Clearinghouse for Modules to Enhance Biomedical Research Workforce Training
The free training modules on this site are provided by the National Institutes of Health (NIH) Institutes and Centers and NIH grantees, including R25 Training Modules for Enhancing Biomedical Research Workforce Training award recipients. This site is managed by the National Institute of General Medical Sciences (NIGMS). Please contact us with any questions or comments.
The external links on this webpage provide additional information that is consistent with the intended purpose of this site. NIGMS cannot attest to the accuracy or accessibility of a nonfederal site.
Training Modules Table of Contents
- NIH Rigor and Reproducibility Training Modules
NIH Office of the Director - Let's Experiment: A Guide for Scientists Working at the Bench
iBiology - Improving Reproducibility in Research
Indiana University School of Medicine - Sex as a Biological Variable: A Primer
NIH Office of Research on Women’s Health - Addressing Sex as a Biological Variable in Preclinical Research
Cohen Veterans Bioscience - Course on Pragmatic and Group-Randomized Trials in Public Health and Medicine
NIH Office of Disease Prevention - Statistical Topics for Reproducible Animal Research
Indiana University School of Public Health and University of Alabama at Birmingham - Improving Reproducibility of Computational Microbiome Analyses
University of Michigan School of Medicine - edX Course: Principles, Statistical and Computational Tools for Reproducible Science
Harvard School of Public Health - The BD2K Guide to the Fundamentals of Data Science Series
Big Data to Knowledge (BD2K) Initiative - Society for Neuroscience Rigor and Reproducibility Training Webinars
Rutgers University and Michigan State University - Improving reproducibility of recording and pre-processing experimental biomedical data
Colorado State University - Podcasts to Address the Impact of the Culture of Science on Reproducibility of Research
Indiana University at Indianapolis - Education Pathways for Biomedical Data Science
Children’s Hospital of Philadelphia - Foundations of rigorous neuroscience research
Society for Neuroscience - How Much Data?
Dominican University - Responsible Conduct of Research in Biomedical Data Science for Dental, Oral and Craniofacial Research
UCLA School of Dentistry
Training Modules

NIH Rigor and Reproducibility Training Modules
These modules, comprised of videos and accompanying discussion materials, were developed by NIH, and focus on integral aspects of rigor and reproducibility in the research endeavor, such as bias, blinding, and exclusion criteria. The modules are not meant to be comprehensive, but rather are intended as a foundation to build on and a way to stimulate conversations.
Module 1: Lack of Transparency
Module 2: Blinding and Randomization
Module 3: Sample Size, Outliers, and Exclusion Criteria
Module 4: Biological and Technical Replicates

Let's Experiment: A Guide for Scientists Working at the Bench
iBiology, R25 GM116704
"Let's Experiment" is a FREE 6-week online course designed for students and practitioners of experimental biology. Scientists from a variety of backgrounds give concrete steps and advice to help you build a framework for how to design experiments. We use case studies to make the abstract more tangible. In science, there is often no simple right answer. However, you can develop a general approach to experimental design and understand what you are getting into before you begin.
By the end of this course, you will have:
- A detailed plan for your experiment(s) that you can discuss with a mentor.
- A flowchart for how to prioritize experiments.
- A lab notebook template that is so impressively organized, it will make your colleagues envious.
- A framework for doing rigorous research.

Improving Reproducibility in Research
Aaron Carroll, Indiana University School of Medicine, R25 GM116146
In order to promote better training and ensure the reliability and reproducibility of research, we developed a series of webisodes (thematically related online videos) targeted at graduate students, postdoctoral fellows, and beginning investigators that addresses critical features of experimental design and analysis/reporting.
Module 1: Experimental Design Learning Module
The Experimental Design Learning Module focuses on the intricacies of designing research that is robust, with an eye toward making it reproducible. It is comprised of four distinct learning units:
- Replication,
- Randomization,
- Pitfalls with Experimental Design, and
- Measurement.
Each of these learning units has one or more sub-topics that is the subject of an individual webisode.
Module 2: Analysis/Reporting Learning Module
The Analysis/Reporting Learning Module covers the various factors that are critical to writing about research with enough clarity to ensure its reproducibility. Very few researchers are given formal education on how to properly report findings to support reproducibility. It is comprised of three distinct learning units:
- Power and P-values,
- Scientific Writing, and
- The Review Process.
Each of these learning units has one or more sub-topics that is the subject of an individual webisode.

