Biophysics
This branch supports studies that apply techniques and principles derived from the physical sciences to examine structures and structure-function relationships in biology. Areas of emphasis in biophysical research include (a) the development and application of physical and theoretical techniques to biological problems from the molecular to cellular level of organization, and (b) the application of engineering science and technology to the development of improved methods of measurement and analysis for physiological and biomedical research. Of interest are new applications of established techniques and modifications to, the modification of existing instrumentation to yield improved resolution, sensitivity, or accuracy. Central problems include the fundamentals of molecular properties and interactions; relationships between sequences and molecular structures, dynamics, and functions; assembly and mechanism of supramolecular structures including cellular membranes, cytoskeleton, and viruses; and discovery of ways to selectively influence biological processes based on these structures. Primary research funded in this branch is from investigator-initiated applications.
Program Areas
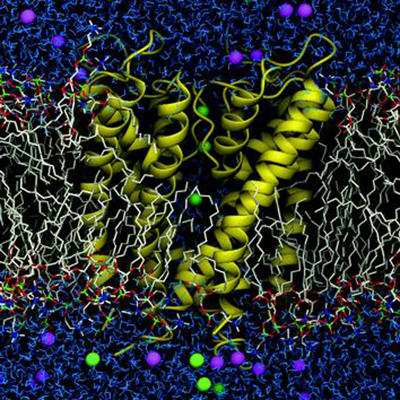
Biophysics of Membranes and Membrane Proteins
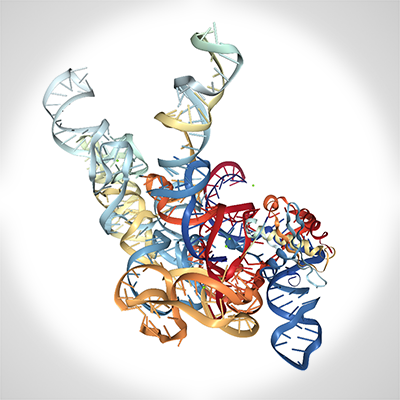
Biophysics of Nucleic Acids and Nucleoprotein Complexes
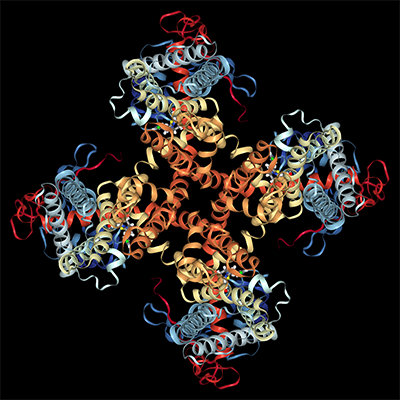
Biophysics of Proteins
Folding, Interactions, Structure, Dynamics & Mechanisms
Biophysical Studies of Supramolecular Complexes
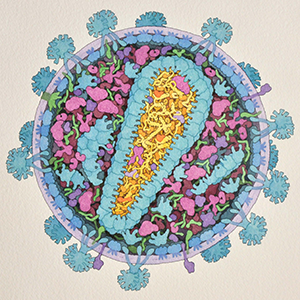
Biophysical Studies of the Viral Life Cycle
Molecular Modeling, Theory, and Design
Funding Opportunities, Research Resources, and Items of Interest
For more information about biophysics programs, contact:
Paula Flicker, Ph.D.
Chief, Biophysics Branch
Division of Biophysics, Biomedical Technology, and Computational Biosciences
National Institute of General Medical Sciences
National Institutes of Health
45 Center Drive MSC 6200
Bethesda, MD 20892-6200
