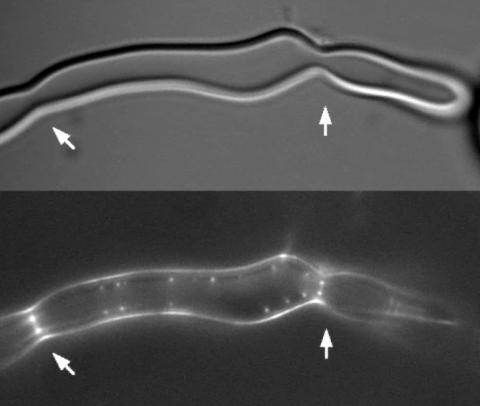
2456: Z rings in bacterial division
Lab-made liposomes contract where Z rings have gathered together and the constriction forces are greatest (arrows). The top picture shows a liposome, and the bottom picture shows fluorescence from Z rings (arrows) inside the same liposome simultaneously.
Masaki Osawa, Duke University
View Media

2455: Golden gene chips
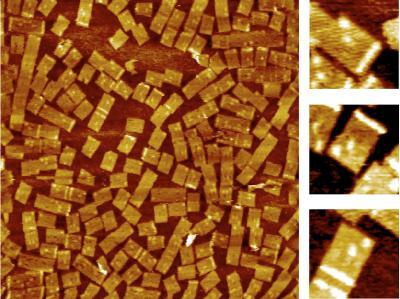
2455: Golden gene chips
A team of chemists and physicists used nanotechnology and DNA's ability to self-assemble with matching RNA to create a new kind of chip for measuring gene activity. When RNA of a gene of interest binds to a DNA tile (gold squares), it creates a raised surface (white areas) that can be detected by a powerful microscope. This nanochip approach offers manufacturing and usage advantages over existing gene chips and is a key step toward detecting gene activity in a single cell. Featured in the February 20, 2008, issue of Biomedical Beat.
Hao Yan and Yonggang Ke, Arizona State University
View Media
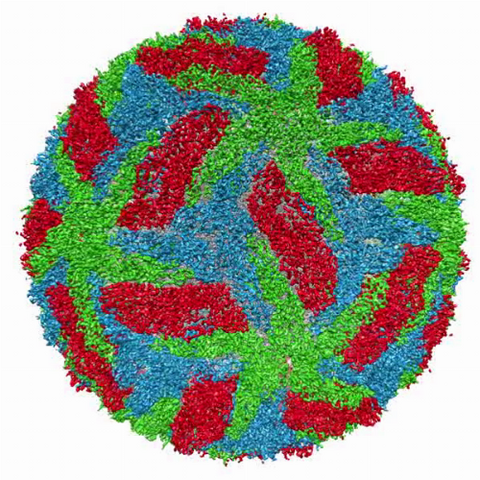
3748: Cryo-electron microscopy of the dengue virus showing protective membrane and membrane proteins
3748: Cryo-electron microscopy of the dengue virus showing protective membrane and membrane proteins
Dengue virus is a mosquito-borne illness that infects millions of people in the tropics and subtropics each year. Like many viruses, dengue is enclosed by a protective membrane. The proteins that span this membrane play an important role in the life cycle of the virus. Scientists used cryo-EM to determine the structure of a dengue virus at a 3.5-angstrom resolution to reveal how the membrane proteins undergo major structural changes as the virus matures and infects a host. For more on cryo-EM see the blog post Cryo-Electron Microscopy Reveals Molecules in Ever Greater Detail. Related to image 3756.
Hong Zhou, UCLA
View Media
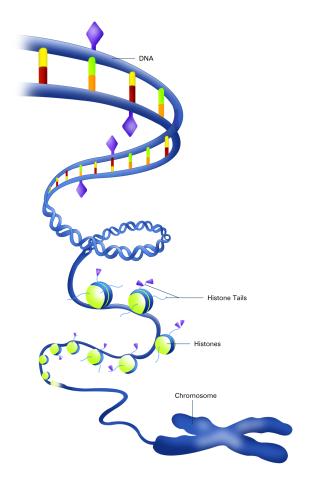
2563: Epigenetic code (with labels)
2563: Epigenetic code (with labels)
The "epigenetic code" controls gene activity with chemical tags that mark DNA (purple diamonds) and the "tails" of histone proteins (purple triangles). These markings help determine whether genes will be transcribed by RNA polymerase. Genes hidden from access to RNA polymerase are not expressed. See image 2562 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
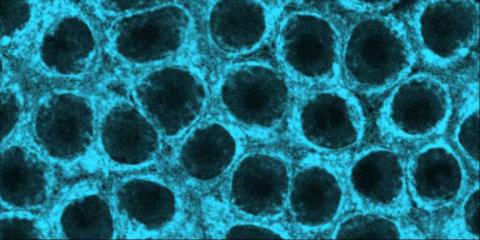
2324: Movements of myosin
2324: Movements of myosin
Inside the fertilized egg cell of a fruit fly, we see a type of myosin (related to the protein that helps muscles contract) made to glow by attaching a fluorescent protein. After fertilization, the myosin proteins are distributed relatively evenly near the surface of the embryo. The proteins temporarily vanish each time the cells' nuclei--initially buried deep in the cytoplasm--divide. When the multiplying nuclei move to the surface, they shift the myosin, producing darkened holes. The glowing myosin proteins then gather, contract, and start separating the nuclei into their own compartments.
Victoria Foe, University of Washington
View Media
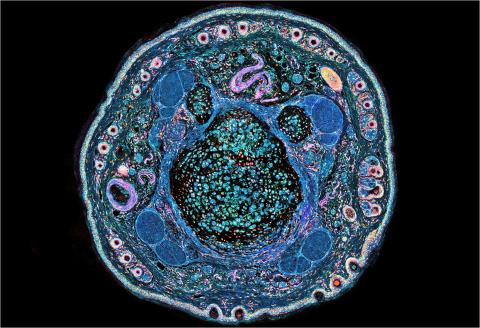
3395: NCMIR mouse tail
3395: NCMIR mouse tail
Stained cross section of a mouse tail.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
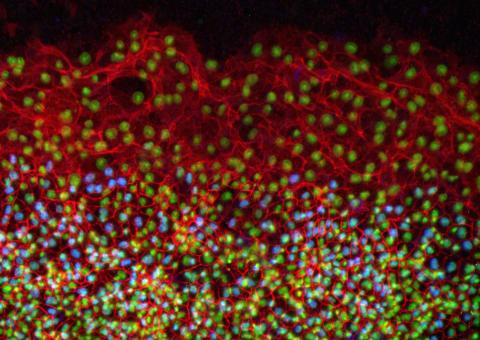
2808: Cell proliferation in a quail embryo
2808: Cell proliferation in a quail embryo
Image showing that the edge zone (top of image) of the quail embryo shows no proliferating cells (cyan), unlike the interior zone (bottom of image). Non-proliferating cell nuclei are labeled green. This image was obtained as part of a study to understand cell migration in embryos. More specifically, cell proliferation at the edge of the embryo was studied by examining the cellular uptake of a chemical compound called BrDU, which incorporates into the DNA during the S-phase of the cell cycle. Here, the cells that are positive for BrDU uptake are labeled in cyan, while other non-proliferating cell nuclei are labeled green. Notice that the vast majority of BrDU+ cells are located far away from the edge, indicating that edge cells are mostly non-proliferating. An NIGMS grant to Professor Garcia was used to purchase the confocal microscope that collected this image. Related to image 2807 and video 2809.
Andrés Garcia, Georgia Tech
View Media