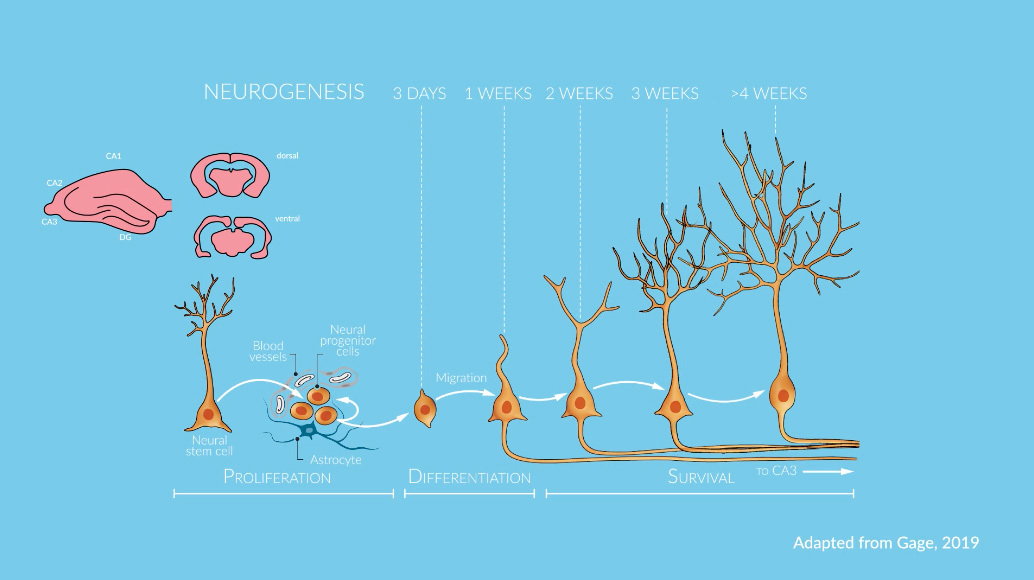
Sex as a Biological Variable: A Primer
Sex as a Biological Variable: A Primer was developed by the NIH Office of Research on Women’s Health with funding support from NIGMS to help the biomedical research community account for and appropriately integrate sex as a biological variable (SABV) across the full spectrum of biomedical sciences. The modules cover the role of SABV in the health of women and men, experimental design, analysis, and research reporting.
Addressing Sex as a Biological Variable in Preclinical Research
The goal of the video training series is to provide practical guidance to preclinical researchers on how to navigate the NIH's 2015 policy on incorporating sex as a biological variable into their current and future research.
NIH Office of Disease Prevention (ODP) Course on Pragmatic and Group-Randomized Trials in Public Health and Medicine
This 7-part online course aims to help researchers design and analyze group-randomized trials (GRTs). It includes video presentations, slide sets, suggested reading materials, and guided activities. The course is presented by ODP's Director, Dr. David M. Murray.

Statistical Topics for Reproducible Animal Research
Andrew W. Brown and David B. Allison, Indiana University School of Public Health-Bloomington; Tapan Mehta and Stephen Watts, University of Alabama at Birmingham, R25 GM116167
Preclinical research involving animal models can be improved when appropriate experimental, analytical, and reporting practices are used. We produced a series of animated vignettes with quantitative experts and laboratory scientists discussing aspects of study design, interpretation, and reporting. Each vignette introduces viewers to key concepts that can stimulate the important conversations needed between quantitative experts and laboratory scientists to enhance rigor, reproducibility, and transparency in pre-clinical research.

Improving Reproducibility of Computational Microbiome Analyses
Patrick Schloss, University of Michigan School of Medicine, R25 GM116149
A series of 14 tutorials on improving the reproducibility of data analysis for those doing microbial ecology research. Although the materials focus on issues in microbiome research, the principles are broadly applicable to other areas of microbiology and science. This series of lessons focus on the importance of command line practices (e.g. bash), scripting languages (e.g. mothur, R), version control (e.g. git), automation (e.g. make), and literate programming (e.g. markdown). These are the tools that are used by a growing number of microbiome researchers to help improve the reproducibility of their research. By completing the activities in the tutorials, you will be listed on the Reproducible Research Tutorial Honor Roll, which provides a certification of your training.

edX Course: Principles, Statistical and Computational Tools for Reproducible Science
Xihong Lin, Harvard School of Public Health, T32GM074897-12S1
Learn skills and tools that support data science and reproducible research to ensure you can trust your research results, reproduce them yourself, and communicate them to others.
This free course covers fundamentals of reproducible science, case studies, data provenance, statistical methods for reproducible science, computational tools for reproducible science, and reproducible reporting science. These concepts are intended to translate to fields throughout the data sciences: physical and life sciences, applied mathematics and statistics, and computing.
Consider this course a survey of best practices that will help you create an environment in which you can easily carry out reproducible research and integrate with similar situations for your collaborators and colleagues.
The BD2K Guide to the Fundamentals of Data Science Series
The Big Data to Knowledge (BD2K) Initiative presents this virtual lecture series on the data science underlying modern biomedical research. Since its beginning in September 2016, the webinar series consists of presentations from experts across the country covering the basics of data management, representation, computation, statistical inference, data modeling, and other topics relevant to "big data" in biomedicine. The webinar series provides essential training suitable for individuals at an introductory overview level. All video presentations from the seminar series are streamed for live viewing, recorded, and posted online for future viewing and reference. These videos are also indexed as part of TCC's Educational Resource Discovery Index (ERuDIte). This webinar series is a collaboration between the TCC (BD2K Training Coordinating Center), the NIH Office of the Associate Director for Data Science, and BD2K Centers Coordination Center (BD2KCCC).
Society for Neuroscience Rigor and Reproducibility Training Webinars
Manny DiCiccio-Bloom, Rutgers University; Cheryl Sisk, Michigan State University, R25 DA041326
Webinar 1: Improving Experimental Rigor and Enhancing Data Reproducibility in Neuroscience
The topics of scientific rigor and data reproducibility have been increasingly covered in the scientific and mainstream media, and they are being addressed by publishers, professional organizations, and funding agencies. This webinar addresses topics of scientific rigor as they pertain to preclinical neuroscience research.
Webinar 1
Webinar 2: Minimizing Bias in Experimental Design and Execution
Investigations into the lack of reproducibility in preclinical research often identify unintended biases in experimental planning and execution. This webinar covers random sampling, blinding, and balancing experiments to avoid sources of bias.
Webinar 2
Webinar 3: Best Practices in Post-Experimental Data Analysis
Proper data handling standards, including appropriate use of statistical tests, are integral to rigorous and reproducible neuroscience research. This webinar covers best practices in post-experimental data analysis.
Webinar 3
Webinar 4: Best Practices in Data Management and Reporting
Efforts to enhance scientific rigor, reproducibility, and robustness critically depend on archiving and retrieving experimental records, protocols, primary data, and subsequent analyses. In this webinar, presenters discuss best practices and challenges for data management and reporting, particularly when dealing with information security and sensitive material; archiving and disclosure of pre- and post-hoc data analytics; and data management on multidisciplinary teams that include collaborators around the globe.
Webinar 4
Webinar 5: Statistical Applications in Neuroscience
How can neuroscientists improve their "statistical thinking" and make full and effective use of their data? This webinar covers common applications of statistics in neuroscience, including the types of research questions statistics are best positioned to address, modeling paradigms, and exploratory data analysis. The presenters also share examples and case studies from their research.
Webinar 5
Webinar 6: Experimental Design to Minimize Systemic Biases: Lessons from Rodent Behavioral Assays and Electrophysiology Studies
Common sources of bias in animal behavior and electrophysiology experiments can be minimized or avoided by following best practices of unbiased experimental design and data analysis and interpretation. In this webinar, presenters discuss experimental design and hypothesis testing for mouse behavioral assays, as well as sampling, interpretational bias, and referencing in in vitro and in vivo electrophysiology recording studies.
Webinar 6
Workshop 7: Tackling Challenges in Scientific Rigor: The (Sometimes) Messy Reality of Science
This webinar explores practical examples of the challenges and solutions in conducting rigorous science from neuroscientists at various career stages. It focuses on development of the interpersonal, scientific, and technical skills needed to address various issues in scientific rigor, such as what to do when you can't replicate a published result, how to get support from a mentor, and how to cope with various career pressures that might affect the quality of your science.
Workshop 7

Improving reproducibility of recording and pre-processing experimental biomedical data
Brooke Anderson, Colorado State University, R25GM132797
These training modules teach laboratory-based biomedical researchers how simple computational reproducibility principles can improve reproducibility at the stages of data recording and pre-processing.

Podcasts to Address the Impact of the Culture of Science on Reproducibility of Research
Aaron Carroll, Indiana University at Indianapolis, R25GM132785
A podcast series exploring the varying opinions on the underlying issues of reproducibility and the culture of science. The podcasts are accompanied by lesson guides that will help learners think critically about the discussions. The lesson guides will also help educators to craft curriculum around the podcast series and host small group discussions about various topics with their students.

Education Pathways for Biomedical Data Science
Jeffrey Pennington, Children's Hospital of Philadelphia, R25GM141501
Modules to equip researchers to perform nimble research with massive datasets that span institutional and disciplinary boundaries. There are 42 data science education modules, as well as publicly available code and documentation related to pedagogical approaches.

Foundations of rigorous neuroscience research
Oswald Steward, Society for Neuroscience, R25NS114922
A comprehensive online rigor and reproducibility toolkit containing over 30 multimedia resources organized in 11 subject areas: ethics, lab management, leadership skills development, outreach, professional development, responsible conduct of research, scientific research, scientific rigor and reproducibility, scientific training, training, and workforce development.

How Much Data?
Robert Calin-Jageman, Dominican University, R25GM132784
An online workshop explaining the importance of sample-size determination and what not to do, and then discusses both planning for power and planning for precision, providing model sample-size plans, tips for optimization, and useful resources. The course consists of short videos, text, and simulations for exploration. The course is provided in Quarto format on GitHub so that elements can easily be re-mixed and adopted for training programs.

Responsible Conduct of Research in Biomedical Data Science for Dental, Oral and Craniofacial Research
Ichiro Nishimura, UCLA School of Dentistry, R25 DE030574
This learning module will introduce general and specific features to improve reproducibility of an investigation using Biomedical Data Science. With the advent of the rapid development of research technologies, the past decade has seen vast accumulation of acquired data on human health and diseases. Biomedical Data Science is an exciting new research discipline to best analyze the massive data sets to discover new knowledge. We envision that a new and diverse generation of biological and quantitative dentist-scientists, oral health researchers, and practicing dentists will be mining these biomedical data to address current challenges to public health and patient care.
There are 5 modules of video instructions:
- Introduction
- Experimental Design
- Laboratory Practices
- Analysis and Reporting
- Culture of Science
